Team:Calgary/Project/OSCAR/Denitrogenation
From 2012.igem.org
| Line 12: | Line 12: | ||
<p>Carbazole was chosen as the model compound to study nitrogen containing heterocycles as it accounts for about 70% of nitrogen by mass in petroleum tailings ponds, as well as the fact that it is considered one of the more difficult to degrade (Morales & Le Borgne, 2010). A pathway that is capable of degrading carbazole could also have the ability to degrade many other types of nitrogen containing compounds found in the tailings ponds.</p> | <p>Carbazole was chosen as the model compound to study nitrogen containing heterocycles as it accounts for about 70% of nitrogen by mass in petroleum tailings ponds, as well as the fact that it is considered one of the more difficult to degrade (Morales & Le Borgne, 2010). A pathway that is capable of degrading carbazole could also have the ability to degrade many other types of nitrogen containing compounds found in the tailings ponds.</p> | ||
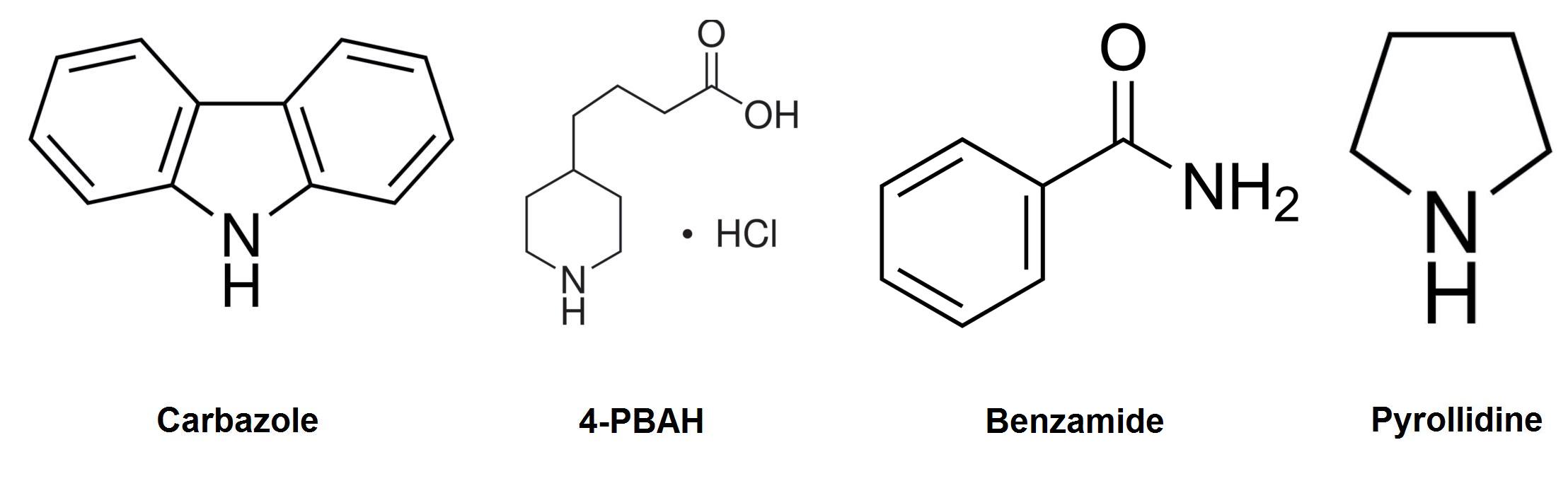
| - | </html>[[File:Model compounds denitrogenation calgary12.jpg|thumb|420px|centre|Model compounds used to test the ability of our system to remove nitrogen containing compounds.]]<html> | + | </html>[[File:Model compounds denitrogenation calgary12.jpg|thumb|420px|centre|Figure 1: Model compounds used to test the ability of our system to remove nitrogen containing compounds.]]<html> |
| Line 19: | Line 19: | ||
<p>We identified a <i>Pseudomonas</i> genus that is capable of converting carbazole into cathecol with the hope that it would also be able to act on similar nitrogen heterocycles. This is accomplished thorugh an enzymatic pathway containing carbazole-1,9-dioxygenase (<i>carA</i>) and an enzyme called anthranilate 1,2-dioxygenase. <i>carA</i> is a 4 part enzyme encoded by three genes: carAa, carAc and carAd. It selectively cleaves the first C-N bond in a nitrogen containing ring structure, converting carbazole into 2'-aminobiphenyl-2,3-diol (Xu et al, 2006,Morales & Le Borgne, 2010). After this step, anthranilate 1,2-dioxygenase enzyme, encoded by the <i>ant</i> operon removes nitrogen from the structure (Diaz & Garcia, 2010). </p>. | <p>We identified a <i>Pseudomonas</i> genus that is capable of converting carbazole into cathecol with the hope that it would also be able to act on similar nitrogen heterocycles. This is accomplished thorugh an enzymatic pathway containing carbazole-1,9-dioxygenase (<i>carA</i>) and an enzyme called anthranilate 1,2-dioxygenase. <i>carA</i> is a 4 part enzyme encoded by three genes: carAa, carAc and carAd. It selectively cleaves the first C-N bond in a nitrogen containing ring structure, converting carbazole into 2'-aminobiphenyl-2,3-diol (Xu et al, 2006,Morales & Le Borgne, 2010). After this step, anthranilate 1,2-dioxygenase enzyme, encoded by the <i>ant</i> operon removes nitrogen from the structure (Diaz & Garcia, 2010). </p>. | ||
