|
|
| (84 intermediate revisions not shown) |
| Line 1: |
Line 1: |
| | {{Team:Calgary/TemplateProjectBlue| | | {{Team:Calgary/TemplateProjectBlue| |
| - | TITLE=Catechol Degradation| | + | TITLE=Decatecholization| |
| | | | |
| - | <h3>Week 1 (May 1-4)</h3> | + | CONTENT=<html> |
| - | <p>The following section covers wetlab aspect of our overall project focusing on microbial conversion of naphthenic acids into economically-valuable hydrocarbons. The approach taken in this endeavour will be from four strategic starting points - ring cleavage, decarboxylation, denitrificaiton, and desulfurization. Overall, the 'hydrocarbons' aspect of the project is a critical one to our overall design and construction of a biosystem capable of not only detecting but also converting naphthenic acids in what will be an economically viable solution to the remediation and recovery of tailings pond water, while removing these toxic compounds before they can have significant detrimental impact on the environment. </p> | + | <img src="https://static.igem.org/mediawiki/2012/1/1c/UCalgary2012_OSCAR_Catechol_Low-Res.png" style="float: right; padding: 10px;"></img> |
| | | | |
| - | <h2>Week 2 (May 7-11)</h2>
| |
| - | <h3>Ring Cleavage</h3>
| |
| - | <p>This week we mainly researched aromatic ring cleaving using intra- and extradiol dioxygenases from species of Pseudomonas and Bacillus but also began literature searches on aliphatic ring cleavage done by monooxygenases. </p>
| |
| | | | |
| - | <h2>Week 3 (May 14-18)</h2> | + | <p>Catechol is a toxic compound found in tailings ponds that is a by-product of polyaromatic hydrocarbon metabolism (Vaillancourt <i>et al.</i>, 2006, Schweigert <i>et al.</i>, 2001)). The chemical properties of catechol allow it to react with biomolecules, causing cellular damage including DNA damage, enzyme inactivation and membrane uncoupling (Schweigert <i>et al.</i>, 2001). </p> |
| - | <h3>Ring Cleavage</h3>
| + | <p> |
| - | <p>In the second week we continued to do more research on the degradation of alicyclic compounds and found two strains of bacteria that contained genes needed for this process. The first strain, Thauera butanivorans, contains the genes required to activate the ring by adding a hydroxyl group. The gene is called butane monooxygenase and is composed of three subunits, a hydroxylase, a reductase, and a regulatory component. The second strain, Acinetobacter sp. SE19, contains the genes needed to oxidize the alcohol, formed by butane monooxygenase, and to cleave the ring. There is a cluster of nine genes that perform this oxidation and cleavage but only six are involved directly.</p> | + | Catechol is characterized as having a benzene ring with two hydroxyl groups at the 2,3 position. It can be converted to 2-hydroxymuconic acid by the enzyme catechol 2,3-dioxygenase, encoded by the <i>xylE</i> gene on the Tol plasmid of <i>Pseudomonas putida</i> (Nakai <i>et al.</i>, 1983).</p> |
| | | | |
| - | <h2>Week 4 (May 22-25)</h2> | + | <p> |
| - | <h3>Ring Cleavage</h3> | + | Currently the registry has two BioBricks available of <i>xylE</i>. One contained <i>xylE</i> with its native ribosome-binding site (<a href=http://partsregistry.org/Part:BBa_J33204>BBa_J33204</a>), while the other part contained <i>xylE</i> under the glucose-repressible promoter <i>cstA </i>(<a href=http://partsregistry.org/Part:BBa_K118021>BBa_K118021</a>). Given that <i>E. coli</i> is grown in the presence of glucose, we designed a new construct to keep <i>xylE</i> expressed by using the <i>tetR</i> promoter (<a href= http://partsregistry.org/Part:BBa_R0040>BBa_R0040</a>).</p> |
| - | <p>We started looking into more organisms that can carry out cyclohexane degradation and found one called Brachymonas petroleovorans. This organism is capable of both hydroxylating cyclohexane and carrying out cyclohexanol oxidation. We are also hoping to find evidence that Pseudomonas fluorescens, Pf-5 has genes that can cleave aliphatic rings, however, first we have to determine if Pf-5 can grow in the presence of butylcyclohexane. To do so we carried out an assay, using LB, testing the toxicity of butylcyclohexane on Pf-5. After 24 hours of incubation we took OD600 readings¬ which suggested that the presence of butylcyclohexane did not have effect on cell growth. We also looked up Pseudomonas fluorescens Pf-5 in the Pseudomonas Genome Database and found genes that code for proteins with similar functions to what we have found for aromatic and aliphatic ring cleavage. </p> | + | |
| | | | |
| - | <h2>Week 5 (May 28 - June 1)</h2> | + | </html>[[File:UCalgary2010_R0040-XylE.png|400px|thumb|Figure 1: BioBrick genetic circuit for catechol degradation showing <i>xylE</i> under the ''tetR'' promoter|center]]<html> |
| - | <h3>Ring Cleavage</h3> | + | <h3></h3> |
| - | <p>This week we redid the previous assay with Pf-5 and butylcyclohexane using glass vials instead of falcon tubes. These vials allowed for more surface and better containment of the volatile compound. We carried out this assay using both LB and M9-MM. After 24 hours we took OD600 readings and obtained similar results as last week. The growth was considerably decreased in the M9-MM, however we are not sure whether the bacteria were able to use the compound as a carbon source because the MM contained glucose. The fact that the growth decreased showed that they might depend on the glucose in the media to grow. </p> | + | |
