Team:Goettingen/Notebook/Results
From 2012.igem.org
| Line 72: | Line 72: | ||
===Electron microscopy=== | ===Electron microscopy=== | ||
| + | [[Image:EM_pics.jpg|thumb|300px|right|'''Fig. 11:''']] | ||
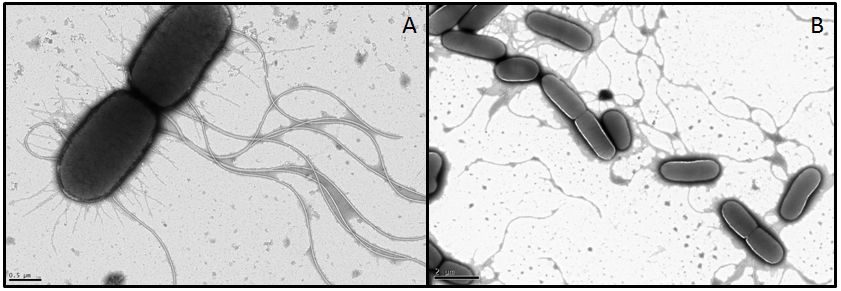
Overexpression of flagellar genes like ''fliC'' or the ''flhDC'' operon should result in alterations of the flagellum structure or number, which can be visualized. In the case of ''fliC'', which encodes flagellin, the building block of the flagellum, we expect the emergence of longer filaments. In contrast to this, ''flhDC'' as the master regulator is thought to increase the number of flagella when overexpressed, since all genes involved in flagellum assembly should be up-regulated.<br> | Overexpression of flagellar genes like ''fliC'' or the ''flhDC'' operon should result in alterations of the flagellum structure or number, which can be visualized. In the case of ''fliC'', which encodes flagellin, the building block of the flagellum, we expect the emergence of longer filaments. In contrast to this, ''flhDC'' as the master regulator is thought to increase the number of flagella when overexpressed, since all genes involved in flagellum assembly should be up-regulated.<br> | ||
In order to visualize these alterations we investigated the ''E. coli'' strain BL21, transformed with ''fliC'' or ''flhDC'' constructs, by using transmission electron microscopy. Both genes were under control of a strong constitutive promoter promoter (J23100). As reference strains we used BL21 cells carrying the same vector with an RFP insert. Furthermore the MG1655 strain, as a more motile strain with non-degenerated flagella, was used. | In order to visualize these alterations we investigated the ''E. coli'' strain BL21, transformed with ''fliC'' or ''flhDC'' constructs, by using transmission electron microscopy. Both genes were under control of a strong constitutive promoter promoter (J23100). As reference strains we used BL21 cells carrying the same vector with an RFP insert. Furthermore the MG1655 strain, as a more motile strain with non-degenerated flagella, was used. | ||
Revision as of 14:23, 26 September 2012
 | |
Deutsch  / English / English  |
 |
Contents |
Homing coli: Engineering E. coli to become tracking dogs
The model organism Escherichia coli is naturally capable of sensing substances in its environment and consequently moves directionally towards these, a phenomenon known as chemotaxis. Here, we apply directed evolution to chemoreceptors by targeting five amino acid residues in the ligand binding site to enable E. coli to perceive novel substances. In order to investigate mobility and directed movement towards a substance, an effective mobility selection method using special "swimming plates" is designed. Additionally, we attempt to improve E. coli's swimming velocity by creating new parts derived from its own motility apparatus. Based on our selection system, we identify variants of chemoreceptors with new binding specificities in the mutant library. By these means, we aim to train the bacterium to detect new molecules such as tumor cell markers. Once having established E. coli as our "tracking dog", the possible applications in medicine but also to environmental issues are virtually countless.
