Team:Calgary/Project/OSCAR/FluxAnalysis
From 2012.igem.org
Emily Hicks (Talk | contribs) |
Emily Hicks (Talk | contribs) |
||
| Line 118: | Line 118: | ||
<h2>Code</h2> | <h2>Code</h2> | ||
| - | <p>This application is a Matlab extension that runs on top of Cobra Toolbox and SBML Toolbox. To run the application, one must have Cobra Toolbox and SBML Toolbox installed. SBML Toolbox can download from <a href="http://sbml.org/Software/SBMLToolbox">SBML.org</a> or <a href="http://sourceforge.net/projects/sbml/files/SBMLToolbox/4.1.0/SBMLToolbox-4.1.0.zip/download">here</a>. Cobra Toolbox can download from <a href="http://opencobra.sourceforge.net/openCOBRA/Welcome.html">openCOBRA</a> or <a href="http://sourceforge.net/projects/opencobra/files/cobra/cobra_2.0.5.zip/download">here</a>. Application Package and source code<a href="https://static.igem.org/mediawiki/2012/5/50/UCalgary2012_OSCAR_optimizer_v1.zip"> download </a>.</p> | + | <p>This application is a Matlab extension that runs on top of the Cobra Toolbox and SBML Toolbox. To run the application, one must have the Cobra Toolbox and SBML Toolbox installed. SBML Toolbox can download from <a href="http://sbml.org/Software/SBMLToolbox">SBML.org</a> or <a href="http://sourceforge.net/projects/sbml/files/SBMLToolbox/4.1.0/SBMLToolbox-4.1.0.zip/download">here</a>. Cobra Toolbox can download from <a href="http://opencobra.sourceforge.net/openCOBRA/Welcome.html">openCOBRA</a> or <a href="http://sourceforge.net/projects/opencobra/files/cobra/cobra_2.0.5.zip/download">here</a>. Application Package and source code<a href="https://static.igem.org/mediawiki/2012/5/50/UCalgary2012_OSCAR_optimizer_v1.zip"> download </a>.</p> |
<br> | <br> | ||
Revision as of 03:19, 4 October 2012


Hello! iGEM Calgary's wiki functions best with Javascript enabled, especially for mobile devices. We recommend that you enable Javascript on your device for the best wiki-viewing experience. Thanks!
Flux-Variability Analysis for Optimization
Flux-variability analysis (FVA) was applied to optimize the bioreactor system of OSCAR for the newly incorporated metabolic pathways (Decarboxylation, Decatecholization, Desulfurization, and Denitrogenation ). FVA combines the framework of metabolic pathways with experimental enzymatic data to provide a computational platform for predicting what changes in metabolite levels will result in increased end-product. We developed a model using MATLAB computer language for predicting what metabolites could be added to your growth media to increase production of hydrocarbons in the E. coli chassis. We also created a graphical user interface to make the the FVA program user-friendly and to allow current and future iGEM teams/scientists to input their own synthetic pathways into the FVA for analysis. Finally, we validated this model in the wet-lab by optimizing the PetroBrick (BBa_K590025) system to increase hydrocarbon production, thereby saving time, and resources.
Click here to download the MATLAB package to test our model!
Click here to view our OSCAR Optimizer Manual.
Background
Why FVA uses Flux Balance Analysis?
Flux Balance Analysis (FBA) uses linear programming of metabolic networks to convert each metabolite into a mathematical coefficient. These coefficients can be connected to each other by changes associated with each enzymatic step of the pathway. FBA can then apply a mathematical method to examine how metabolites relate to each other in the network and allows us to make generalized predictions about organism growth, product output, and metabolite levels inside the cell. FVA is an extension of FBA that determines the range of reactions that result in an optimal flux through the metabolic network. What this means is that unlike FBA which just determines the most optimized network for increasing cell growth, we can modify the system to make something else, such as product output, the component to be optimized.