| | | | |
| - | <h2>Week 6 (June 4 - June 8)</h2>
| + | <p>Catechol 2,3-dioxygenase is an extradiol dioxygenase which cleaves catechol adjacent to the two hydroxyl groups. When this occurs 2-hydroxymuconate semialdehyde is produced, which is yellow in colour. This change in colour allows for visual assay to assess the activity of XylE.</p> |
| - | <h3>Ring Cleavage </h3>
| + | |
| - | <p> This week we made subcultures from Pf-5 in M9-MM into M9-MM without glucose to avoid carryover of glucose, to insure that the bacteria use butylcyclohexane as their only carbon source. We also took GC readings and compared experimental values to standards made with water and butylcyclohexane (butCH). Readings showed a decrease in amount of butCH in the headspace, compared to standards, indicating that the compound was being degraded by Pf-5. Then, we made subcultures from Pf-5 in M9-MM without glucose into the same media and butCH to decrease carry over of glucose and did time zero readings. Furthermore, we made subcultures from original Pf-5 in M9-MM without glucose into the same media and toluene to test for pathways that degrade aromatic structures. To identify possible genes for the pathways we started running blasts on the Pf-5 genome looking for genes that are homologous to ones that we have been looking at for the two ring cleavage pathways (aromatic and aliphatic). In addition we started a verification of XylE (aromatic ring cleavage enzyme) by transforming part K118021 (XylE with a Pcst promoter) from the parts registry and did two colony PCRs. unfortunately none of them showed successful results and only the positive control showed an appropriate length DNA. </p> | + | |
| | | | |
| - | <h2>Week 7 (June 11 - June 15)</h2> | + | </html>[[File:UCalgary2012_Catechol_to_2-HMS.PNG|400px|thumb|Figure 2: Catechol 2,3-dioxygenase (XylE) converts catechol to 2-hydroxymuconate semialdehyde in the presence of oxygen. Adapted from Shu <i>et al</i>., 1995.|center]]<html> |
| | | | |
| - | <h3>Ring Cleavage</h3> | + | <p>The visual assays were performed with <i>E. coli</i> cells transformed with (<a href=http://partsregistry.org/Part:BBa_K118021>BBa_K118021</a>) as well as with <i>E. coli</i> cells transformed with the newly constructed part (<a href=http://partsregistry.org/Part:BBa_K902048 >BBa_K902048</a>) by bringing the supernatant of an overnight culture to a concentration of 0.1 M of catechol. When the part (<a href=http://partsregistry.org/Part:BBa_K118021>BBa_K118021</a>) was used, the pellet was first washed in M9-MM and centrifuged before catechol was added to the supernatant. This was necessary to avoid the glucose in the LB from repressing the cstA promoter (<a href=http://partsregistry.org/Part:BBa_K118011>BBa_K118011</a>). Catechol was added to the supernatant because the reaction takes place outside of the cell. Within minutes of the addition of catechol to the supernatant, the solution turned from the pale yellow of LB to a bright yellow. This was indicative that catechol was breaking down into 2-hydroxymuconate semialdehyde, which was exactly what we expected! This assay was completed by following the protocol written by the 2008 Edinburgh iGEM team.</p> |
| | | | |
| - | <p> Since last week we continued taking GC readings on the Pf-5 incubated with butylcyclohexane and toluene. At this point the Pf-5 had been incubated with butylcyclohexane for 6 days and with toluene for 5 days. The amount of both butylcyclohexane and toluene decreased by a large amount in the experimental cultures but the amount of each in the abiotic standards also largely decreased by about the same amount. This lead us to conclude that the reason for decrease was not due to the bacteria metabolizing the compound, but due to unknown abiotic factors. To measure relatie amounts of bacterial growth we took OD600 readings for both cultures which were very low and indicated that there was little bacterial growth. The verification of XylE from the parts registry was continued using a miniprep on three overnight cultures from part K118021. The concentrations were very high likely due to genomic contamination. A second part (J33204) containing XylE and an rbs site was transformed and a colony PCR was done on part J33204 but the gel did not indicate successful results, again only the positive control worked. No positive colonies were found.</p> | + | </html>[[File:UCalgary2012_Catechol_assay.jpg|500px|thumb|Figure 3: Results of the catechol visual assay using ''xylE'' [http://partsregistry.org/Part:BBa_K118021 BBa_K118021]. Cultures were grown overnight in LB and the pellets were washed with M9-MM at various times (From left to right: 0 min, 5 min, 10 min, 15 min, and 20 min.). Cells were then spun down and catechol was added to the supernatant to 0.1 M. The amount of time didn't affect the colour change in the cultures containing the <i>xylE</i> gene. The far right tube has <i>E. coli</i> cells without the <i>xylE</i> gene as a negative control and the supernatant remained clear when the catechol was added. |center]]<html> |
| | | | |
| - | <h2>Week 8 (June 18 - June 22)</h2> | + | <a name="Catechol"></a><h2> Converting Catechol into hydrocarbons? </h2> |