#1 - Selection / Swimming
#2 - Speed Improvement
The goal of our group was to enhance the motility of E. coli in order to allow them to reach their targets more efficiently in later applications. This also results in a a faster selection of our Homing Coli and quick generation of results. The BioBricks we designed are therefore primarily directed at flagellum efficiency or number. We selected five different genes that were tested along our project in different strains. E. coli strain BL21 was used as our main object of research because this is a typical lab-strain and therefore usually not very motile. Our goal was to find a way to accelerate BL21 E. coli by transforming them with our constructs. Here is a overview of the genes we tested and the corresponding results.
motA and motB
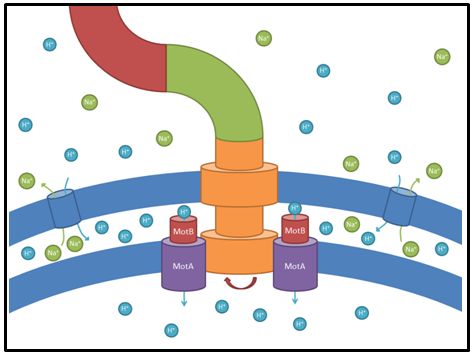
These genes code for proteins that build the stator part of the bacterial flagellum which generates torque by using a proton gradient that exists across the membrane (Fig. 5). Flagellum function depends on copy number of motA and motB (Van Way et al. 2000). Therefore we chose to test these genes, hoping that the number of stator protein copies might have a positive effect on motility (Reid et al. 2006). We first tested our strains with motA and motB constructs containing the natural 3’ and 5’ regions in puc18 because a strong overexpression might have counterproductive effects. High expressed motA might thereby lead to proton leakage and disturb the metabolism of E. coli.
- Results
- For these two stator-genes it was very hard to get reproducible data. However, after several rounds of testing on different swimming agars and conditions we were able to conclude that overexpression of motB often had a mild positive effect on motility of BL21 E. coli (Fig. 6). However, increased expression on motA seemed to have no or even a slightly negative effect.
- We did not see very strong effects after testing these two candidates. A possible explanation could be that the number of functional stator elements at one flagellum could not be increased anymore or might just not have a significant impact. Furthermore the expression of motA and motB is usually regulated in concert and if out of balance, their expression might not cause any positive effect for motility. For future projects it might be interesting to test chimeric stator proteins that could harbor higher efficiencies or use other ions for torque generation.
yhjH
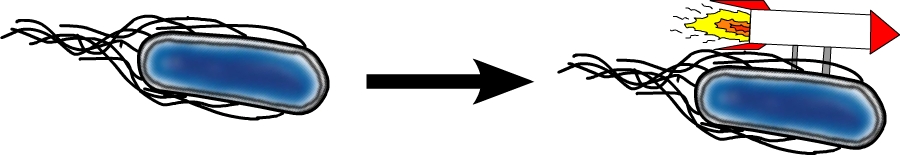
This gene codes for a phosphodiesterase which reduces the levels of c-di-GMP that are involved in cell motility (Ko and Park, 2000). The protein YcgR is able to bind c-di-GMP and act as a flagellar brake which consequently reduces motility (Paul et al. 2010). The mechanism that we would like to exploit is illustrated in Figure 7. Overexpression of yhjH should therefore result in lower levels of c-di-GMP and increase E. coli's speed by diminishing the ability of YcgR to act as a brake on the flagellum. The second messenger c-di-GMP furthermore plays a role in the regulation of biofilm development. Expression of yhjH on the other hand represses biofilm formation (Suzuki et al. 2006) which could also serve our purposes and induce swimming motility.