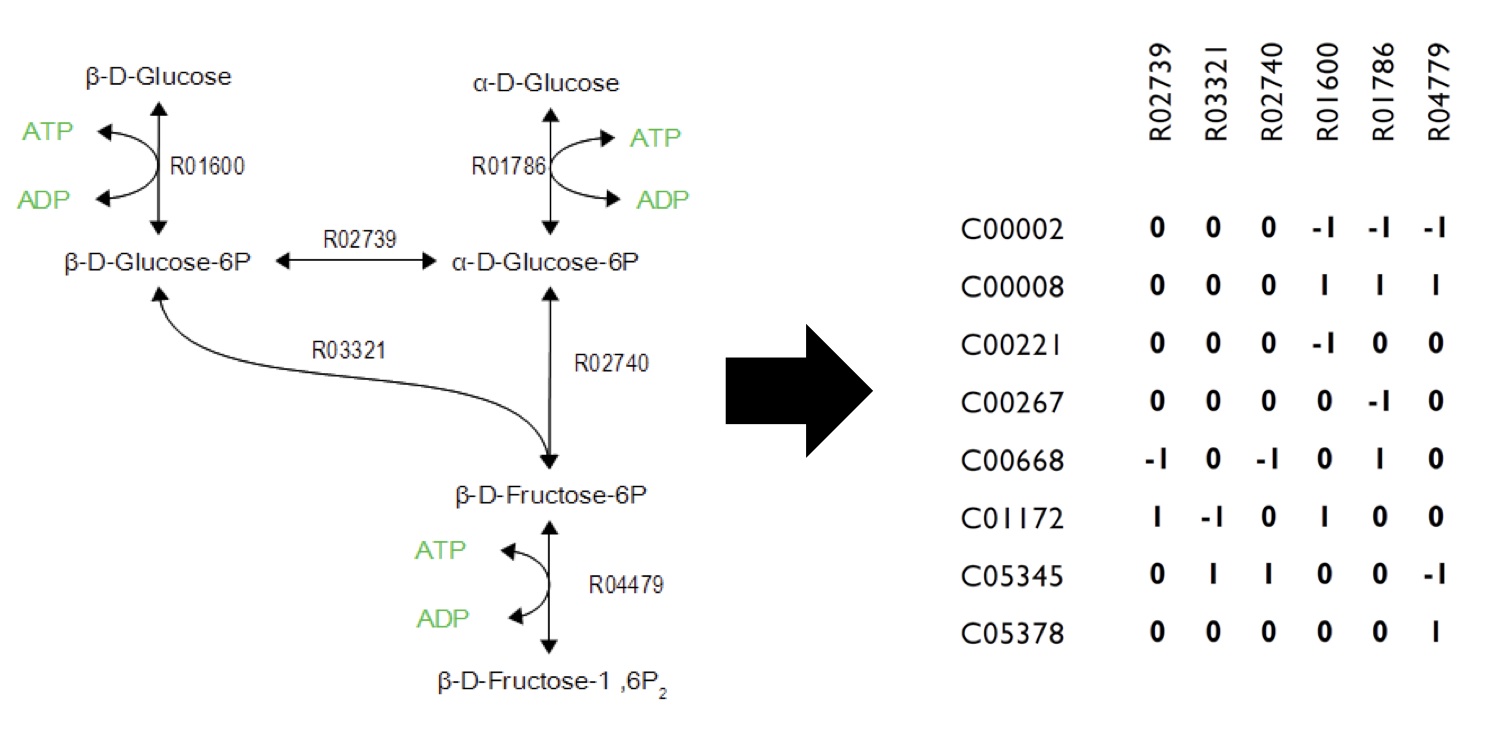
What Are The Constraints In The FVA Model?
As illustrated in Figure 1, metabolic networks can be encoded as stoichiometric matrices, in which each row represents a unique metabolite and each column represents a biochemical reaction. The entries in each column of this matrix are the stoichiometric coefficients of the metabolites in the reaction. Metabolites are consumed have a negative coefficient and metabolites that are produced have a positive coefficient.
Why use Flux Variability Analysis as Opposed To Flux Balance Analysis?
