Team:TU Darmstadt/Labjournal/Transport
From 2012.igem.org
Transport
Overview
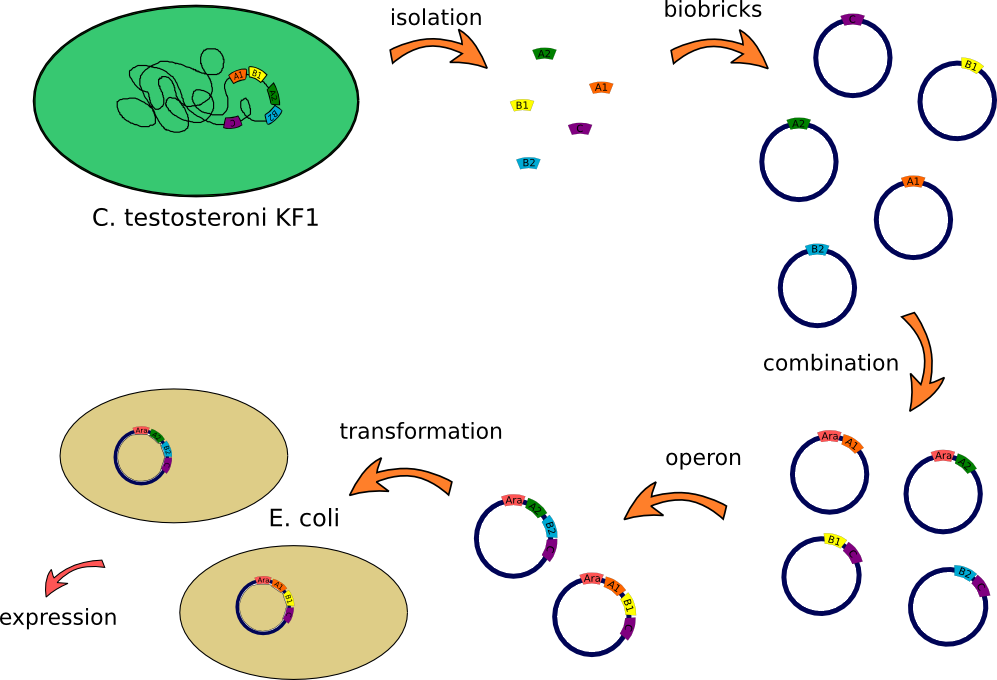
This lab journal describes a isolation and characterisation of the putative terephtalate (TPA) uptake system from [http://www.dsmz.de/catalogues/details/culture/DSM-14576.html?tx_dsmzresources_pi5%5BreturnPid%5D=304Comamonas testosteroni KF-1] (C. testosteroni) in Escherichia Coli (E.Coli). The putative terephtalate system consist of three different domain, two alpha-helical transmembrane protein (11-12 and 4-5 alpha-helical transmembrane protein ) and one periplasmic terephtalate binding protein. The strain was purchased from [http://www.dsmz.de/ DMSZ-German Collection of Microorganism and Cell Cultures] ([http://www.dsmz.de/catalogues/details/culture/DSM-14576.html?tx_dsmzresources_pi5%5BreturnPid%5D=304 DMSZ no.14576]). We designed two putative TPA uptake operons and the intake of TPA was checked by photometry, gas chromatography-mass spectrometry (GC-MS)and energy dispersive X-ray spectroscopy (EDX).
