Team:Calgary/Project/OSCAR/Decarboxylation
From 2012.igem.org


Hello! iGEM Calgary's wiki functions best with Javascript enabled, especially for mobile devices. We recommend that you enable Javascript on your device for the best wiki-viewing experience. Thanks!
Decarboxylation
Why Decarboxylation?
Though there is great diversity between the many possible compounds termed “naphthenic acids” found in tailings ponds, all share the common chemical feature of a carboxylic acid group. This particular group is known to be the primary cause for the toxicity of these compounds, allowing them to enter through cell membranes and destroy cells. Naphthenic acids are a very problematic component of the tailings ponds, and are not easily degraded under regular circumstances. They are known to leak into local environments and cause many problems for wildlife.
Additionally, removing the carboxylic acid group(s) is of great importance in the conversion of these contaminants to biofuel. Since naphthenic acids are a widely varying and unspecific mixture of compounds, an enzymatic process with very low substrate specificity is necessary, as particularly specific enzymes would not be able to universally decarboxylate all naphthenic acid substrates found in tailings ponds. Without a carboxylic acid group, and with the removal of sulphur and nitrogen contaminants, an alkane suitable for use as fuel can be obtained. The goal of this subproject was to find a suitable pathway to accomplish the decarboxylation of naphthenic acids with the broadest specificity possible.
The PetroBrick
As part of their submission for the 2011 iGEM competition, the University of Washington included a part called the PetroBrick, consisting of two genes intended to establish a metabolic pathway to convert fatty acids to alkanes in vivo. The first gene in the pathway encodes acyl-ACP reductase (AAR), reducing fatty acids bound to ACP to fatty aldehydes, while the second encodes aldehyde decarbonylase (ADC) to subsequently cleave the entire aldehyde group, resulting in a hydrocarbon chain.