| | + | <p>After verifying that we could in fact degrade catechol into 2-hydroxymuconate semialdehyde using our <i>xylE</i> construct (<a href=http://partsregistry.org/Part:BBa_J33204>BBa_J33204</a>), we wondered if we could take this any further. What if we could convert this by-product into hydrocarbons too? As catechol is the breakdown product of a number of different degradation pathways in bacteria, this could be particularly useful.</p> |
| | | | |
| - | <h3> Ring Cleavage </h3>
| + | <p>Since 2-hydroxymuconate semialdehyde can be further metabolized to pyruvate and acetaldehyde (Harayama et al., 1987), it seemed possible that these products could be routed into the fatty acid biosynthesis pathway and converted to alkanes using the PetroBrick or the OleT enzyme. Given that the catechol 2,3-dioxygenase reaction is extracellular, it creates a possible scenario in which cells with the <i>xylE</i> construct could be co-cultured with Petrobrick-containing cells to cooperatively metabolize catechol into hydrocarbons. </p> |
| - | <p>This week we started a new assay where Pf-5 was incubated with naphthalene. Since naphthalene is not volatile, GC headspace reading was not an appropriate method of measurement to indicate decrease of the compound. Instead we took OD600 readings to measure bacterial growth. The original cultures were taken from a Pf-5 culture in LB and inoculated into M9-MM without glucose. After 24 hours of growth these were subculutred into fresh M9-MM without glucose. This was to avoid carryover of glucose from the LB. Positive controls were also made that consisted of M9-MM with glucose and Pf-5. OD600 readings of the subcultures were taken over a period of 8 days. GC readings were done on the Pf-5 incubated with butylcyclohexane after 11 days and with toluene after 10 days. The trend remained the same as the measurements done after 5 and 6 days in the previous week. The amount of compound decreased in the experimental cultures but the amount of compound also decreased in the abiotic standards almost in equal proportions. With the results after 10 and 11 days we could see that the decrease was not due to the bacteria and we decided that the strain we were using did not contain the genes necessary for butylcyclohexane or toluene degradation. Throughout the week we continued the verification of XylE by performing a second colony PCR from the transformation plate using part J33204. The gel did not contain DNA of the right lengths and only the positive control worked. </p> | + | |
| | | | |
| - | <h2>Week 9 (June 25 - June 29)</h2> | + | <p> In order to test this, we followed this <a href=https://2012.igem.org/Team:Calgary/Notebook/Protocols/decatecholization>protocol</a>, where we co-cultured cells expressing our <i>xylE</i> construct with either <i>E. coli</i> cells expressing the PetroBrick, or <i>Jeotgalicoccus</i> sp. ATCC 8456 cells expressing <i>oleT</i> in the presence of catechol. |
| | | | |
| - | <h3>Ring Cleavage</h3>
| |
| | | | |
| - | <p> After the 8 day incubation the readings of the naphthalene assay were graphed and there was a definite increase in the cell growth of the cultures containing naphthalene as a carbon source with indicated that the bacteria were most likely were metabolizing the compound. A verification of another part, Alkane hydroxylase system, from the registry was started. The system consists of 4 genes that have been shown to hydroxylate C5-C8 cycloalkanes. We thought that this part would assist in the degradation of cyclohexane and cyclopentane. The transformation was successful and the gel of the colony PCR showed one positive colony but since there was contamination in the gel we were not sure if we could conclude that the PCR had worked. Final GC readings on the butch and toluene cultures were taken and results were graphed. | + | </html>[[File:Calgary PetrobrickCatechol.jpg|600px|thumb|centre|Figure 4: Gas chromatograph of catechol degradation assay using the PetroBrick. While there is limited differences between <i>xylE</i> incubated with and without the PetroBrick, there was one peak with a retention time of 10.5 min which was dramatically increased in the co-culture.]]<html> |
| - | </p> | + | |
| | | | |
| - | </html>[[File:UCalgary2012_Butchgraph.png|frame|center]] | + | </html>[[File:Calgary MSCatecholPetroPeak.jpg|450px|thumb|centre|Figure 5: Mass spectrum of the Petrobrick/<i>xylE</i> co-culture retention peak at 10.5 min as shown in Figure 4. While the identity of this compound is currently unknown, there are changes occuring to some of the catechol breakdown products.]]<html> |
| - | [[File:UCalgary2012_Toluene_assay.png|frame|center]]
| + | |
| - | [[File:UCalgary2012Naphthalene_assay.png|frame|center]]
| + | |
| - | [[File:UCalgary2012Week9GelTry2.png|500px|thumb|Alkane Hydroxylase system colony PCR gel (K398014). Lanes 2-8 (Colonies A-E) with a 1Kb ladder (L).|center]]<html>
| + | |
| | | | |
| - | <h2>Week 10 (July 2-July 6)</h2> | + | </html>[[File:Calgary CatechololeTGC.jpg|600px|thumb|centre|Figure 6: Gas chromatograph of catechol degradation assay using <i>Jeotgalicoccus</i> sp. ATCC 8456 a species of bacteria that converts fatty acids into alkenes. This identified a similar peak change in the PetroBrick with a retention time of 10.5 min as shown in Figure 4. This provides additional support that the PetroBrick and this organism can further degrade catechol into breakdown products.]]<html> |
| | | | |