Week 1 (21.-25.05.12)
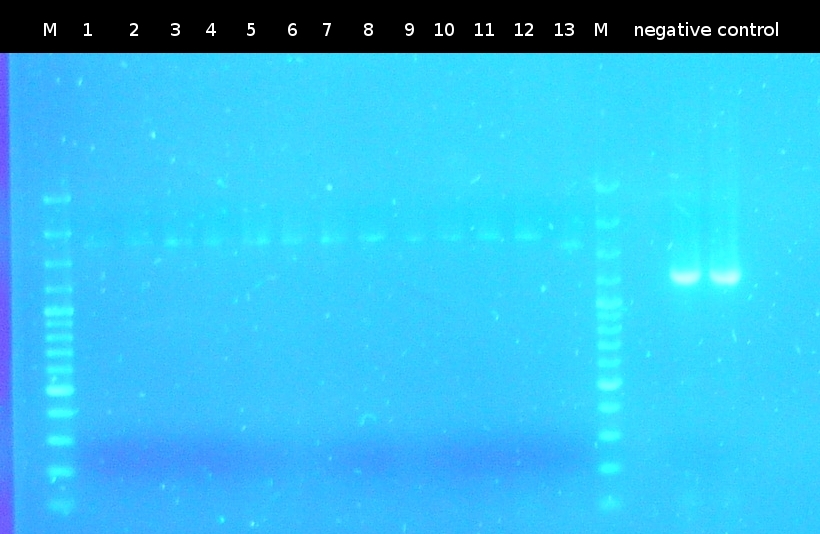
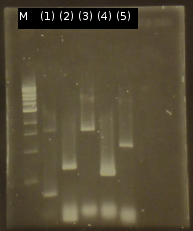
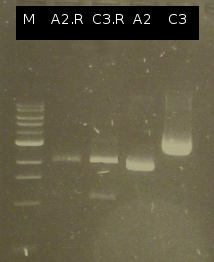
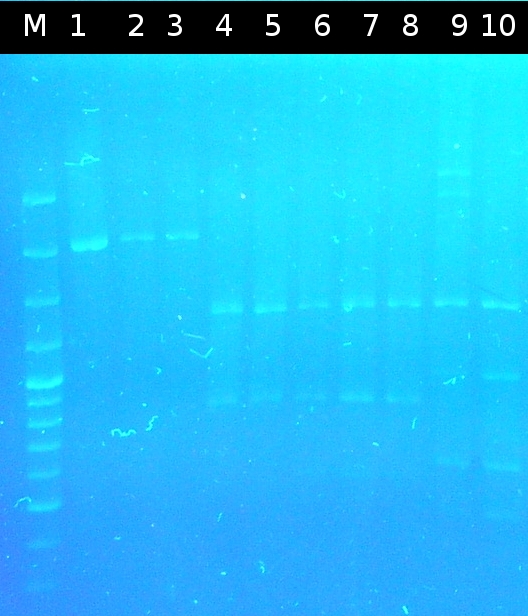
- First colony PCR of Comamonas testosteroni for isolation of the following genes: [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808004 tctA 505aa] (1), [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808005 tctB 197aa] (2), [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808002 tctA_503aa] (3), [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808003 tctB 162aa] (4), [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808001 tphC 322aa] (5). (Primers: tctA_503 Biobrick, tctA_505 Biobrick, tctB_162 Biobrick, tctB_197 Biobrick, tphC Biobrick). Analysis of [http://en.wikipedia.org/wiki/Polymerase_chain_reaction PCR] products by Agarose gel electrophoresis
- PCR product purification using the Promega-Kit
- Nanodrop measurements of the purified PCR products:
- (1) : 19.5 ng/µl 260/280=1.58
- (2) : 61.9 ng/µl 260/280=1.75
- (3) : 75.1 ng/µl 260/280=1.75
- (4) : 93.3 ng/µl 260/280=1.85
- (5) : 91.3 ng/µl 260/280=1.58
- Restriction digest of PCR products by SpeI and EcoRI, heat inactivation after digestion
- Analysis of restriction digest by Agarose gel electrophoresis
- Purification of the bands gained form gel electrophoresis (Promega-Kit)
- Nanodrop measurements of the purified products:
- (1) : 8.3 ng/µl 260/280=1.94
- (2) : 8.9 ng/µl 260/280=1.80
- (3) : 13.0 ng/µl 260/280=1.57
- (4) : 1.5 ng/µl 260/280=3.08
- (5) : 3.7 ng/µl 260/280=2.26
Results
- We got the purified Genes from C. testosteroni but the yield after purification is to low for futher use
week 2 (28.05.-01.06.12)
- Purified products from 1. week were ligated into pSB1A2 and subsequently transformed into DH5a (in spite of low concentration)
- Overnight incubation on LB agar plates; no growth detectable
- Second approach to isolate the five genes from Comamonas t. using colony-PCR (Primers: tctA_503 Biobrick, tctA_505 Biobrick, tctB_162 Biobrick, tctB_197 Biobrick, tphC Biobrick)
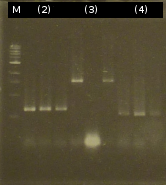
- Analysis of PCR products by Agarose gel electrophoresis; without results; modification of the PCR protocols: adding DMSO
- Third approach to isolate the five genes from Comamonas t. by colony-PCR (Primers: tctA_503 Biobrick, tctA_505 Biobrick, tctB_162 Biobrick, tctB_197 Biobrick, tphC Biobrick)
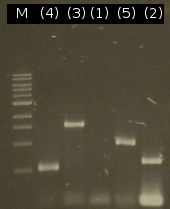
- PCR product purification using the Promega-Kit: ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K808005 tctB 197aa] (2), [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808002 tctA_503aa] (3), [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808003 tctB 162aa] (4), [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808001 tphC 322aa] (5))
- Nanodrop measurements of the purified PCR products:
- (2) : 56.1 ng/µl 260/280=1.87
- (3) : 66.8 ng/µl 260/280=1.92
- (4) : 72.8 ng/µl 260/280=1.87
- (5) : 68.8 ng/µl 260/280=1.82
Results
- Finally we got the genes 2, 3, 4 and 5 in acceptable concentrations
week 3 (04.-08.06.12)
- Restriction digest of the purified PCR products and of [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808004 tctA 505aa] (leftover of the 1. week) by SpeI and EcoRI
- Restriction digest of pSB1A2 using SpeI and EcoRI
- Dephosphorylation of the restriction reactions by using antarctic Phosphatase
- Ligation of the five genes into pSB1A2 and subsequently transformation into DH5a (incubation at 37°C)
- colony-PCR of [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808004 tctA 505aa] (1), 2 approaches (Primers:tctA_505 Biobrick)
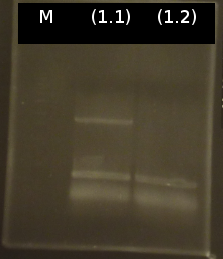
- Analysis of PCR products by Agarose gel electrophoresis. Only one of them showed the expected band. The band was excluded and purified via the Promega-Kit.