Since the end goal of the decarboxylation project was fuel production, and because the University of Washington’s project provided an easily accessible route to a decarboxylation pathway already available in the BioBrick registry, the PetroBrick was selected as a starting point to this issue. The problem with using the PetroBrick to decarboxylate naphthenic acids was immediately apparent: the first enzyme in particular (AAR) was highly specific for fatty acids bound to ACP. While this is not a concern for producing alkanes from fatty acid biosynthesis, it was not considered to be particularly compatible with naphthenic acid degradation. Therefore we sought an alternate reductase to accompany AAR in the pathway, which would be the CAR gene.
Meanwhile, testing proceeded with the PetroBrick. Cultures were grown in a production media designed by Washington in 2011 to be conducive towards alkane production and incubated for 48 hours. Afterwards, an extraction with ethyl acetate was performed to separate the alkanes. Not only was production of alkanes from glucose successful, as the University of Washington had earlier demonstrated, but we were able to optimize the PetroBrick to produce alkane and alkene hydrocarbons from naphthenic acids.
Results of the PetroBrick
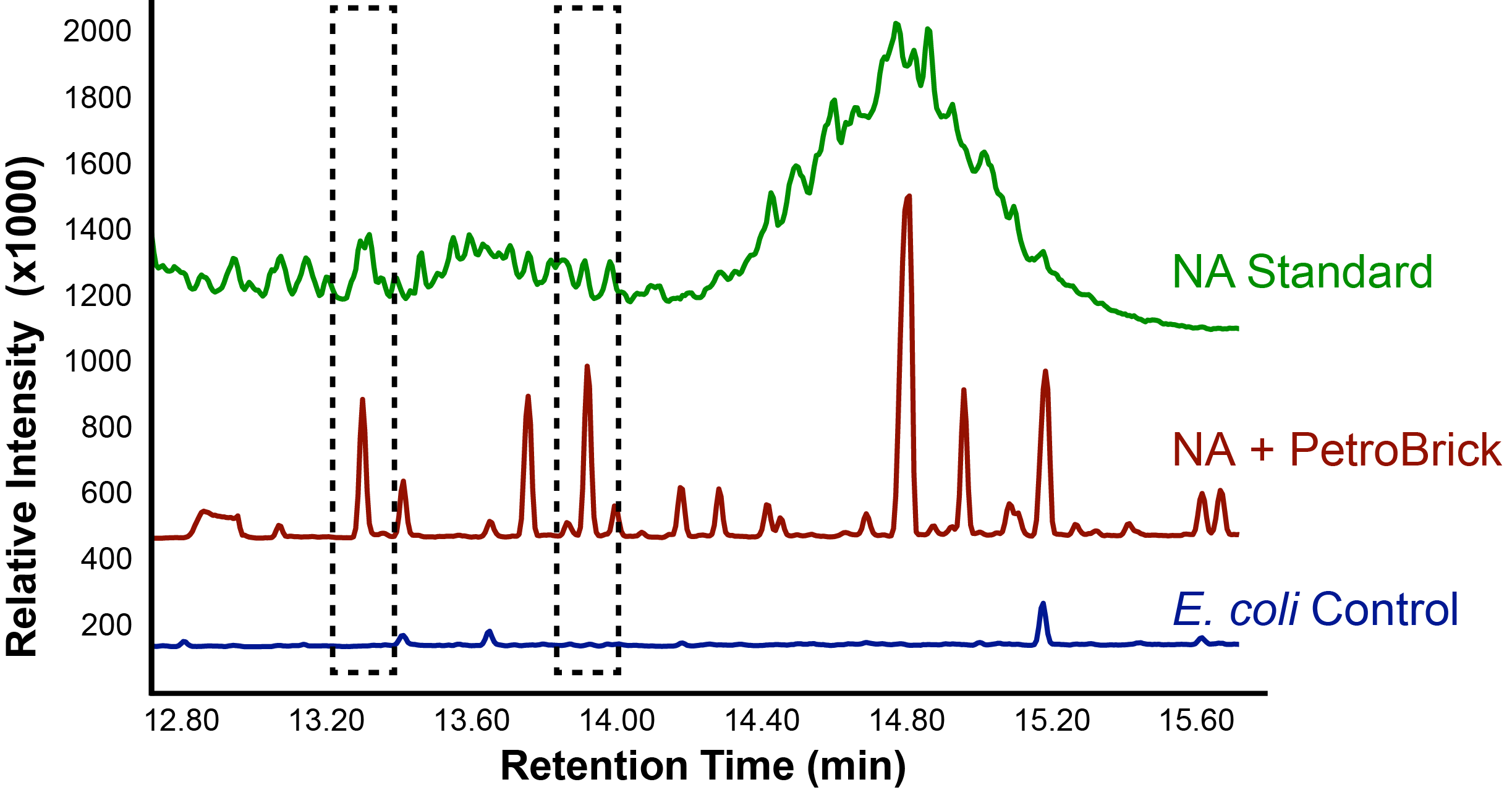
Figure 1: The relative intensity of alkane production over a retention time in both E.coli that contain the PetroBrick, and in E.coli that are lacking the PetroBrick, as measured with GC-MS. Naphthenic acids were used as a substrate. A naphthenic acid standard was required to compare peaks.
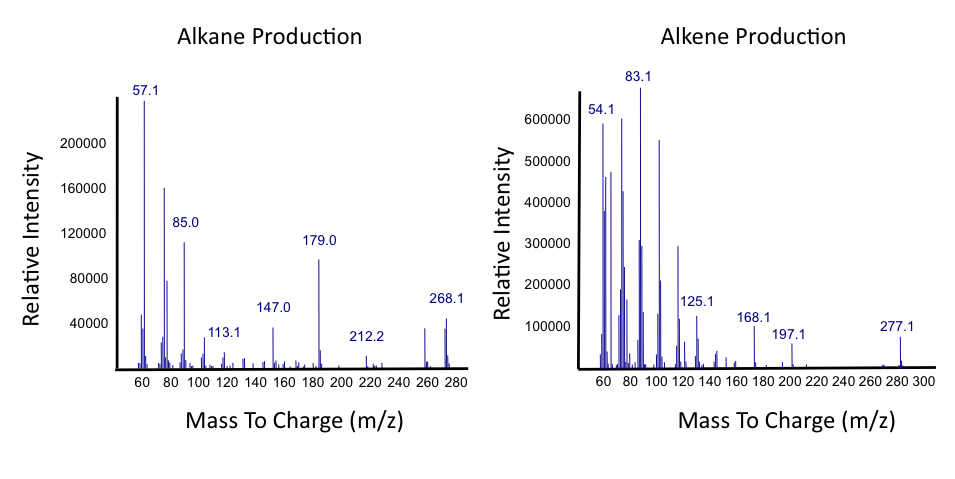
The above graphs indicate that hydrocarbons were successfully produced from E.coli that contained the PetroBrick plasmid, as analysed with GC-MS. In Figure 1, E.coli containing the PetroBrick had significantly higher hydrocarbon peaks than in a control of E.coli that did not contain the PetroBrick plasmid. Not only was the PetroBrick able to degrade naphthenic acids into alkanes, but it was also able to produce alkenes as shown by Figure 2, indicating that the PetroBrick had more potential than previously indicated.