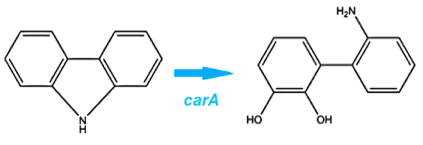
| - | </html>[[File:CarA pathway Calgary12.jpg|thumb|420px|center|Action of the <i>carA</i> enzyme complex.]]<html> | + | </html>[[File:CarA pathway Calgary12.jpg|thumb|420px|center|Figure 2: Action of the <i>carA</i> enzyme complex.]]<html> |
<h2> Does it work? </h2> | <h2> Does it work? </h2> | ||
| Line 25: | Line 25: | ||
<p>Upon obtaining the carbazole-degrading pseudomonas strain, we wanted to validate that this approach was valid. We performed an initial carbazole degradation assay where we incubated a <i>pseudomonas</i> culture with carbazole and monitored degradation over time. Our detailed protocol can be found <a href="https://2012.igem.org/Team:Calgary/Notebook/Protocols/carbazole">here.</a></p> | <p>Upon obtaining the carbazole-degrading pseudomonas strain, we wanted to validate that this approach was valid. We performed an initial carbazole degradation assay where we incubated a <i>pseudomonas</i> culture with carbazole and monitored degradation over time. Our detailed protocol can be found <a href="https://2012.igem.org/Team:Calgary/Notebook/Protocols/carbazole">here.</a></p> | ||
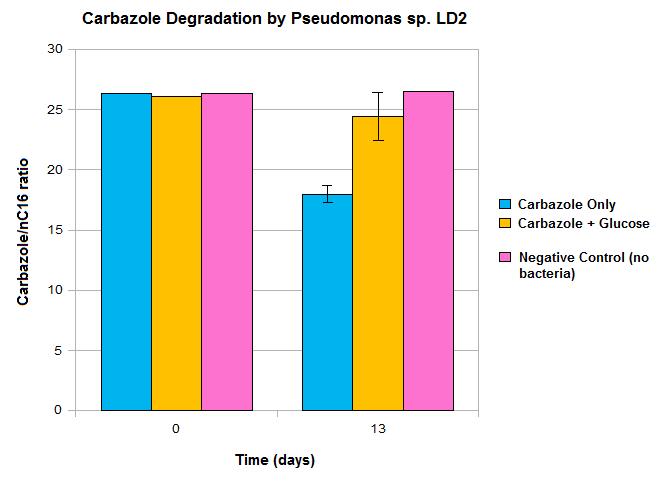
| - | </html>[[File:LD2 degradation september data calgary12.jpg|thumb|420px|right|Carbazole loss demonstrated after culturing with <i>Pseudomonas sp. LD2</i>, the source of the <i>car</i> and <i>ant</i> genes.]]<html> | + | </html>[[File:LD2 degradation september data calgary12.jpg|thumb|420px|right|Figure 3: Carbazole loss demonstrated after culturing with <i>Pseudomonas sp. LD2</i>, the source of the <i>car</i> and <i>ant</i> genes.]]<html> |
<p>Our data suggests that over time, we do see a decrease in carbazole levels, shown by the difference seen between day 0 and day 13. However, these results also show some limitations in the native pathway that we hope our synthetic approach will be able to overcome. Firstly, we only see modest loss of carbazole after 13 days, which would not be quick enough for OSCAR's large-scale bioreactor. Secondly, the system must be glucose free for any significant carbazole loss to occur, as we didn’t see a significant decrease in carbazole levels when glucose was also present in the system. This also may not be realistic in our bioreactor. This may be due to the P<sup>ant</sup> promoter, the native promoter upstream of the any genes becoming deactivated in the presence of a preferred fuel source such as glucose. As such, we wanted to introduce an alternative promoter. This promoter may also gave the potential to upregulate transcription levels, making the degradation process quicker.