Team:UNAM Genomics Mexico/Modeling/Parameters
From 2012.igem.org
| Line 49: | Line 49: | ||
</tr> | </tr> | ||
<tr> | <tr> | ||
| - | <td id="contentcolumnwhite" align= "center"><br />[[File:UGM_Par5.png|870px]]<br /><br /><p> | + | <td id="contentcolumnwhite" align= "center"><br />[[File:UGM_Par5.png|870px]]<br /><br /><p>'''NOTE: mol refers to molecules, not the unit mol''' |
</p></td> | </p></td> | ||
</tr> | </tr> | ||
Latest revision as of 02:28, 27 October 2012

Parameters and considerations
Our team made both a deterministic and a stochastic model. There is a strong emphasis on exploring the dynamics given by the system's structure and the individual component's parameters in either type of model. The parameters were obtained from articles and databases such as Harvard Medical School's BioNumbers. Another important source of information was the general binding rates included in the Kappa Simulator manual.
General assumptions:
●This model's scope is single cell. While the regulation steps were modeled through Hill equations, other reactions were treated under Michaelis-Menten assumptions, notably isomerizations and transport processes (i.e. those that involved energy consumption).
●The maximal transcription rate for the system species was obtained by dividing the maximum number of nucleotides processed by the RNA polymerase (i.e. maximum polymerase activity), by the length of the nucleotide sequence.