Nocardia Carboxylic Acid Reductase (CAR)
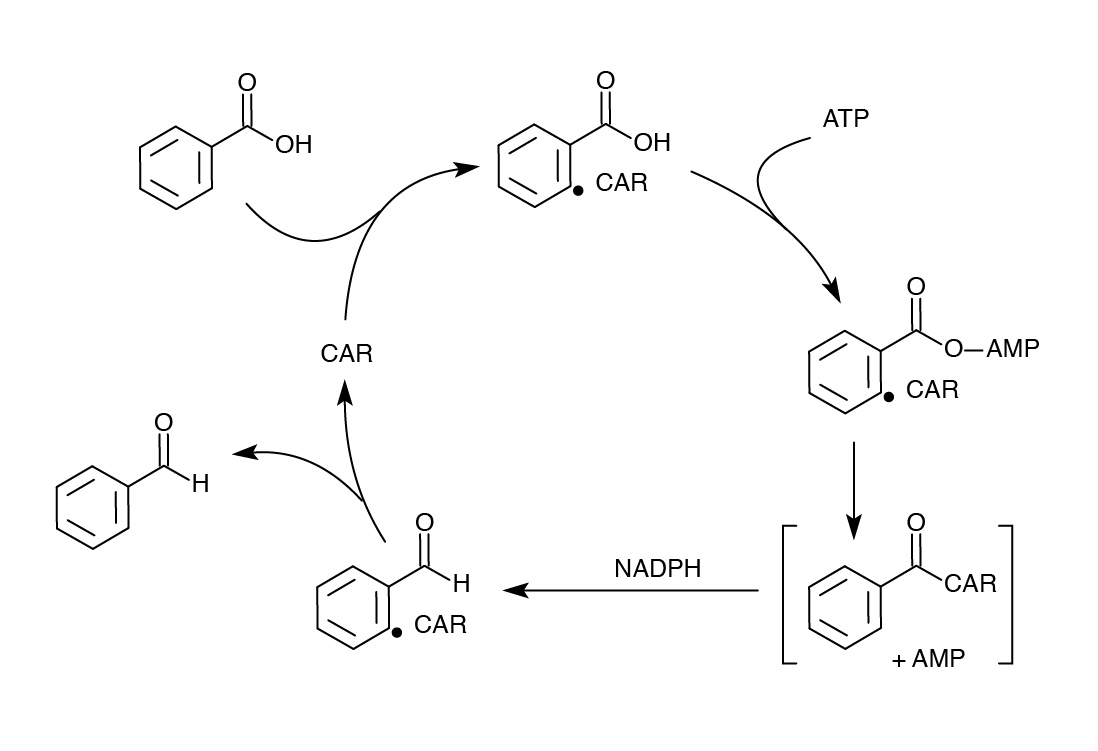
In order to model our part after the PetroBrick, an enzyme was needed to replace AAR that would not require covalent attachment to ACP and would have low enough specificity to accommodate diverse naphthenic acids. The enzyme chosen for this purpose was carboxylic acid reductase (CAR), found in Nocardia iowensis. CAR was selected for its remarkably low specificity in converting carboxylic acids to aldehydes. It was determined that a second gene from N. iowensis, called Nocardia phosphopantetheinyl transferase (NPT) was also necessary to append a 4’-phosphopantetheine prosthetic group to CAR required for its full function.
N. iowensis (NRRL 5646) was purchased from DSMZ and rehydrated, then grown in both solid and liquid brain heart infusion (BHI) media. CAR and NPT were successfully cloned out and amplified, and CAR was verified after ligation with the PET vector by a restriction digest and subsequent gel. NPT was likewise verified.
CAR was also found to have six cut sites deemed illegal for BioBrick construction: one XpaI site, two EcoRI sites, and three NotI sites. These were first to be addressed by a multi-site mutagenesis derived from the QuikChange® Multi Site Directed Mutagenesis Kit, but this showed little success. Instead, a more time-consuming series of conventional single-site mutagenesis procedures was favoured, using the KAPA amplification system. A PET vector was chosen as an alternate vector to pSB1C3 to ligate with CAR for the duration of mutagenesis, while NPT was still ligated with pSB1C3. The XpaI and EcoRI sites were eliminated first, and following these steps and subsequent verification of their success, the mutated CAR+PET plasmid was successfully amplified with BioBrick primers after multiple attempts in preparation for the final construct. NPT was ligated in a construct following part J13002 from the iGEM parts registry, containing a ribosomal binding site and a TetR promoter.
How do the PetroBrick and CAR+NPT work together?