Biological systems often contain redundancies that contribute to their robustness. However, FBA only returns a single flux distribution that corresponds to maximal growth under given growth conditions regardless alternate optimal solutions may exist. FVA is capable of examining these redundancies by calculating the full range of numerical values for each reaction flux in a network. Consequently, FVA can be employed to study the entire range of achievable cellular functions as well as the redundancy in optimal phenotypes. FVA can also examine different ranges of bacterial growth vs. product output which is valuable in assessing validity of models in the wetlab.
Using FVA to optimize OSCAR
What Are We Trying To Model?
FVA provides a method to modulate the inputs in endogenous metabolic pathways or investigate what chemicals can be added to the growth media to upregulate an introduced pathway in E. coli. Development of a tool to model the optimal flux would also benefit numerous iGEM teams who engineer bacteria with novel pathways. To do this, we need to specifically model the flux rate of metabolic pathways responding to different growth media conditions and generate an optimal set of metabolites that should be added to growth media in order to improve production rate.
How Could Systems Like OSCAR Benefit From the Model?
Same as chemical reactions need optimal environmental conditions to achieve maximum production rate, microbes also require optimized growth conditions to accomplish their tasks in maximum speed. During industrial scale up, the optimal conditions for production needs to be maximized while reducing cost of production to a minimum. In microbiological bioreactor systems the conditions of growth media is much more crucial than in chemical synthesis reactors. Furthermore, the selection of media compounds is one of the most significant conditions for growth media and selecting a mix of compounds is very important for this process. If a model can predict an optimal set of metabolites that need to be added into media, this will save time, resources, and funds.
How Does The Program Work?
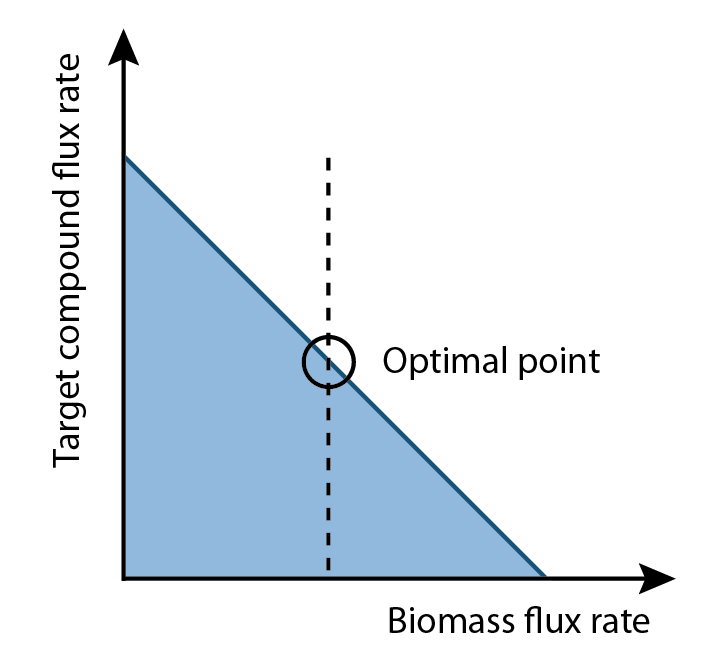
Our program is built upon flux variability analysis applied by the functions in the Cobra Toolbox. It uses the published E.coli iAF1260 and E.coli core models provided from the Palsson Group University of San Diego. Using this as a base, we constructed reactions and metabolites for our hydrocarbon production component of our project. Specifically, new reactions corresponding to the PetroBrick as well as the upgrading (desulfurization and denitrogenation) pathways were engineered into the E.coli base chassis. By running flux variability analysis, the program will give an output correlated to a optimized growth rate. By running the FVA algorithm at optimal biomass flux rates you can get a range of output for your product of interest (as illustrated in Figure 2). This allows you to determine an optimum point where compound production is maximized. Finally, the program will analysis the data with an algorithm to generate a set of media compounds that is expected to accelerate production rate.