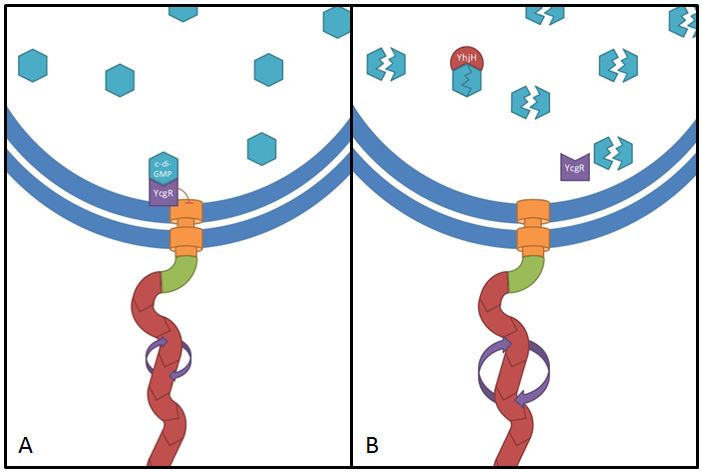 Fig. 7: The yhjH gene encodes for a c-di-GMP phosphodiesterase. The substrate, c-di-GMP, is a second messenger that binds to YcgR, a protein that functions as a flagellar brake and thus down-regulates the motor (A). We suggest that the overexpression of yhjH results in increased c-di-GMP degradation and hence in narrowed braking force. Due to a stronger rotation of the flagellum higher motility could be achieved (B). References: http://www.uniprot.org/uniprot/P37646; http://www.uniprot.org/uniprot/P76010 | 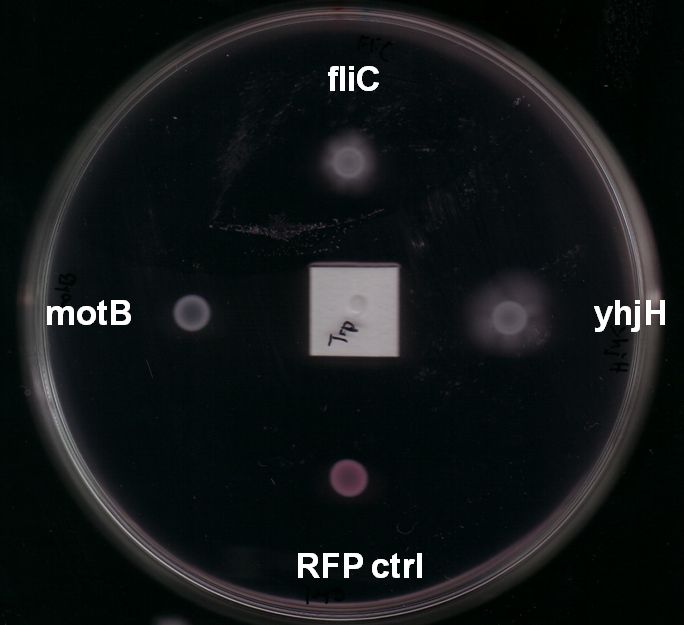 Fig. 8: BL21 E. coli carrying different constructs on Tryptone swimming agar after 12h incubation at 33°C. Cells expressing yhjH in puc18 under the natural promoter travelled approximately 0.5cm (radius) whereas no swimming could be detected for the control plasmid carrying [http://partsregistry.org/Part:BBa_K777125 K777125]. |
- Results:
- Expression of yhjH under its natural promoter in puc18 increased motility in almost all of our assays on LB or Tryptone swimming agar (Fig. 6 & Fig. 8). Not only were the yhjH expressing strains among the fastest on these swimming agar plates but also usually the first colonies to start swimming. This might be an indicator that indeed biofilm formation was inhibited and the E. coli were able to spread faster. Compared to our other tested constructs we got the most reliable results for yhjH. However the positive effect on motility was not present when we used M9 minimal medium swimming agar. Here, yhjH-transformed cells usually did not show a strong tendency for swimming. This result suggests that the motility-effects of YhjH are somehow nutrient-dependent and therefore depend on the used media.