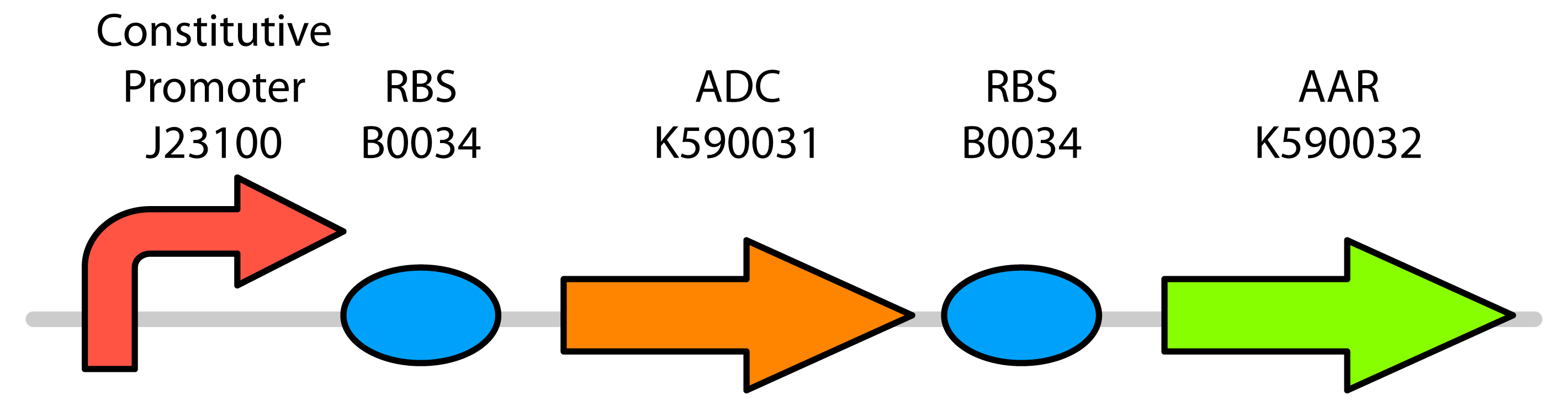
Our team decided to created a BioBrick construction containing three major parts. These included: the PetroBrick (as received from Washington), the CAR gene, and the NPT gene. This construction can be seen below.
Both CAR and NPT use the part JI3002, which consists of pTetR (a constitutive promotor), as well as an RBS (ribosomal binding site). This constitutive promoter allows the gene to always be expressed, so that decarboxylation may consistently occur. The construction also includes Washington’s PetroBrick, which contributes the genes ADC and AAR. While AAR is useful to help convert some carboxylic acids and fatty acids to aldehydes, it is not known to be highly specific like the CAR enzyme. We are primarily interested in using the PetroBrick for its ADC gene, which is responsible for producing the only enzyme in our system that is capable of converting aldehydes into hydrocarbons. Meanwhile, both AAR and CAR in our construct will work to convert the initial carboxylic acids into the aldehydes required to produce hydrocarbons.
What about an alternative? (Jeotgalicoccus olefin-forming fatty acid decarboxylase, or OleT)
In the early stages of work with CAR, an alternate approach to naphthenic acid decarboxylation was proposed that was entirely distinct from the PetroBrick+CAR/NPT system. The idea behind this method was to use olefin-forming fatty acid decarboxylase (OleT) from Jeotgalicoccus sp. ATCC 8456, a decarboxylase of the cytochrome P450 family acting on fatty acids, but with low substrate specificity. We chose to investigate OleT to determine if it was capable of directly decarboxylating naphthenic acids.
OleT was successfully amplified from the Jeotgalicoccus sp. ATCC 8456. Like CAR, OleT was inserted in a PET vector before placing it into a BioBrick, as two illegal cut sites adjacent to one another needed to be mutagenized. The final construct for OleT is very simple, containing the J13002 promotor/RBS, and the OleT gene. This simple construct should be able to convert carboxylic acids to terminal alkenes on its own.
References
https://www.dsmz.de/catalogues/details/culture/DSM-45197.html (retrieved 8/28/2012)
https://2011.igem.org/Team:Washington/Alkanes/Background (retrieved 8/28/2012)
http://partsregistry.org/wiki/index.php?title=Part:BBa_K590025 (retrieved 8/28/2012)
He, A., Li, T., Daniels, L., Fotheringham, I., and Rosazza, J.P.N. 2004. Nocardia sp. Carboxylic Acid Reductase: Cloning, Expression, and Characterization of a New Aldehyde Oxidoreductase Family. Applied and Environmental Microbiology 70:1874–1881.
Venkitasubramanian, P., Daniels, L., and Rosazza, J.P.N. 2006. Reduction of Carboxylic Acids by Nocardia Aldehyde Oxidoreductase Requires a Phosphopantetheinylated Enzyme. Journal of Biological Chemistry 282:478-485.
Rude, M.A., Baron, T.S., Brubaker, S., Alibhai, M., Del Cardayre, S.B., and Schirmer, A. 2011. Terminal olefin (1-alkene) biosynthesis by a novel p450 fatty acid decarboxylase from Jeotgalicoccus species. Applied and Environmental Microbiology 77:1718–1727.
 "
"