●Translational rates were obtained by getting the maximum number of amino acids added by the ribosome divided by the length of the protein in amino acids.
●The protein degradation rate is considered equal for all the protein species in the model.
●The degradation rate for mRNAs is equal for all the different species of mRNAs.
●The different binding sites of the same transcription factor have the same affinity to the TF.
●For each mRNA species, its concentration will be the sum of the production as affected by its respective promoter and the transcription factors that regulate it, minus the degradation of the mRNA at that time.
●For each protein, the change in its concentration depends in the amount of protein produced by translation minus the degradation rate of the protein.
●The Hill coefficients tend to be 2.
●The parameters of concentration were expressed in terms of molecules per cell (volume of a B.subtilis cell ~10^-14) and the time units were converted sec.
Parameters
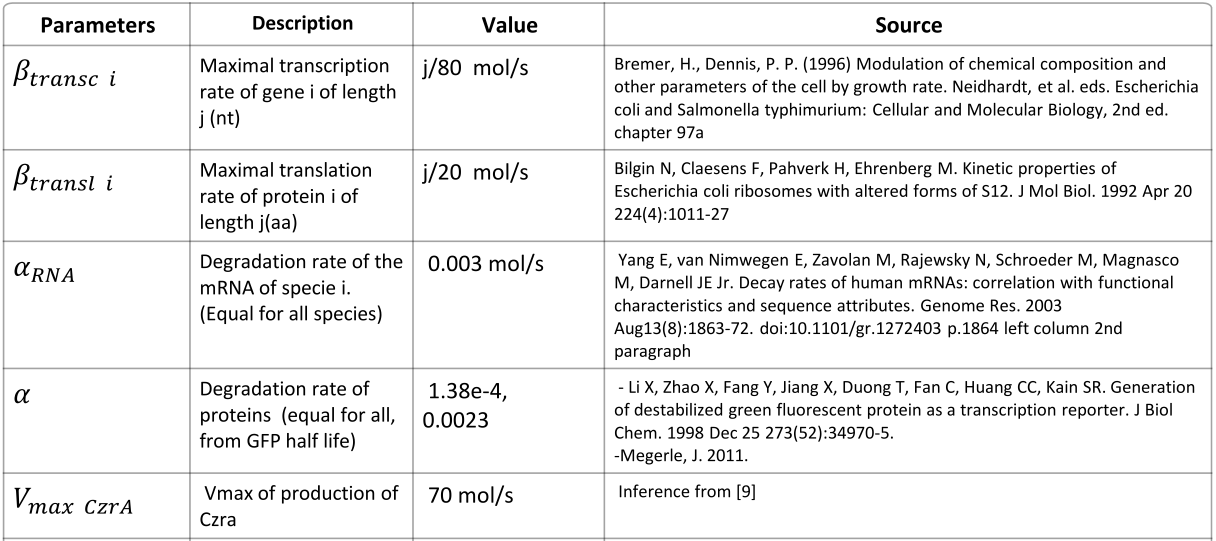
|
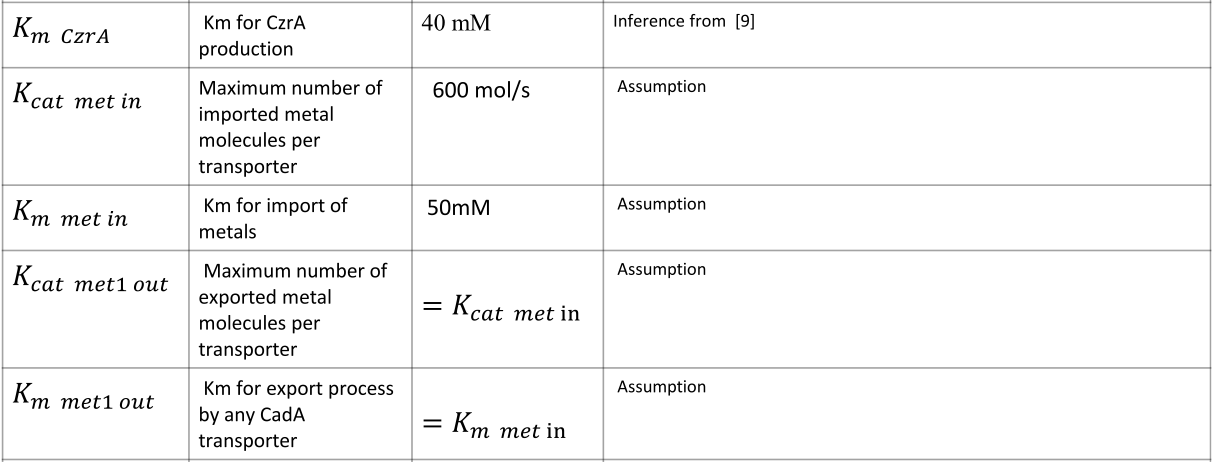
|
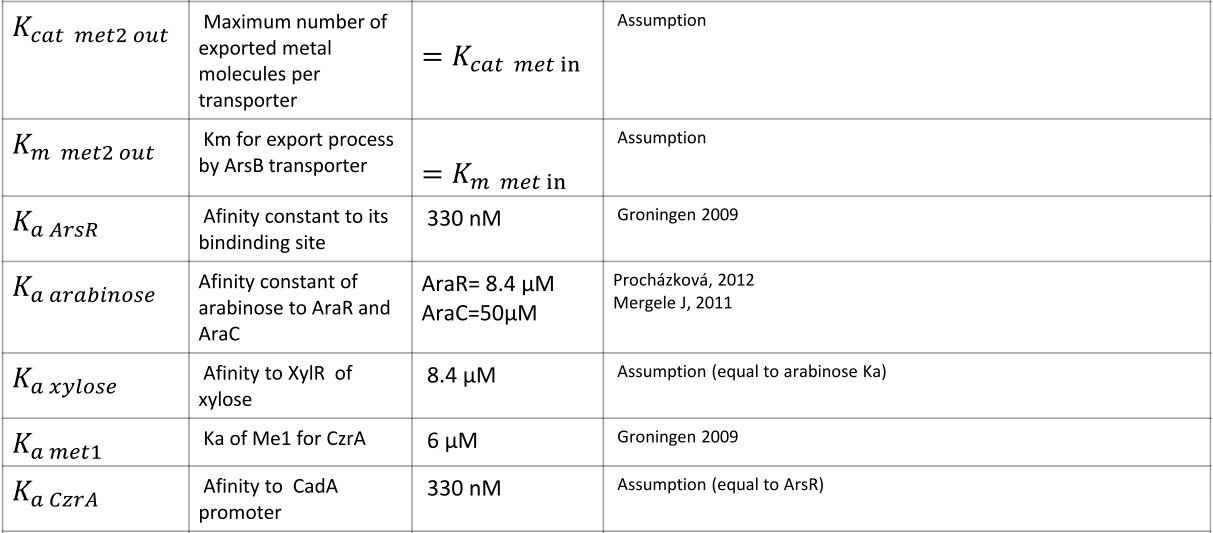
|
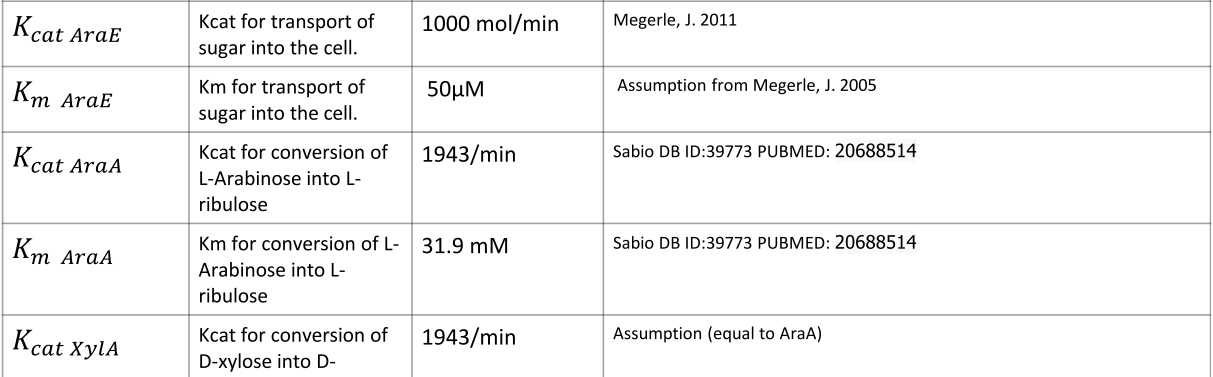
|
 NOTE: mol refers to molecules, not the unit mol |
References
[1]Rust, L.; E.C. Pcsi, B.H. Iglewski. Analysis of the Pseudomonas aeruginosa elastase (lasB) regulatory region.J. Bacteriol. February 1996 vol. 178 no. 4 1134-1140.