fliC
References:
http://ecocyc.org/ECOLI/NEW-IMAGE?type=POLYPEPTIDE&object=EG10321-MONOMER
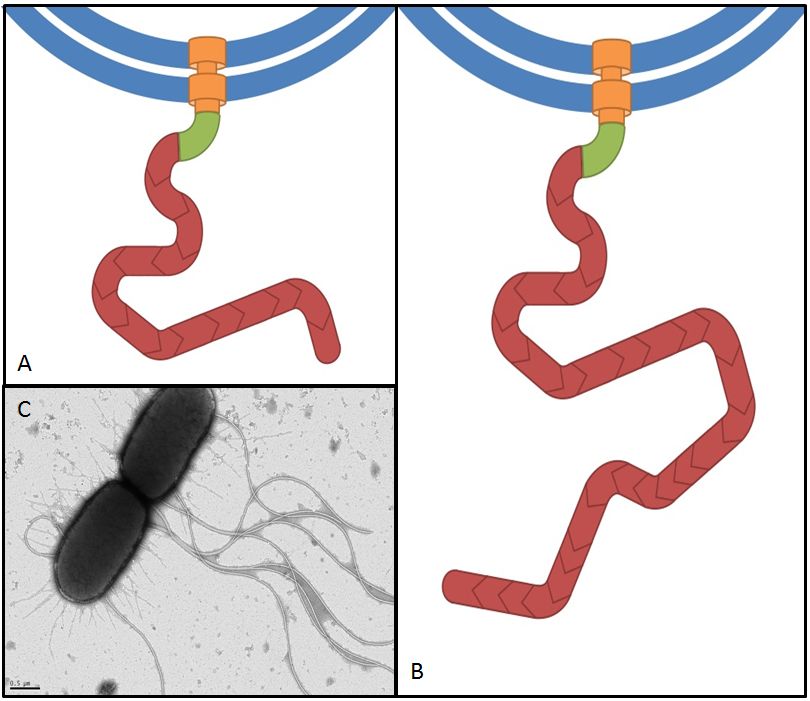
Flagellin, the product of the fliC gene is the structural protein that builds up the filament of the bacterial flagellum (Fig. 9). By overexpressing fliC we hope to construct longer flagella which might have a positive impact on swimming speed (Furuo et al. 1997). At the same time we were aware that overlong flagella might cause adverse effects due to an obstructive architecture.
- The fliC gene was amplified from DH10B E. coli genomic DNA and contained four forbidden restriction sites. We had to mutate all these sites via overlap PCR. A description for this method can be found here.
- Results
- Our overlap PCR for the mutation of three PstI and one SpeI sites worked efficiently and fast. The resulting PCR product was not cut by any BioBrick standard 10 enzymes.
- In order to avoid too strong levels of expression we included 1kb of the upstream region for most of the assays to allow a certain level of regulation in the cells. Like yhjH-transformants these strains showed increased motility in almost all of our assays (see Fig. 8). On the minimal medium M9 agar plates they were usually significantly faster and showed swimming motility earlier than any other cells. On Tryptone swimming agar they were in general the runner up behind yhjH-transformed E. coli. Unfortunately, we had trouble cloning our fliC constructs into pSB1C3 because of a problem in our reverse primer suffix. When we had located the source of the problem it was too late to send this part to the Registry in time.