</p> | <p>Our data suggests that over time, we do see a decrease in carbazole levels, shown by the difference seen between day 0 and day 13. However, these results also show some limitations in the native pathway that we hope our synthetic approach will be able to overcome. Firstly, we only see modest loss of carbazole after 13 days, which would not be quick enough for OSCAR's large-scale bioreactor. Secondly, the system must be glucose free for any significant carbazole loss to occur, as we didn’t see a significant decrease in carbazole levels when glucose was also present in the system. This also may not be realistic in our bioreactor. This may be due to the P<sup>ant</sup> promoter, the native promoter upstream of the any genes becoming deactivated in the presence of a preferred fuel source such as glucose. As such, we wanted to introduce an alternative promoter. This promoter may also gave the potential to upregulate transcription levels, making the degradation process quicker.</p> | ||
| Line 34: | Line 34: | ||
<p> As we want our system to act on a variety of substrates, we were also concerned about the specificity of the anthranilate 1,2-dioxygenase enzyme. It is designed to act on the substrate anthranilate, which is produced in the native pathway by the actions of the <i>carB</i> and <i>carC</i> enzymes, however since our synthetic pathway does not include these genes as they do not directly attack nitrogen there is a chance that this enzyme will not interact with our remaining nitrogen groups. To this end, we also wanted to explore a deaminase enzyme (<i>amdA</i>) from <i>Rhodococcus erythropolis</i> that has been shown to selectively cleave the 2nd C-N bond from a variety of nitrogen heterocycles (Kilbane et al., 2002, Kayser & Kilbane, 2005). This allows an alternative pathway for the 2nd step of nitrogen removal, possibly making the system more efficient and should show less substrate specificity than the <i>ant</i> genes allowing us to degrade a wider range of nitrogen containing rings.