| - | <h3>Ring Cleavage</h3> | + | </html>[[File:Calgary CatecholMSoleT.jpg|600px|thumb|centre|Figure 7: Mass spectrum of <i>Jeotgalicoccus</i> sp. ATCC 8456/<i>xylE</i> co-culture retention peak at 10.5 min as shown in Figure 6. This peak is similar to the peak from the Petrobrick/<i>xylE</i> co-culture, suggesting the breakdown product for both of these cultures is modified catechol from <i>xylE</i>. The identification of this compound is ongoing.]]<html> |
| | | | |
| - | <p> This week we performed another colony PCR of 5 colonies from the transformation plate of part K398014 and ran the products on a gel. We obtained 4 positive colonies | + | <p> Based on our GC-MS results, we were able to show the appearance of a new peak when cells expressing <i>xylE</i> and the PetroBrick were co-cultured. Although we don't know the exact identity of this peak, it is distinct form our control. It resembles standards that contain a toluene substituent suggesting that this compound maybe similar in structure. This structure is also made possible by the fact that the catechol degradation product is capable of forming polymers and potentially ring type structures. It may very well be that these ring structures are being further processed by the Petrobrick into useful products, which we hope to identify through GC-MS/MS in the future. Interestingly, a similar peak appeared when cells expressing our <i>xylE</i> construct were co-cultured with <i>Jeotgalicoccus</i> sp. ATCC 8456 cells. This suggests that although we don't know the exact identity of this new peak, it is likely that it may be in fact a further breakdown product of catechol. This is a very promising result, as it suggests that in addition to converting naphthenic acids into hydrocarbons, we may also be able to break down catechol, one of the other major toxic components in tailings ponds.</p> |
| - | and made overnight cultures. We decided to stop using this part for the project and therefore did not require a further miniprep on these overnight cultures. However we made an overnight culture of the positive colony from the alkane hydroxylase system gel (colony 5) and miniprepped the overnight culture. This resulted in a good concentration of 345.6 ng/μL as we continued verification of XylE by doing two more colony PCRs of each part (K118021 and J33204) but only the positive control worked when we ran the gel.
| + | |
| - | </p> | + | |
| | | | |
| - | </html>[[File:UCalgary2012_Week10XylEPCRgel.png|500px|thumb|Alkane Hydroxylase system colony PCR gel (K398014) Lanes 2-8 (Colonies A-E) with a 1Kb ladder (L).|center]]<html>
| |
| | | | |
| - | <h2>Week 11 (July 9-July 13)</h2>
| |
| | | | |
| - | <h3>Ring Cleavage</h3>
| |
| | | | |
| - | <p> Since we could not obtain any positive colonies from last weeks colony PCRs we began the transformation of each part again. The transformations were successful and when the colony PCR products were run on a gel there were two positive colonies for each part. Overnight cultures of the positive colonies were made. Minipreps were done on the overnight cultures and a restriction digest was performed on the products of the minipreps. The restriction digest products were run on a gel and there was one that looked like it had been successful but due to a poor quality ladder on this gel we could not make any final conclusions on the success of the restriction digest.</p>
| |
| | | | |
| - | </html>[[File:UCalgary2012_Week11GEl.png|500px|thumb|XylE Restriction Digest Gel (J33204: lanes 2,3 – colonies J2,J4) (K118021: lanes 4,5 – colonies: K3,K5) with a poor 1Kb ladder (L).|center]]<html> | + | </html>}} |
| - | | + | |
| - | <h2>Week 12 (July 16 -July 20)</h2>
| + | |
| - | | + | |
| - | <h3>Ring Cleavage</h3>
| + | |
| - | | + | |
| - | <p> We began the week by rerunning the restriction digest products on a gel with a good ladder and obtained one positive result with two bands. One band was around 1000 bp for the part and the other was at about 2000 bp for the plasmid backbone. We made a streak plate of the colony that had worked for the restriction digest and sent the miniprep product for sequencing. The colony that worked was from the transformation plate of part K118021. We also repeated a colony PCR of the transformants from part J33204 using 10 colonies and all of them looked like positive colonies. Overnight culutres were made and a miniprep and restriction digest was completed.</p>
| + | |
| - | | + | |
| - | </html>[[File:UCalgary2012_Week12Gel1.png|500px|thumb|XylE Restriction Digest Gel (J33204: lanes 6,7 – colonies J2,J4) (K118021: lanes 8-11 – colonies: K3,K5,2,3) with a 1Kb ladder (L).|center]]
| + | |
| - | [[File:UCalgary2012Week12Gel2.png|500px|thumb|Colony PCR gel of XylE with rbs site (J33204). Lanes 9-13 (Colonies A,B,D,J) with a 1Kb ladder (L).|center]]
| + | |
| - | [[File:UCalgary2012Week12(3).png|500px|thumb|Restriction digest gel of XylE gene with rbs site (J33204). Lanes 10-14 (Colonies A-J). Successfully digested plasmids from colonies B and D are at an expected size of 1000 bp with a 1Kb ladder (L).|center]] <html>
| + | |
| - | | + | |
| - | <h2>Week 13 (July 23 - July 27)</h2>
| + | |
| - | | + | |
| - | <h3>Catechol Degradation</h3>
| + | |
| - | | + | |
| - | <p>The colony PCR of part J33204 (XylE with a native rbs site) was done again and all five colonies looked like they contained the plasmid after running the products on a gel. After miniprepping and running a restriction digest on these five colonies two of them were sent to sequencing.