Algorithm
Conceptualization
FVA can determine the full range of numerical values for each reaction flux within the network. Additionally, it allows for a broader analysis of growth and production rates compared to FBA. Since biomass rate reflects the growth condition, cells must have positive values of biomass flux rate in order to survive and proliferate. This positive growth rate is indicative of a real system as cells are optimized to prefer increases in growth rather than increases in product output. On the other hand, our goal is to increase the production flux rate above a zero value. This implied among all possible set of fluxes, the optimal flux set should locate a place where growth and production rate are at the optimum point for cell survival and compound output.
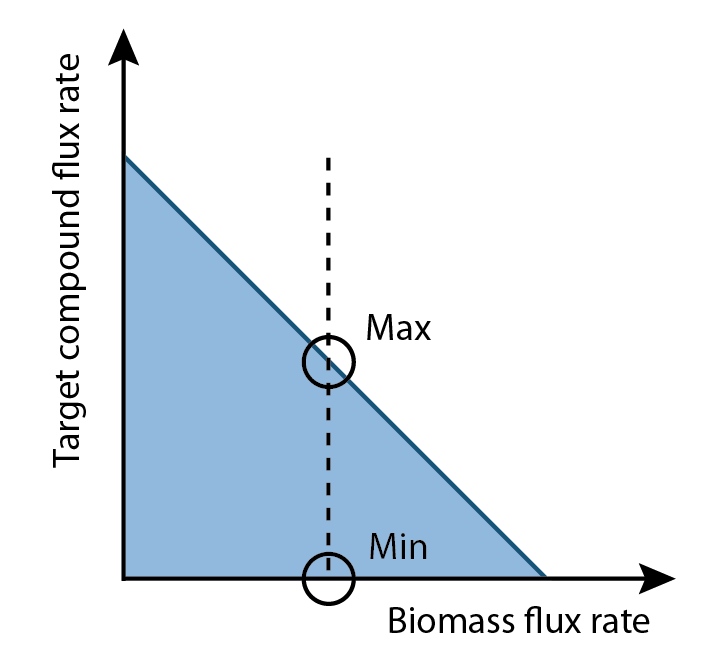
The differences of values for each reaction in a set of fluxes that maximize and minimize production rates can be compared. Reactions that demonstrated higher flux in the maximum production rate versus the minimum production rate represented substrates targets that could be increased in order to increase product output (as identified in Figure 3). Therefore we can create a list of compounds that should increase product output by increasing their concentration in solution.
However not all substrates can be uptaken by the cell. Hence, only the metabolites that had natural transporters in the cell were considered in our finalized list of compounds. We used a model based on glucose minimal media, however, this could be applied to any type of media.
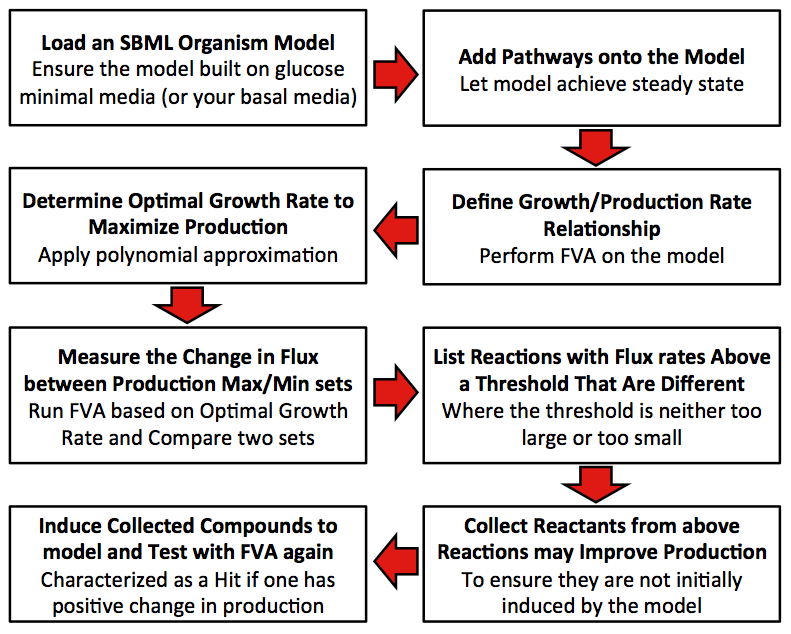
Model Steps
Precondition: The original model is built with glucose minimum media.