</p> | <p> As we want our system to act on a variety of substrates, we were also concerned about the specificity of the anthranilate 1,2-dioxygenase enzyme. It is designed to act on the substrate anthranilate, which is produced in the native pathway by the actions of the <i>carB</i> and <i>carC</i> enzymes, however since our synthetic pathway does not include these genes as they do not directly attack nitrogen there is a chance that this enzyme will not interact with our remaining nitrogen groups. To this end, we also wanted to explore a deaminase enzyme (<i>amdA</i>) from <i>Rhodococcus erythropolis</i> that has been shown to selectively cleave the 2nd C-N bond from a variety of nitrogen heterocycles (Kilbane et al., 2002, Kayser & Kilbane, 2005). This allows an alternative pathway for the 2nd step of nitrogen removal, possibly making the system more efficient and should show less substrate specificity than the <i>ant</i> genes allowing us to degrade a wider range of nitrogen containing rings.</p> | ||
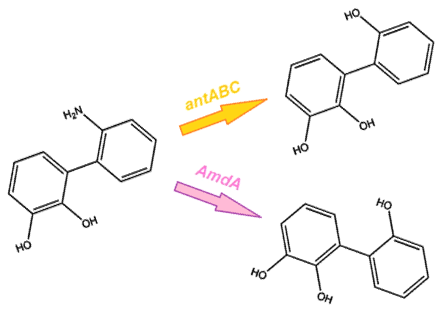
| - | </html>[[File:Carbazole degradation lower pathway calgary12.png|thumb|420px|centre|2 options for the final step in the pathway.]]<html> | + | </html>[[File:Carbazole degradation lower pathway calgary12.png|thumb|420px|centre|Figure 4: 2 options for the final step in the pathway.]]<html> |
<h2> Putting it all together- Where are we at? </h2> | <h2> Putting it all together- Where are we at? </h2> | ||
| Line 40: | Line 40: | ||
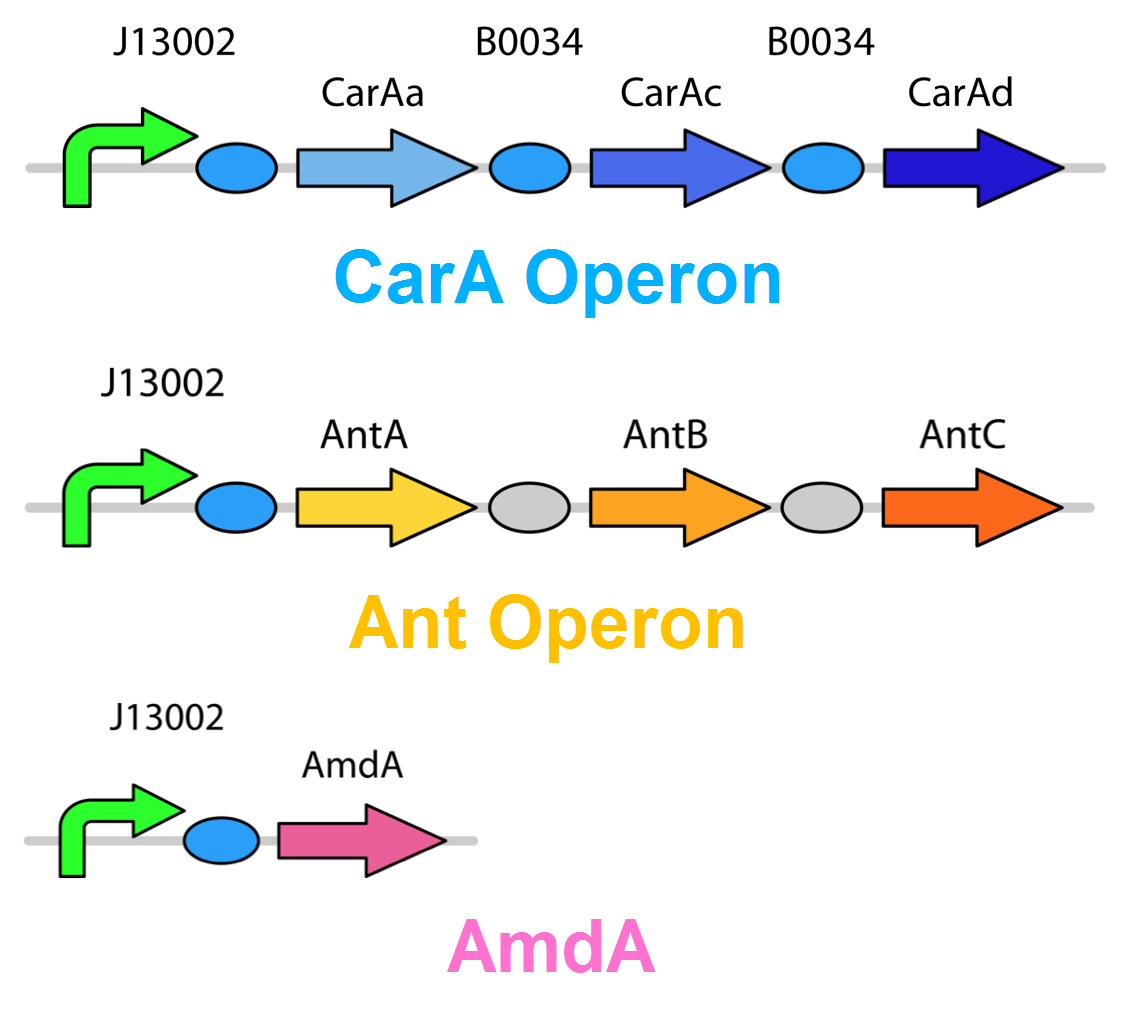
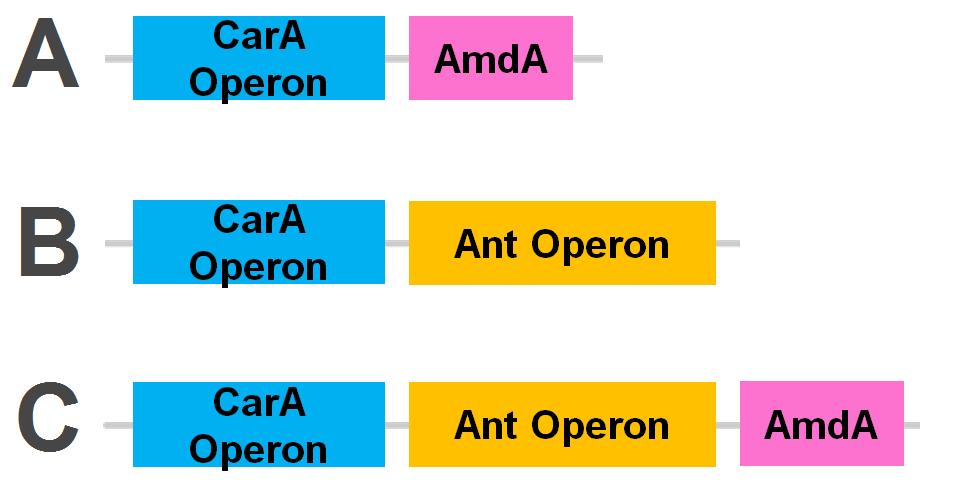
<p>The <i>carA</i> genes have all been biobricked individually (<a href="http://partsregistry.org/Part:BBa_K902032">BBa_K902032</a>,<a href="http://partsregistry.org/Part:BBa_K902033">BBa_K902033</a> and <a href="http://partsregistry.org/Part:BBa_K902034">BBa_K902034</a>) and the <i>carAd</i> gene was given a silent mutation to eliminate an illegal NotI site in the native sequence via <a href="https://2012.igem.org/Team:Calgary/Notebook/Protocols/mutagenesis">site directed mutagenesis</a>. We are now in the process of adding ribosome binding sites (<a href="http://partsregistry.org/Part:BBa_B0034">BBa_B0034</a>) and promoter elements (<a href="http://partsregistry.org/Part:BBa_J13002">BBa_J13002</a>). The <i>ant</i> genes were biobricked as a whole operon with native ribosome binding sites in between each gene and have been constructed downstream of a TteR repressible promoter and ribosomal binding site (<a href="http://partsregistry.org/Part:BBa_J13002">BBa_J13002</a>). The <a href="http://partsregistry.org/Part:BBa_K902035"><i>AmdA</i></a> gene was also biobricked and constructed downstream of a TetR repressible promoter and ribosomal binding site (<a href="http://partsregistry.org/Part:BBa_J13002">BBa_J13002</a>). We are now