| + | |
| - | An assay using the compound catechol was also started this week. E.coli cells (top 10) transformed with the part K118021 (XylE with a Pcst promoter) were used in this assay. Six overnight cultures were made, 3 using M9-MM and 3 using LB. These were spun down and resuspended in fresh M9-MM and LB and brought to a concentration of 0.1 M catechol by using a 1M stock solution. There seemed to be an initial colour change in the reactions from clear to yellow which was only visible in the cultures using M9-MM. All of the cultures were incubated overnight and they turned a black green colour. For this assay we should have used the supernatant because the reaction takes place outside of the cell so this is what we did next week.</p>
| + | |
| - | | + | |
| - | <h2>Week 14 (July 30 - August 3)</h2>
| + | |
| - | | + | |
| - | <h3>Catechol Degradation</h3>
| + | |
| - | | + | |
| - | <p>The sequencing results for the part J33204 matched the part so a construction was started. The construction was a 3-way ligation putting the tetR promoter (R0040) and XylE (J33204) into a pSB1C3 vector. The ligation product was transformed into competent cells and a colony PCR was done on the transformants using both bio brick and R0040 primers.
| + | |
| - | The primers for XylT, which is a ferredoxin that allows for XylE to be activated after a reaction with catechol, arrived and a colony PCR on four different strains of Pseudomonas putida was done. None of the strains seemed to contain the TOL plasmid, which is where XylT is found in certain strains of P. putida, because no bands showed up in the gel.
| + | |
| - | The catechol assay was continued this week. Six overnight cultures were made, 3 using M9-MM and 3 using LB. They were spun down and the supernatant was brought to a catechol concentration of 0.1M, 0.2 M and 0.5 M using a 1M stock solution. The solutions turned light yellow initially, but after a few hours they turned brown.
| + | |
| - | The assay was repeated with the same procedure as above but a control, a colony that did not contain the part K118021, was added. After the catechol was added to the supernatants there was a colour change from clear to yellow in all of the tubes, even the control. For a negative control catechol was added to M9-MM and this solution became pink. The tubes were incubated on the bench and checked regularly. The colour changed from yellow to pink to brown when they were left on the bench overnight.</p>
| + | |
| - | | + | |
| - | <h2>Week 15 (August 7 - August 10)</h2>
| + | |
| - | | + | |
| - | | + | |
| - | | + | |
| - | | + | |
| - | }} | + | |
Hello! iGEM Calgary's wiki functions best with Javascript enabled, especially for mobile devices. We recommend that you enable Javascript on your device for the best wiki-viewing experience. Thanks!
Decatecholization

Catechol is a toxic compound found in tailings ponds that is a by-product of polyaromatic hydrocarbon metabolism (Vaillancourt et al., 2006, Schweigert et al., 2001)). The chemical properties of catechol allow it to react with biomolecules, causing cellular damage including DNA damage, enzyme inactivation and membrane uncoupling (Schweigert et al., 2001).
Catechol is characterized as having a benzene ring with two hydroxyl groups at the 2,3 position. It can be converted to 2-hydroxymuconic acid by the enzyme catechol 2,3-dioxygenase, encoded by the xylE gene on the Tol plasmid of Pseudomonas putida (Nakai et al., 1983).
Currently the registry has two BioBricks available of xylE. One contained xylE with its native ribosome-binding site (BBa_J33204), while the other part contained xylE under the glucose-repressible promoter cstA (BBa_K118021). Given that E. coli is grown in the presence of glucose, we designed a new construct to keep xylE expressed by using the tetR promoter (BBa_R0040).
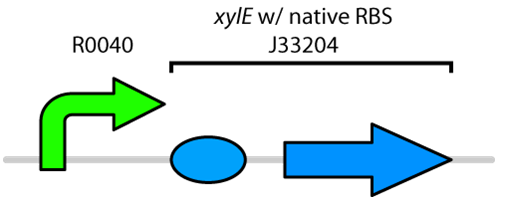
Figure 1: BioBrick genetic circuit for catechol degradation showing
xylE under the
tetR promoter
Catechol 2,3-dioxygenase is an extradiol dioxygenase which cleaves catechol adjacent to the two hydroxyl groups. When this occurs 2-hydroxymuconate semialdehyde is produced, which is yellow in colour. This change in colour allows for visual assay to assess the activity of XylE.