flhDC
References:
[1] Wei et al. (2001) Positive regulation of motility and flhDC expression by the RNA-binding protein CsrA of Escherichia coli. Molecular Microbiology, 40(1):245-256
[2] Gauger et al. (2007) Role of motility and the flhDC operon in Escherichia coli MG1655 colonization of the mouse intestine. Infect immune 75(7):3315-3324
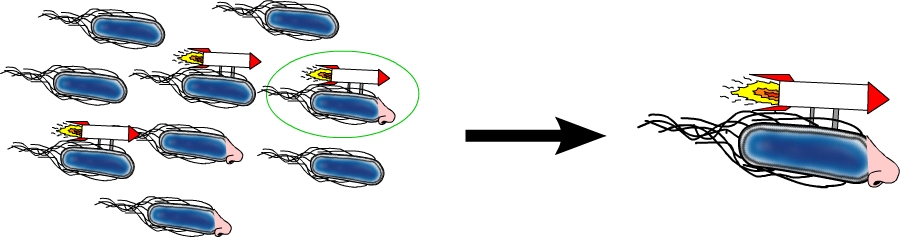
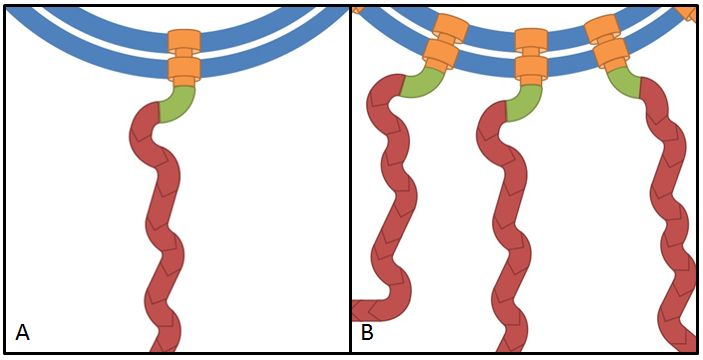
The flhDC operon is the master regulator of motility and chemotaxis in E. coli. This means that it is the main control instance for flagellar synthesis and starts the complex process of flagellar gene synthesis and flagellum assembly (Chevance and Hughes 2008). The flhDC operon codes for the transcriptional regulator FlhD4C2 which forms heterotetramers and activates class II operons in concert with sigma factor 70. Among the gene products of class II operons are several components of the flagellum and the alternative sigma factor FliA which is essential for the transcription of class III genes. It has been shown that increased expression of flhDC also enhances motility in E. coli (Ling et al. 2010). We wanted to see how different levels of flhDC expression affect the speed of our E. coli lab-strains. In theory, the number of assembled flagellums should increase (Fig. 10) and thereby have a positive impact on motility.
- Results
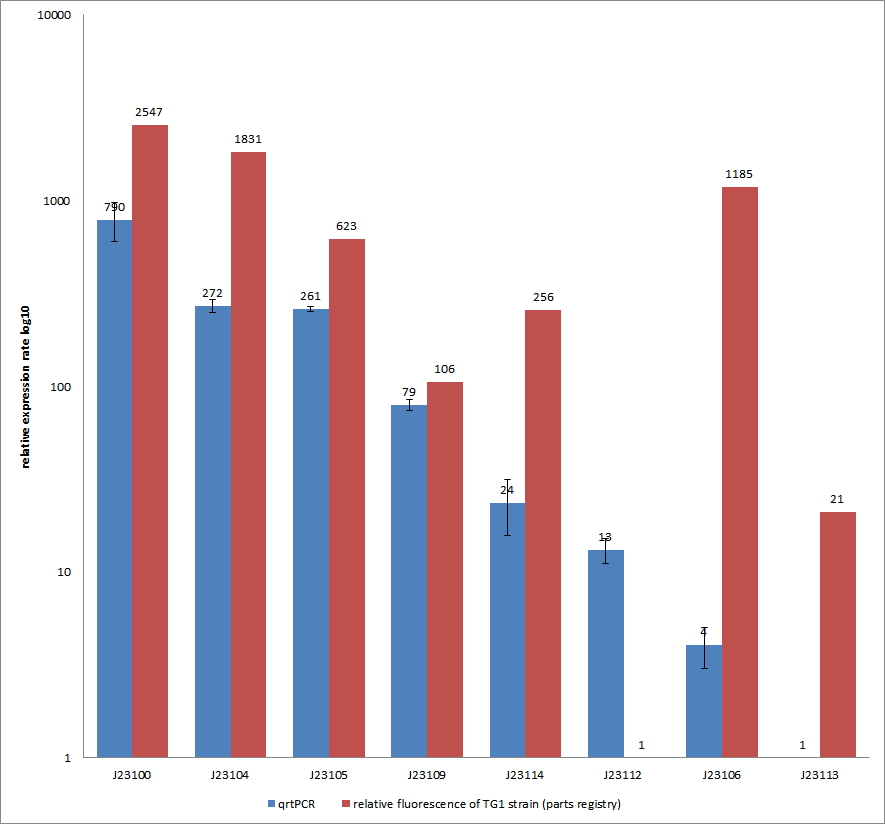
- Our plan was to express flhDC constitutively with a selection of 8 different Anderson-promoters to achieve different levels of transcription. And indeed BL21 E. coli transformed with these constructs showed a higher motility than empty vector controls or WT cells. However, our results were very inconsistent, especially concerning the different promoter strengths when compared to each other. The troubleshooting revealed that we were missing a RBS between the promoters and the ATG within all our constructs. This probably explains the inconsistent results that did not allow us to quantify the effect of our constructs. We hope to finish these experiments with correct constructs soon because we are eager for presentable results!