almost done constructing these into lager test constructs allowing us to compare the <i>ant</i> pathway vrsus the <i>amdA</i> pathway. Final constructs are illustrated below. </p> | <p>The <i>carA</i> genes have all been biobricked individually (<a href="http://partsregistry.org/Part:BBa_K902032">BBa_K902032</a>,<a href="http://partsregistry.org/Part:BBa_K902033">BBa_K902033</a> and <a href="http://partsregistry.org/Part:BBa_K902034">BBa_K902034</a>) and the <i>carAd</i> gene was given a silent mutation to eliminate an illegal NotI site in the native sequence via <a href="https://2012.igem.org/Team:Calgary/Notebook/Protocols/mutagenesis">site directed mutagenesis</a>. We are now in the process of adding ribosome binding sites (<a href="http://partsregistry.org/Part:BBa_B0034">BBa_B0034</a>) and promoter elements (<a href="http://partsregistry.org/Part:BBa_J13002">BBa_J13002</a>). The <i>ant</i> genes were biobricked as a whole operon with native ribosome binding sites in between each gene and have been constructed downstream of a TteR repressible promoter and ribosomal binding site (<a href="http://partsregistry.org/Part:BBa_J13002">BBa_J13002</a>). The <a href="http://partsregistry.org/Part:BBa_K902035"><i>AmdA</i></a> gene was also biobricked and constructed downstream of a TetR repressible promoter and ribosomal binding site (<a href="http://partsregistry.org/Part:BBa_J13002">BBa_J13002</a>). We are now almost done constructing these into lager test constructs allowing us to compare the <i>ant</i> pathway vrsus the <i>amdA</i> pathway. Final constructs are illustrated below. </p> | ||
| - | </html>[[File:Carbazole degradation constructs calgary12.png|thumb|320px|left|Genetic circuits for nitrogen removal. 2 operons and an additional gene have been biobricked together under 1 TetR promoter each.]]<html></html>[[File:Carbazole Super constructs calgary12.jpg|thumb|320px|right|3 "superconstructs" to be tested for their nitrogen removal ability.]]<html> | + | </html>[[File:Carbazole degradation constructs calgary12.png|thumb|320px|left|Figure 5: Genetic circuits for nitrogen removal. 2 operons and an additional gene have been biobricked together under 1 TetR promoter each.]]<html></html>[[File:Carbazole Super constructs calgary12.jpg|thumb|320px|right|3 "superconstructs" to be tested for their nitrogen removal ability.]]<html> |
Revision as of 11:02, 3 October 2012


Hello! iGEM Calgary's wiki functions best with Javascript enabled, especially for mobile devices. We recommend that you enable Javascript on your device for the best wiki-viewing experience. Thanks!