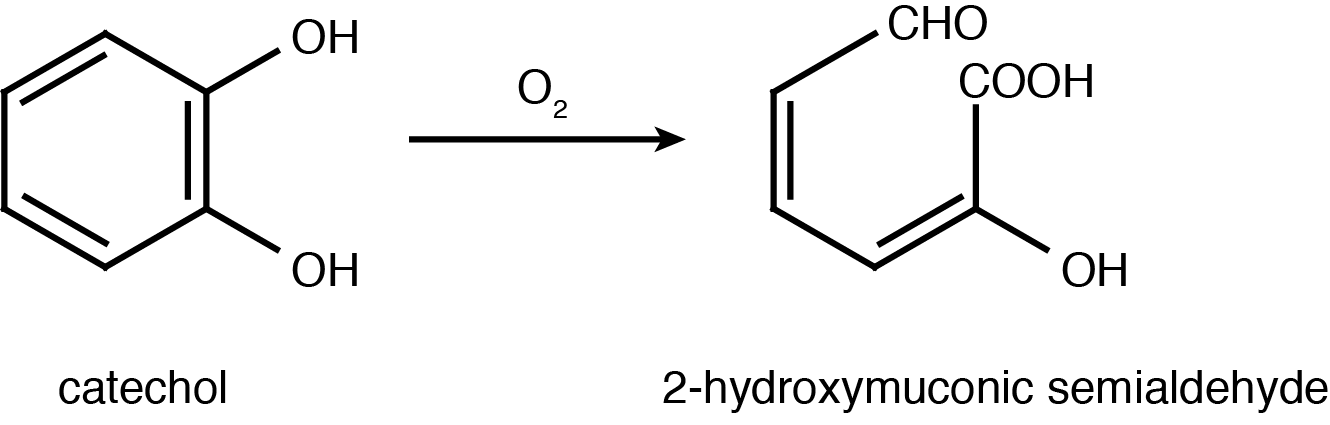
Figure 2: Catechol 2,3-dioxygenase (XylE) converts catechol to 2-hydroxymuconate semialdehyde in the presence of oxygen. Adapted from Shu
et al., 1995.
The visual assays were performed with E. coli cells transformed with (BBa_K118021) as well as with E. coli cells transformed with the newly constructed part (BBa_K902048) by bringing the supernatant of an overnight culture to a concentration of 0.1 M of catechol. When the part (BBa_K118021) was used, the pellet was first washed in M9-MM and centrifuged before catechol was added to the supernatant. This was necessary to avoid the glucose in the LB from repressing the cstA promoter (BBa_K118011). Catechol was added to the supernatant because the reaction takes place outside of the cell. Within minutes of the addition of catechol to the supernatant, the solution turned from the pale yellow of LB to a bright yellow. This was indicative that catechol was breaking down into 2-hydroxymuconate semialdehyde, which was exactly what we expected! This assay was completed by following the protocol written by the 2008 Edinburgh iGEM team.
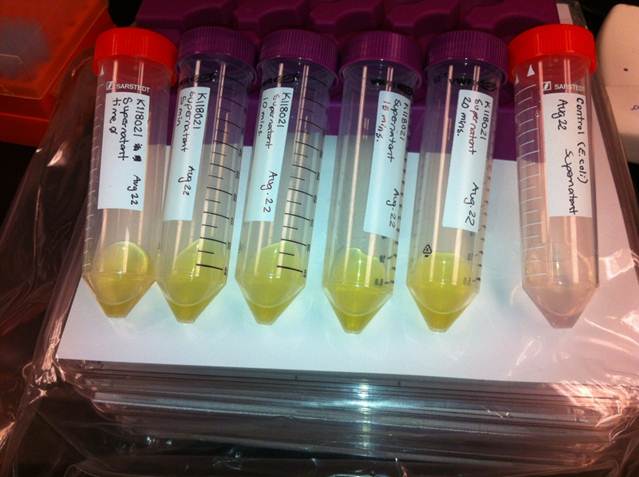
Figure 3: Results of the catechol visual assay using
xylE [http://partsregistry.org/Part:BBa_K118021 BBa_K118021]. Cultures were grown overnight in LB and the pellets were washed with M9-MM at various times (From left to right: 0 min, 5 min, 10 min, 15 min, and 20 min.). Cells were then spun down and catechol was added to the supernatant to 0.1 M. The amount of time didn't affect the colour change in the cultures containing the
xylE gene. The far right tube has
E. coli cells without the
xylE gene as a negative control and the supernatant remained clear when the catechol was added.
Converting Catechol into hydrocarbons?
After verifying that we could in fact degrade catechol into 2-hydroxymuconate semialdehyde using our xylE construct (BBa_J33204), we wondered if we could take this any further. What if we could convert this by-product into hydrocarbons too? As catechol is the breakdown product of a number of different degradation pathways in bacteria, this could be particularly useful.
Since 2-hydroxymuconate semialdehyde can be further metabolized to pyruvate and acetaldehyde (Harayama et al., 1987), it seemed possible that these products could be routed into the fatty acid biosynthesis pathway and converted to alkanes using the PetroBrick or the OleT enzyme. Given that the catechol 2,3-dioxygenase reaction is extracellular, it creates a possible scenario in which cells with the xylE construct could be co-cultured with Petrobrick-containing cells to cooperatively metabolize catechol into hydrocarbons.
In order to test this, we followed this protocol, where we co-cultured cells expressing our xylE construct with either E. coli cells expressing the PetroBrick, or Jeotgalicoccus sp. ATCC 8456 cells expressing oleT in the presence of catechol.