- Nanodrop measurement of the purified product:
- (1.1) : 8.9 ng/µl 260/280= 2,22
- Because of low concentration and contamination the PCR was repeated and the product was purified again by the Promega-Kit.
- Performing a colony-PCR-screen of the transformed cells using the Taq-Polymerase. Verification of 4 colonies per plate. (Primers: tctA_503 Biobrick, tctA_505 Biobrick, tctB_162 Biobrick, tctB_197 Biobrick, tphC Biobrick)
- PCR did not work
- Testing the Taq-Polymerase for function. The Phu-Polymerase was used for comparison. Only the approach using the Phu-Polymerase showed the expected bands in Agarose gel electrophoresis.
- Screening was repeated with Phu-Polymerase; no bands visible
Results
- We ligated the purified and restricted genes into pSB1A2 and transformed it into DH5a but the colony-PCR didn't work.
- We tried to isolate gene 1 again but after purification the concentration was too low
week 4 (11.-15-06.12)
- Inoculation of LB-media with DH5a_pSB1A2 and DH5a_pSB1C3; overnight incubation at 37°C
- Miniprep of DH5a_pSB1A2 and DH5a_pSB1C3 using the Promega-Kit
- Nanodrop measurements of the preparation:
- pSB1A2 : 136.1 ng/µl 260/280= 1.93
- pSB1C3 : 265.1 ng/µl 260/280= 1.89
- Restriction digest of pSB1A2 and pSB1C3 with EcoRI and SpeI. Afterwards dephosphorylation by using Antarctic phosphatase.
- Analysis of the restriction products through Agarose gel electrophoresis; for comparison the uncropped vectors were also analyzed.
- pSB1C3 shows several bands (Insert, vector without insert, linearized vector with insert); concentration of pSB1A2 is very low, only one band visible
- Approach to isolate the following genes from Comamonas t. by colony-PCR: [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808004 tctA 505aa] (1), [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808005 tctB 197aa] (2), [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808002 tctA_503aa] (3), [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808003 tctB 162aa] (4) and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808001 tphC 322aa] (5) by [http://en.wikipedia.org/wiki/Polymerase_chain_reaction PCR] (Primers: tctA_503 Biobrick, tctA_505 Biobrick, tctB_162 Biobrick, tctB_197 Biobrick, tphC Biobrick)
- Analysis of PCR products by Agarose gel electrophoresis
- PCR product purification by using the Promega-Kit
- Nanodrop measurements of the purified PCR products:
- (2) : 18.3 ng/µl 260/280=2.03
- (3) : 21.2 ng/µl 260/280=1.95
- (4) : 23.9 ng/µl 260/280=2.05
- Restriction digest of purified PCR products (2), (3) and (4) using SpeI and EcoRI
- Ligation of the digested genes ((2), (3), (4)) into pSB1A2 and pSB1C3
- Transformation of DH5a with pSB1A2_197, pSB1A2_503, pSB1A2_162, pSB1C3_197, pSB1C3_503, pSB1C3_162 (6 different transformation approaches). Overnight incubation at 37°C
- Screening of the transformants by colony-PCR (Primers: tctA_503 Biobrick, tctA_505 Biobrick, tctB_162 Biobrick, tctB_197 Biobrick, tphC Biobrick). Verification of 6 colonies for every transformation. Afterwards the PCR products were analyzed by Agarose gel electrophoresis
- Colony-PCR of Comamonas t. to isolate [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808004 tctA 505aa] (1) und [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808001 tphC 322aa] (5) (Primers: tctA_505 Biobrick, tphC Biobrick)
- Analysis of PCR products (Colony-PCR) and the ligation approach by Agarose gel electrophoresis
- Restriction digest of [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808001 tphC 322aa] (5) and subsequent Ligation into pSB1A2 and pSB1C3
- Transformation of DH5a with pSB1A2_322 und pSB1C3_322. [http://en.wikipedia.org/wiki/Polymerase_chain_reaction PCR] (Primers: tphC Biobrick) used to check for successful transformation
- Isolation of C. testosteroni genome, because it seems impossible to amplificate [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808004 tctA 505aa] (1) from a fluid Comamonas t. culture.