Denitrogenation

Why Focus On Nitrogen Bioremediation?
The removal of nitrogen heterocycles from the petroleum tailings ponds is important for both of the main goals of the OSCAR project. Since nitrogen containing compounds lower the quality and stability of fuel (Katzer & Sivasubramanian, 1979) , their presence would be detrimental to the usefulness of the hydrocarbons OSCAR could create. The presence of nitrogen compounds in our products also hinders our environmental goals as they have been shown to undergo radical changes, yielding highly genotoxic byproducts (Xu et al., 2006). The combination of these two factors makes the removal of nitrogen very important to the success of OSCAR.
Our Model Compounds
Carbazole was chosen as the model compound to study nitrogen containing heterocycles as it accounts for about 70% of nitrogen by mass in petroleum tailings ponds, as well as the fact that it is considered one of the more difficult to degrade (Morales & Le Borgne, 2010). A pathway that is capable of degrading carbazole could also have the ability to degrade many other types of nitrogen containing compounds found in the tailings ponds.
How Are We Going To Accomplish This?
We identified a Pseudomonas genus that is capable of converting carbazole into cathecol with the hope that it would also be able to act on similar nitrogen heterocycles. This is accomplished thorugh an enzymatic pathway containing carbazole-1,9-dioxygenase (carA) and an enzyme called anthranilate 1,2-dioxygenase. carA is a 4 part enzyme encoded by three genes: carAa, carAc and carAd. It selectively cleaves the first C-N bond in a nitrogen containing ring structure, converting carbazole into 2'-aminobiphenyl-2,3-diol (Xu et al, 2006,Morales & Le Borgne, 2010). After this step, anthranilate 1,2-dioxygenase enzyme, encoded by the ant operon removes nitrogen from the structure (Diaz & Garcia, 2010).
.Does it work?
Upon obtaining the carbazole-degrading pseudomonas strain, we wanted to validate that this approach was valid. We performed an initial carbazole degradation assay where we incubated a pseudomonas culture with carbazole and monitored degradation over time. Our detailed protocol can be found here.
Our data suggests that over time, we do see a decrease in carbazole levels, shown by the difference seen between day 0 and day 13. However, these results also show some limitations in the native pathway that we hope our synthetic approach will be able to overcome. Firstly, we only see modest loss of carbazole after 13 days, which would not be quick enough for OSCAR's large-scale bioreactor. Secondly, the system must be glucose free for any significant carbazole loss to occur, as we didn’t see a significant decrease in carbazole levels when glucose was also present in the system. This also may not be realistic in our bioreactor. This may be due to the Pant promoter, the native promoter upstream of the any genes becoming deactivated in the presence of a preferred fuel source such as glucose. As such, we wanted to introduce an alternative promoter. This promoter may also gave the potential to upregulate transcription levels, making the degradation process quicker.
An alternative?
As we want our system to act on a variety of substrates, we were also concerned about the specificity of the anthranilate 1,2-dioxygenase enzyme. It is designed to act on the substrate anthranilate, which is produced in the native pathway by the actions of the carB and carC enzymes, however since our synthetic pathway does not include these genes as they do not directly attack nitrogen there is a chance that this enzyme will not interact with our remaining nitrogen groups. To this end, we also wanted to explore a deaminase enzyme (amdA) from Rhodococcus erythropolis that has been shown to selectively cleave the 2nd C-N bond from a variety of nitrogen heterocycles (Kilbane et al., 2002, Kayser & Kilbane, 2005). This allows an alternative pathway for the 2nd step of nitrogen removal, possibly making the system more efficient and should show less substrate specificity than the ant genes allowing us to degrade a wider range of nitrogen containing rings.
Putting it all together- Where are we at?
The carA genes have all been biobricked individually (BBa_K902032,BBa_K902033 and BBa_K902034) and the carAd gene was given a silent mutation to eliminate an illegal NotI site in the native sequence via site directed mutagenesis. We are now in the process of adding ribosome binding sites (BBa_B0034) and promoter elements (BBa_J13002). The ant genes were biobricked as a whole operon with native ribosome binding sites in between each gene and have been constructed downstream of a TteR repressible promoter and ribosomal binding site (BBa_J13002). The AmdA gene was also biobricked and constructed downstream of a TetR repressible promoter and ribosomal binding site (BBa_J13002). We are now almost done constructing these into lager test constructs allowing us to compare the ant pathway vrsus the amdA pathway. Final constructs are illustrated below.
 "
"