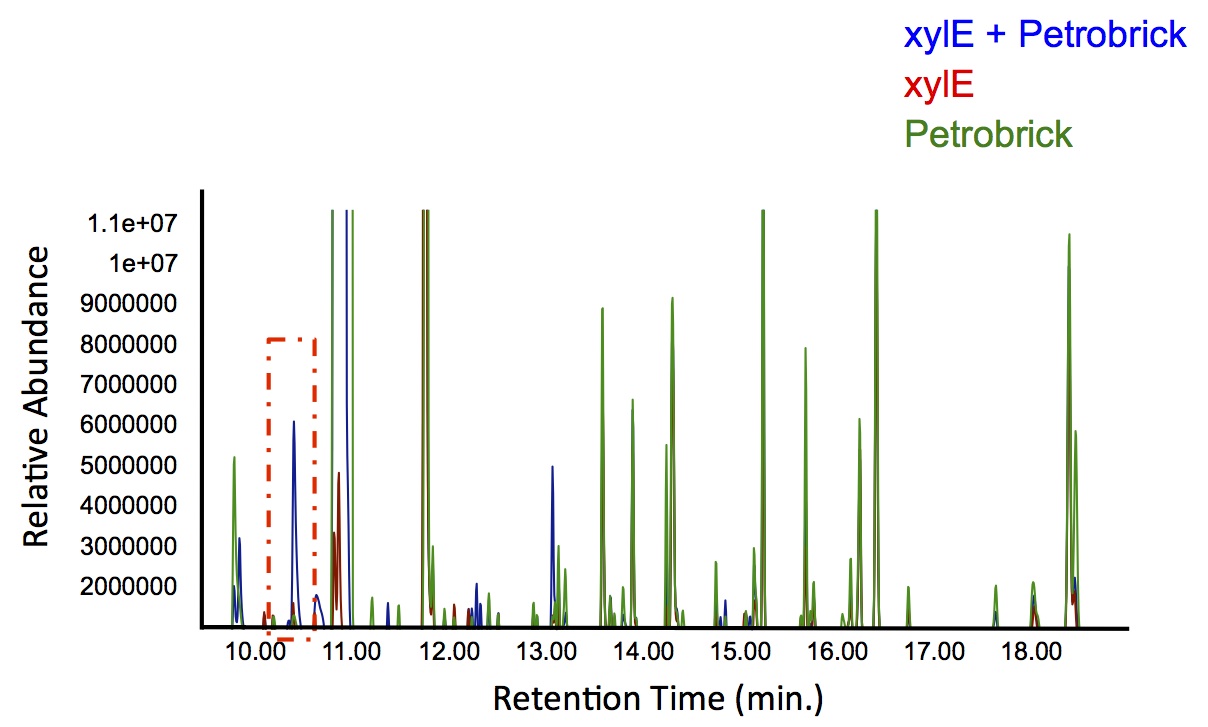
Figure 4: Gas chromatograph of catechol degradation assay using the PetroBrick. While there is limited differences between
xylE incubated with and without the PetroBrick, there was one peak with a retention time of 10.5 min which was dramatically increased in the co-culture.
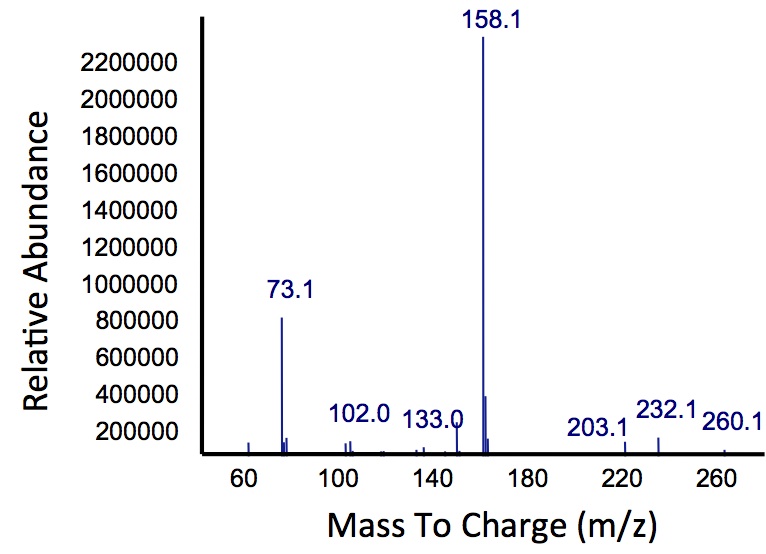
Figure 5: Mass spectrum of the Petrobrick/
xylE co-culture retention peak at 10.5 min as shown in Figure 4. While the identity of this compound is currently unknown, there are changes occuring to some of the catechol breakdown products.