Electron microscopy
Overexpression of flagellar genes like fliC or the flhDC operon should result in alterations of the flagellum structure or number, which can be visualized. In the case of fliC, which encodes flagellin, the building block of the flagellum, we expect the emergence of longer filaments. In contrast to this, flhDC as the master regulator is thought to increase the number of flagella when overexpressed, since all genes involved in flagellum assembly should be up-regulated.
In order to visualize these alterations we investigated the E. coli strain BL21, transformed with fliC or flhDC constructs, by using transmission electron microscopy. Both genes were under control of a strong constitutive promoter promoter (J23100). As reference strains we used BL21 cells carrying the same vector with an RFP insert. Furthermore the MG1655 strain, as a more motile strain with non-degenerated flagella, was used.
- Results
- For MG1655 nice pictures were obtained since the strain grew very fast and seems to be quite robust (Fig. 11). However, we were not able to obtain meaningful pictures showing flagella of the BL21 strain. Under stress conditions bacteria tend to lose their flagella very easily (Blum, 1971). Apparently the transport form our Lab to the EM-facility was enough to stress the BL21 E. coli. It appears that this demands quite something for MG1655 however, or the pictures we got from this strain were already in a stressed state and the flagella are usually even longer or more in number. We had a look at the different strains with a light microscope in our lab before the EM sessions and they were all motile. This consideration, that the cells were stressed during the transport was supported by their subsequent investigation under a light microscope. The cells were not motile anymore and featured very heterogeneous shapes whereby some formed very long rods. All these appearances are indications for stressed cells. Another problem with BL21 was the very low growth rate, since a high optical density is required in order to obtain an adequate number of cells during preparation.
- In conclusion, we were not able to get informative data about the flagellar structures and morphologies, but we found out, that BL21 is very sensitive and drops its flagella in no time.
- For MG1655 nice pictures were obtained since the strain grew very fast and seems to be quite robust (Fig. 11). However, we were not able to obtain meaningful pictures showing flagella of the BL21 strain. Under stress conditions bacteria tend to lose their flagella very easily (Blum, 1971). Apparently the transport form our Lab to the EM-facility was enough to stress the BL21 E. coli. It appears that this demands quite something for MG1655 however, or the pictures we got from this strain were already in a stressed state and the flagella are usually even longer or more in number. We had a look at the different strains with a light microscope in our lab before the EM sessions and they were all motile. This consideration, that the cells were stressed during the transport was supported by their subsequent investigation under a light microscope. The cells were not motile anymore and featured very heterogeneous shapes whereby some formed very long rods. All these appearances are indications for stressed cells. Another problem with BL21 was the very low growth rate, since a high optical density is required in order to obtain an adequate number of cells during preparation.
References
- Blum J.J.(1971) Existence of a breaking point in cilia and flagella. J. Theor. Biol. 33:257–263.
- Chevance F. F., Hughes K. T. 2008. Coordinating assembly of a bacterial macromolecular machine. Nat Rev Microbiol. 6: 455–465.