[2]Lyons T, David J E. Transport and storage of metal ions in Biology. Chapter V. "http://www.ffame.org/pubs/Transport%20and%20Storage%20of%20Metal%20Ions%20in%20Biology.pdf ".
[3]Fujita M, Tanaka K, Takahashi H, Amemura A. Transciption of hte principal sigma-factor genes, rpoD and rpoS, in Pseudomonas aeruginosa is controlled according to the growth phase. Mol Microbiology. 1994 Sep;13 (6):1071-7
[4]Laval Université. Chapitre 2 Régulation transcriptionnelle chez Pseudomonas aeruginosa. Collection Mémoires et théses électroniques. "http://archimede.bibl.ulaval.ca/archimede/fichiers/24237/ch02.html ".
[5]Kreuzer P, Gartner D, Allmansberger R, Hillen W. Identification and sequence analysis of the Bacillus subtilis W23 xylR gene and xyl operator.J Bacteriol. 1989 Jul ; 171(7): 3840-5
[6]Kreuzer P, Gärtner D, Allmansberger R, Hillen W. Identification and sequence analysis of the Bacillus subtilis W23 xylR gene and xyl operator. J Bacteriol. 1989 Jul;171(7):3840-5.
[7]Buchler N, Gerland U, Hwa T. On schemes of combinatorial transcription logic. PNAS April 29, 2003 vol. 100no. 9 5136-5141 DOI:10.1073
[8]Bhavsar A, Zhao X, Brown E. Development and characterization of a xylose-dependent systme for expression of cloned genes in Bacillus subtilis: Conditional complementation of a Teichoic acid mutant. Appl Environ Microbiol. 2001 January; 67(1): 403–410. doi: 10.1128/AEM.67.1.403-410.2001
[9]Moore C, Gablla A, Hui M, Ye R W, Helmann J D. Genetic and physiological responses of Bacillus subtillis to metal ion stress. Molecular Microbiology. July 2005 Vol 57(1): 27–40.
[10]Fujita M, Gonzáles-Pastor J E, Losick R. High- and low-threshold genes in the Spo0A regulon of Bacillus subtilis. J. Bacteriol. February 2005 vol. 187 no. 4 1357-1368. DOI: 10.1128/JB.187.4.1357-1368.2005.
[11]Klaus A, Hueck C, Hillen W. Catabolite repression of the Bacillus subtilis xyl operon involves a cis element functional in the context of a unrelated sequence, and glucose exerts additional xylR-dependent repression. J Bacteriol. 1994 March; 176(6): 1738–1745. PMCID: PMC205262
[12]Jarmer H, Larsen T S, et al. Sigma A recognition sites in the Bacillus subtilis genome. Microbiology. September 2001. Vol 147(9):2417-2424.
[13]Helmann J D. Sigma factors in gene expression. Nature. 2001. Encyclopedia of life sciences.
[14]Gärtner D, Geissendörfer, Hillen W. Expression of the Bacillus subtilis xyl operon is repressed at the level of transcription and is induced by xylose. J Bacteriol. 1988. 170(7):3102-3109.
[15]Bintu L, Buchler N, Garcia H G, Gerland U, Hwa T. Transcriptional regulation by the numbers: Models. Genetics & Development. Opinion. 2005. 15(2):116-124.DOI:10.1016
[16]Goryachev A B, Toh D J, Lee J. System analysis of a quorum sensing network: Design constraints imposed by the functional requirements, network topology and kinetic constraints. BioSystems. 2006. 83:178-187.
[17]Procházková K, Cermáková K, Pachl P, et al. Structure of the effector-binding domain of the arabinose repressor AraR from Bacillus subtilis.
[18]Megerle, Judith. Cell to cell variability of gene expression dynamics in inducible regulatory networks. Dissertation of Physics Faculty of Ludwig Maximilians University of Munich. January 2011.
[19]Krispin O, Allmansberger R. The Bacillus subtilis AraE protein displays a broad substrate specificity for several different sugars. J Bacteriol. 1998 June; 180(12): 3250–3252. PMCID: PMC107832.
[20]Gu Y, Ren C, Sun Z, Rodionov D A, Zhang W, Yang S, Yang C, Jiang W. Reconstruction of xylose utilization pathway and regulons in Firmicutes. BMC Genomics 2010, 11:255 doi:10.1186/1471-2164-11-25.
[21]Moore C, Helmann J D. Metal ion homeostasis in Bacillus subtilis. Microbiology. April 2005. 8(2):188-195 DOI: 10.1016/j.mib.2005.02.007
[22]Sá-Nogueira I, Nogueira T V, Soares S, Lencastre H. The Bacillus subtilis L-arabinose (ara) operon: nucleotide sequence, genetic organization and expression. Microbiology. 1997. 143:957-969.
[23]Chakravorty D, Wang B, Won Lee C, Giedroc D P, Merz K M. Simulations of allosteric motions in the zinc sensor CzrA. J. Am. Chem. Soc. 2012, 134(7):3367-3376.
 "
"