1. Define relationship between growth rate and production rate.
2. Find out the optimal growth rate that can maximize the production.
3. Get the difference percentage of flux rate for each reaction between production maximum set and production minimum set.
4. Collect all reactions with different percentages between two sets that exceed the user input threshold. (The input threshold determines the size of the difference in flux that the user is interested in. We used a 100% difference in our model.).
5. Score each compound in all collected reactions (Initial score is zero for each compound). (Scoring is determined by the difference in the value of a particular compound's value when the user's production of the compound of interest is maximized and minimized flux sets as determined by the FVA) This process is additive if the particular compound is found in more than one reaction (i.e. ATP is found in many reactions sets) and only includes the reactions identified from step 4.
6. Determine whether compounds with positive scores have natural transporters in cell. If so, mark the compound as candidate.
7. Add each candidate to the growth media and the run FVA under optimal growth rate computed in Step 2. Compare the production rate from novel model (step 7) to that from the raw model (step 2), if the rate is improved, mark the compound as an effector.
Graphical User Interface (GUI) Development
A graphical user interface allows for easier use of our program by everyone in iGEM and beyond. This will open up our program to be used by individuals who do not have experience with programs such as MATLAB or the Cobra Toolbox. We developed the GUI using the GUIDE program designed in MATLAB. For more information please see our manual.
Demo
Here we have uploaded a video showing some of the screens for using the model as a basic tutorial for teams to see how our program is used. The GUI interface allows for easy building of different synthetic constructs into the E. coli network but this could use any model from any organism that is available in SBML format.
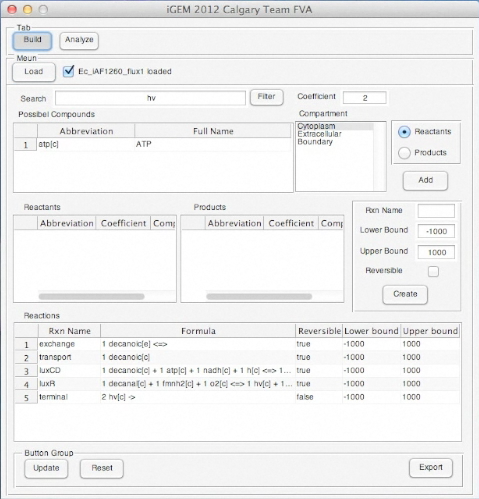
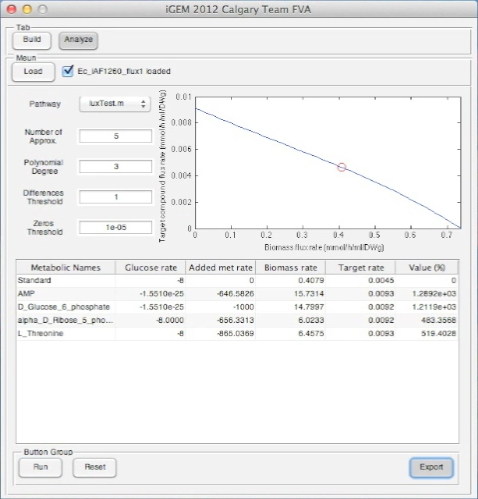
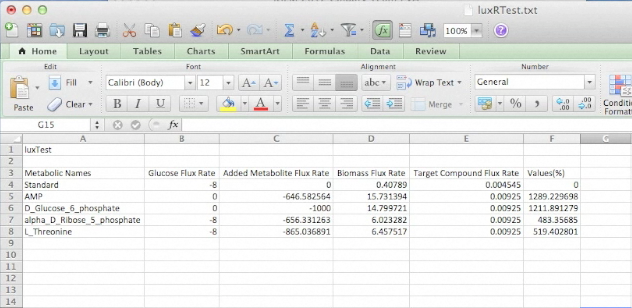
Screen Shots of Our Application
Wetlab Validation of the Model
In order to assess if our program could be used to identify compounds that would selectively increase the output of a target compound, we thought it would be important to test this in a wetlab system. To do this we used the PetroBrick construct and attempted to increase it's output by supplementing the media with particular components predicted by our model to increase hydrocarbon output. Therefore we ran the model with the AAR and ADC gene components of the Petrobrick system and looked at the predicted metabolites that we should add to solution. This is illustrated in the figure below.