- Furuno, M., T. Atsumi, T. Yamada, S. Kojima, N. Nishioka, I. Kawagishi, and M. Homma. 1997. Characterization of polar-flagellar-length mutants in Vibrio alginolyticus. Microbiology. 66: 3632–3636.
- Ko M., Park C. 2000. Two novel flagellar components and H-NS are involved in the motor function of Escherichia coli. J. Mol. Biol. 303: 371-382.
- Ling H., Kang A., Tan M.H., Qi X., Chang M.W. 2010. The absence of the luxS gene Increases swimming motility and flagella synthesis in Escherichia coli K12. Biochem Biophys Res Commun. 401: 521-526.
- Paul, K., Nieto, V., Carlquist, W.C., Blair, D.F., Harshey, R.M. The c-di-GMP Binding Protein YcgR Controls Flagellar Motor Direction and Speed to Affect Chemotaxis by a “Backstop Brake” Mechanism. 2012. Mol Cell. 38: 128–139.
- Reid, S. W., M. C. Leake, J. H. Chandler, C. J. Lo, J. P. Armitage, and R. M. Berry. 2006. The maximum number of torque-generating units in the flagellar motor of Escherichia coli is at least 11. Proc. Natl. Acad. Sci. USA 103: 8066-8071.
- Suzuki, K., Babitzke, P., Kushner, S. R., Romeo, T. 2006. Identification of a novel regulatory protein (CsrD) that targets the global regulatory RNAs CsrB and CsrC for degradation by RNase E. Genes Dev. 20: 2605–2617.