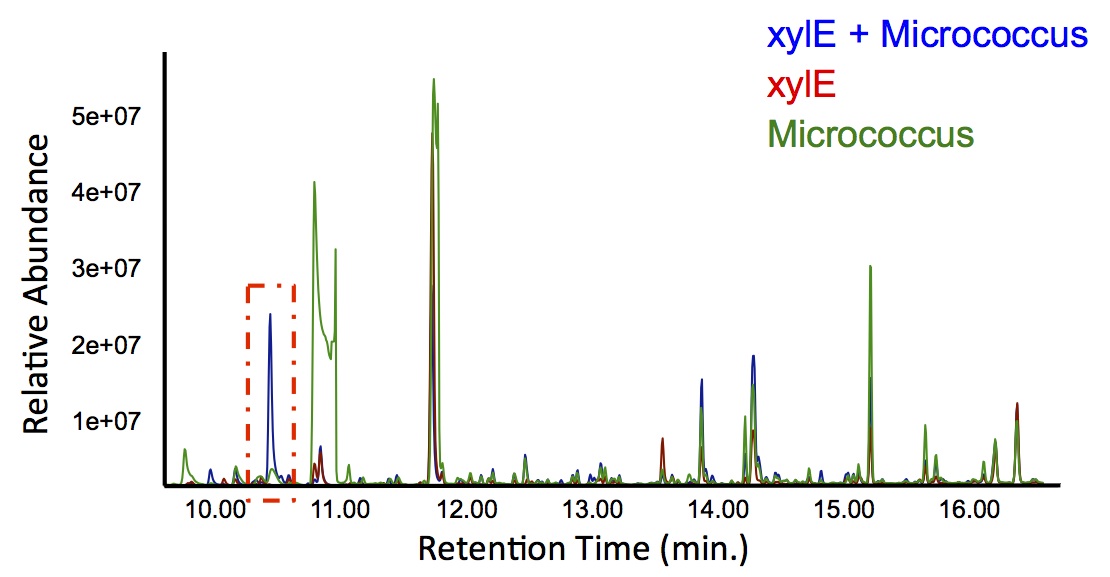
Figure 6: Gas chromatograph of catechol degradation assay using
Jeotgalicoccus sp. ATCC 8456 a species of bacteria that converts fatty acids into alkenes. This identified a similar peak change in the PetroBrick with a retention time of 10.5 min as shown in Figure 4. This provides additional support that the PetroBrick and this organism can further degrade catechol into breakdown products.
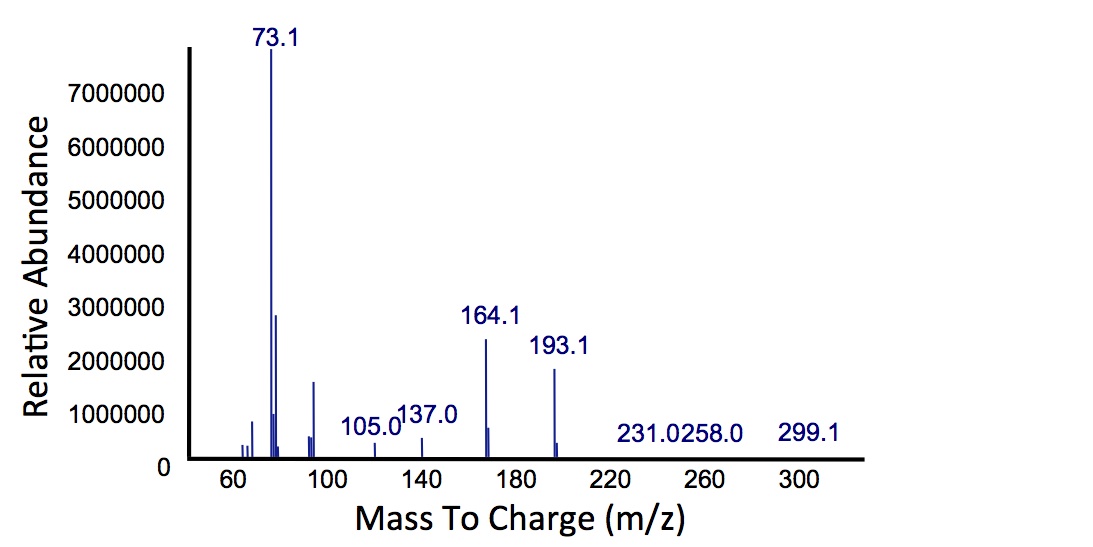
Figure 7: Mass spectrum of
Jeotgalicoccus sp. ATCC 8456/
xylE co-culture retention peak at 10.5 min as shown in Figure 6. This peak is similar to the peak from the Petrobrick/
xylE co-culture, suggesting the breakdown product for both of these cultures is modified catechol from
xylE. The identification of this compound is ongoing.
Based on our GC-MS results, we were able to show the appearance of a new peak when cells expressing xylE and the PetroBrick were co-cultured. Although we don't know the exact identity of this peak, it is distinct form our control. It resembles standards that contain a toluene substituent suggesting that this compound maybe similar in structure. This structure is also made possible by the fact that the catechol degradation product is capable of forming polymers and potentially ring type structures. It may very well be that these ring structures are being further processed by the Petrobrick into useful products, which we hope to identify through GC-MS/MS in the future. Interestingly, a similar peak appeared when cells expressing our xylE construct were co-cultured with Jeotgalicoccus sp. ATCC 8456 cells. This suggests that although we don't know the exact identity of this new peak, it is likely that it may be in fact a further breakdown product of catechol. This is a very promising result, as it suggests that in addition to converting naphthenic acids into hydrocarbons, we may also be able to break down catechol, one of the other major toxic components in tailings ponds.






 "
"