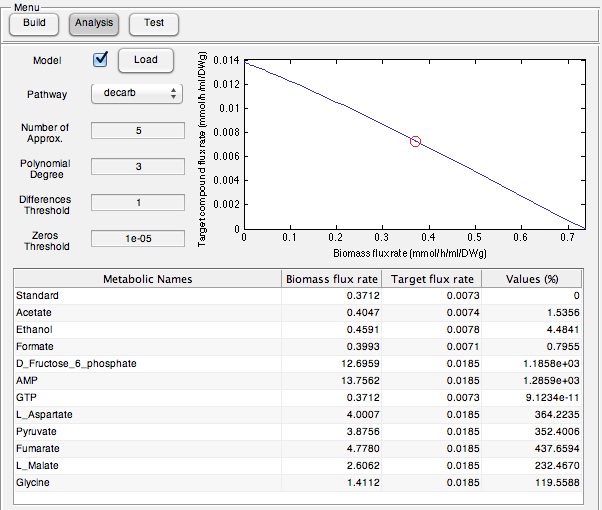
Once these compounds were identified, we set up an assay where we supplemented minimal M9 media and glucose with each of the compounds alone, or in combinations. Compounds were added at concentrations of 50 mM except for Ethanol (2.5% v/v), AMP (100mg/L), and L-aspartate (100mg/L). As a positive control, instead of using minimal media a solution of 50:50 LB:Washington Production Media (see protocols section for composition) was used to exhibit what normal production looked like. Additionally, we found a compound that was predicted to NOT increase the production of hydrocarbons, formate, and used this as a control. These compounds were allowed to incubate with the PetroBrick starting at an OD600 of ~0.05 and grown for 72 hours at 37oC. Once grown, OD600 measurements were taken prior to sonication of the samples and extraction of any produced hydrocarbons using 1mL of ethyl acetate. This was quantitated based on the peak area of a C15 hydrocarbon product which is described on our decarboxylation wiki page (also see protocols for relevant procedures dealing with this section). Once quantitated these result yielded the following:
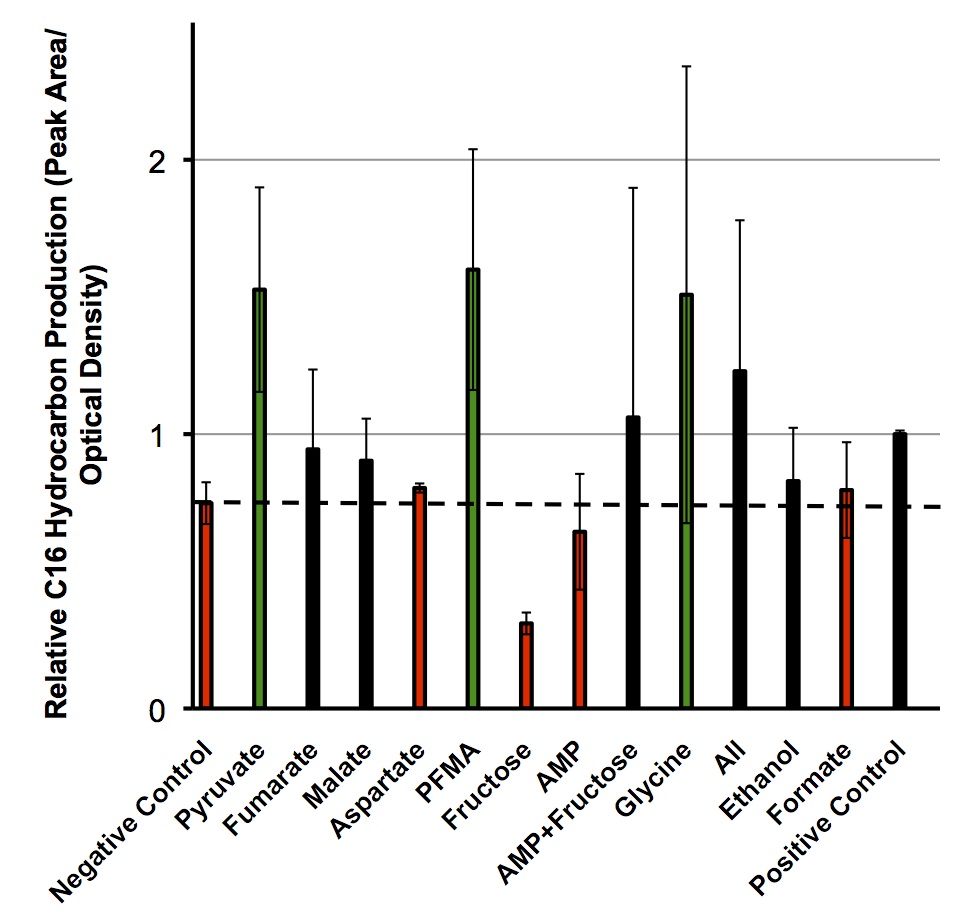
These results suggest that there is some natural variability in the output of hydrocarbons from the PetroBrick with different compounds. It was interesting to observe that five of the compounds demonstrated production levels higher than that of the minimal media control which suggested that our model was correct for predicting these compounds. However AMP and Fructose which both were thought to have shown increases in hydrcarbon output demonstrated large decreases suggesting that while our model may make some correct predictions there is clearly some error in assessing these predictions. What was very exciting however, is that for two of these compounds (pyruvate and glycine) their addition to the growth media increased the relative number of hydrocarbons higher than that of a complex media as in the positive control. This suggests that we can indeed use our model to optimize OSCAR along with other metabolic systems, however, these results should be tested in the wetlab to ensure the model is predicting them accurately.