- Van Way, S. M., E. R. Hosking, T. F. Braun, and M. D. Manson. 2000. Mot protein assembly into the bacterial flagellum: a model based on mutational analysis of the motB gene. J. Mol. Biol. 297: 7-24.
#3 - Chemoreceptor Library


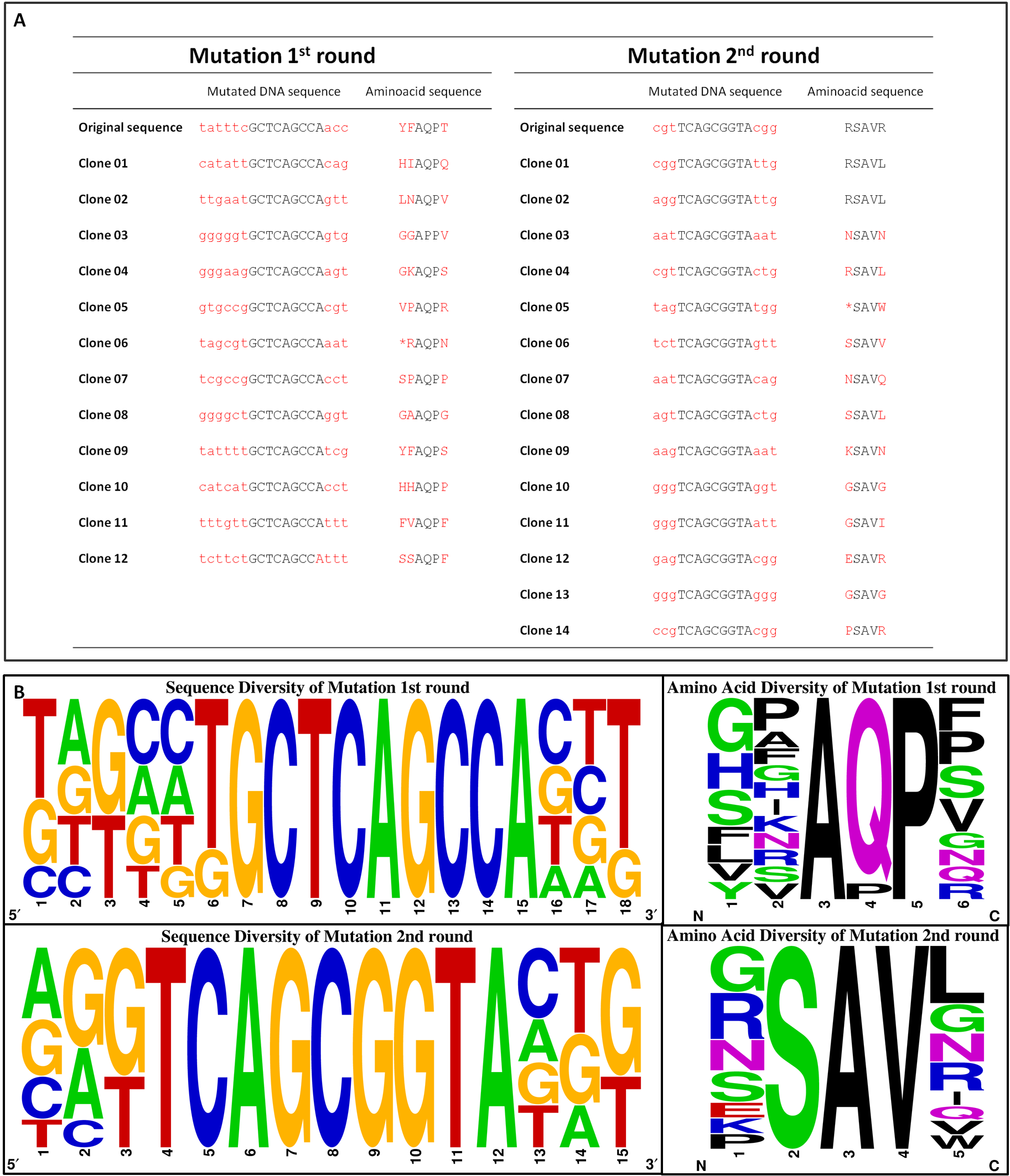
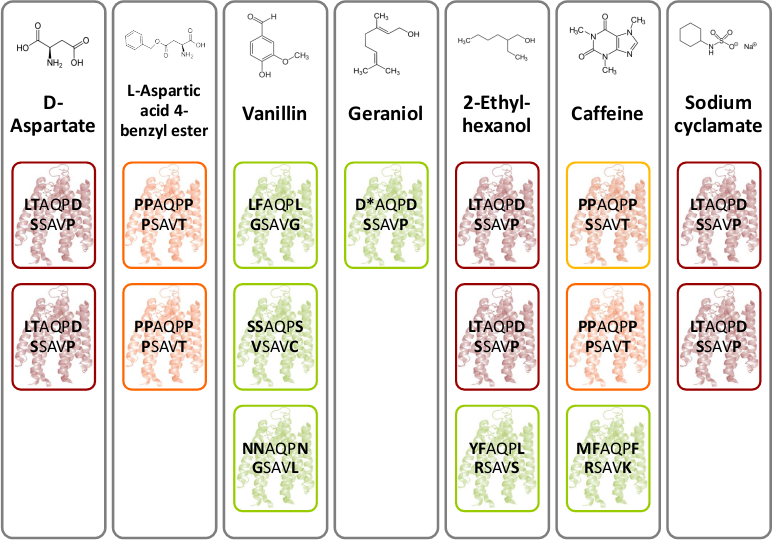
To get rid of BioBrick standard restriction sites, the QuikChange reaction is applied. In the case of the Tar receptor, the BsaI site will be exchanged while keeping the codon for the same amino acid.
To generate a chemoreceptor library our method of choice requires the absence of the BsaI restriction site in both, the insert and the vector backbone, of the cloned plasmid. Therefore, pUC18 containing one BsaI site in the bla gene seems to be not appropriate. Thus, we moved on cloning our promoter constructs with the quik changed Tar (TAR_QC) at the XbaI restriction site into the pSB1C3 BioBrick vector. There are two advantages coming along: Firstly, we need to send our designed biobricks this year in this particular vector and secondly, this vector contains a chloramphenicol resistance gene and hence, lacking the undesired BsaI site.
| ↑ Back to top! |

|
 "
"