Drawbacks
This application is built upon the Cobra Toolbox, and the SBML Toolbox. As a consequence, any flaws in the Cobra Toolbox and SBML Toolbox will affect this application.
In this program, pathways added to base chassis model (E. Coli iAF1260) which contains constraints that rely on the Stoichiometric Matrix (such as Stoichiometric coefficients), lower bounds and upper bounds of reactions. They lack genetic and enzymatic regulation, which makes the connections between reactions in the network much weaker than those in the wetlab. The missing rules could lead to inaccuracy results.
At current stage, the algorithm can only pick metabolites with natural transporters in cell. Many other intermediate metabolites are ignored. The algorithm has no power to trace the intermediate metabolites back to initial metabolites and take those initial metabolites into account. This model does not take into consideration rate limiting steps of reactions, toxic intermediate accumulation, or any form of regulation of the enzymes (an example being negative feedback). The model should be used as a starting point to decrease a bottleneck from the amount of naturally available metabolites in E. coli to increase the amount of starting compound your synthetic circuit can use.
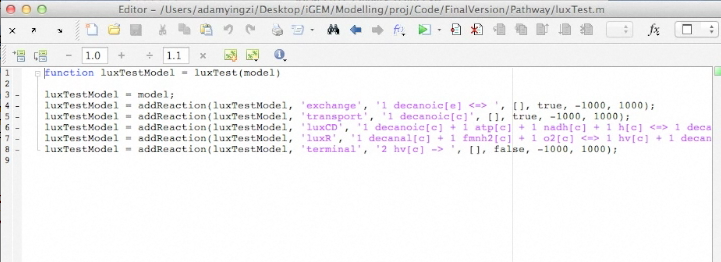
Code
This application is a Matlab extension that runs on top of the Cobra Toolbox and SBML Toolbox. To run the application, one must have the Cobra Toolbox and SBML Toolbox installed. SBML Toolbox can download from SBML.org or here. Cobra Toolbox can download from openCOBRA or here. Application Package and source code download .
Documents
Download the OSCAR Optimizer Manual for more information on how to use the program!
 "
"