Results
- We started an other approach to isolate the five genes. All but gene 1 worked, so we isolated the C. testosteroni genome
- We purified, restricted and ligated the genes 2-4 into pSB1A2 and pSB1C3 and transformed them into DH5a
- We checked them by colony-PCR and made fluid cultures of the positive colonies
week 5 (18.-22.06.12)
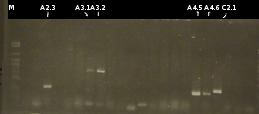
- Introduction of a nomenclature: A/C gene/colony: pSB1A2 = A; pSB1C3 = C; gene: tctA 505aa = 1, tctB 197aa = 2, tctA 503aa = 3, tctB 162aa = 4, tphC 322aa = 5; colony = 1...X
- Producing fluid cultures of the positive transformants
- Miniprep of the fluid culures by using the Frementas-Kit. Afterwards Nanodrop measurements of the preparations
- A2.3 : 125.9 ng/µl 260/280= 1.62
- A3.1 : 103.3 ng/µl 260/280= 1.72
- A4.6 : 147.2 ng/µl 260/280= 1.70
- A4.5 : 108.7 ng/µl 260/280= 1.86
- A5.12 : 88.0 ng/µl 260/280= 1.87
- A5.8 : 85.01 ng/µl 260/280= 1.78
- A3.2 : 87.3 ng/µl 260/280= 1.81
- C4.6 : 70.5 ng/µl 260/280= 1.81
- C4.1 : 41.2 ng/µl 260/280= 1.85
- C4.4 : 104.2 ng/µl 260/280= 1.85
- C2.1 : 132.2 ng/µl 260/280= 1.85
- C2.6 : 188.1 ng/µl 260/280= 1.75
- [http://en.wikipedia.org/wiki/Polymerase_chain_reaction PCR] using a rest of purified [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808004 tctA 505aa] (1) (Primers: tctA_505 Biobrick), subsequent Agarose gel electrophoresis
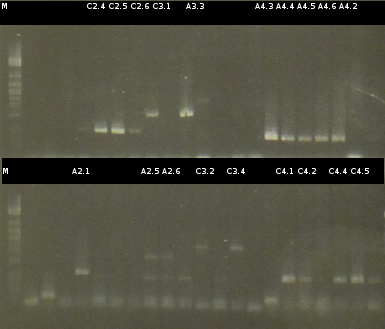
- Another colony-PCR using the transformants (2)-(5) as template (Primers: tctA_503 Biobrick, tctB_162 Biobrick, tctB_197 Biobrick, tphC Biobrick); analyzing the PCR product by Agarose gel electrophoresis
- 3 positive transformants were picked for inoculation of fluid media (A34, A43; C513)
- Miniprep (Fermentas Kit) of the fluid culures and Nanodrop measurements:
- A2.4 : 202,1 ng/µl 260/280= 1,63
- A2.5 : 126,5 ng/µl 260/280= 1,90
- A2.6 : 107,5 ng/µl 260/280= 1,81
- A4.2 : 133,8 ng/µl 260/280= 1,59
- C2.3 : 187,8 ng/µl 260/280= 1,87
- C2.4 : 270,7 ng/µl 260/280= 1,80
- C4.1 : 103,8 ng/µl 260/280= 1,80
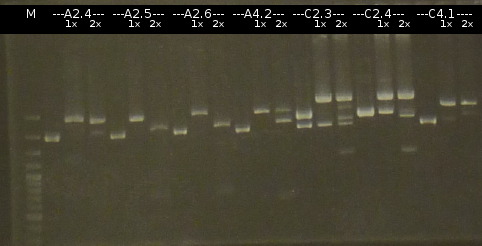
- The Miniprep products were Restriction digested by restriction enzymes, each was cut once with EcoRI and twice with EcoRI/SpeI. Analisys of the digested products by Agarose gel electrophoresis
week 6 (25.-29.06.12)
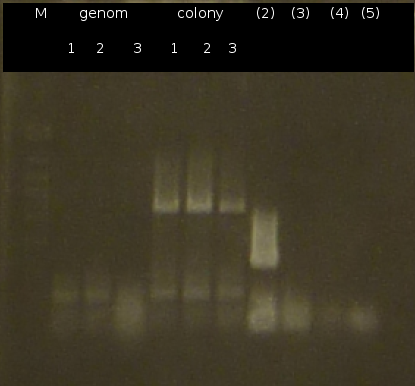
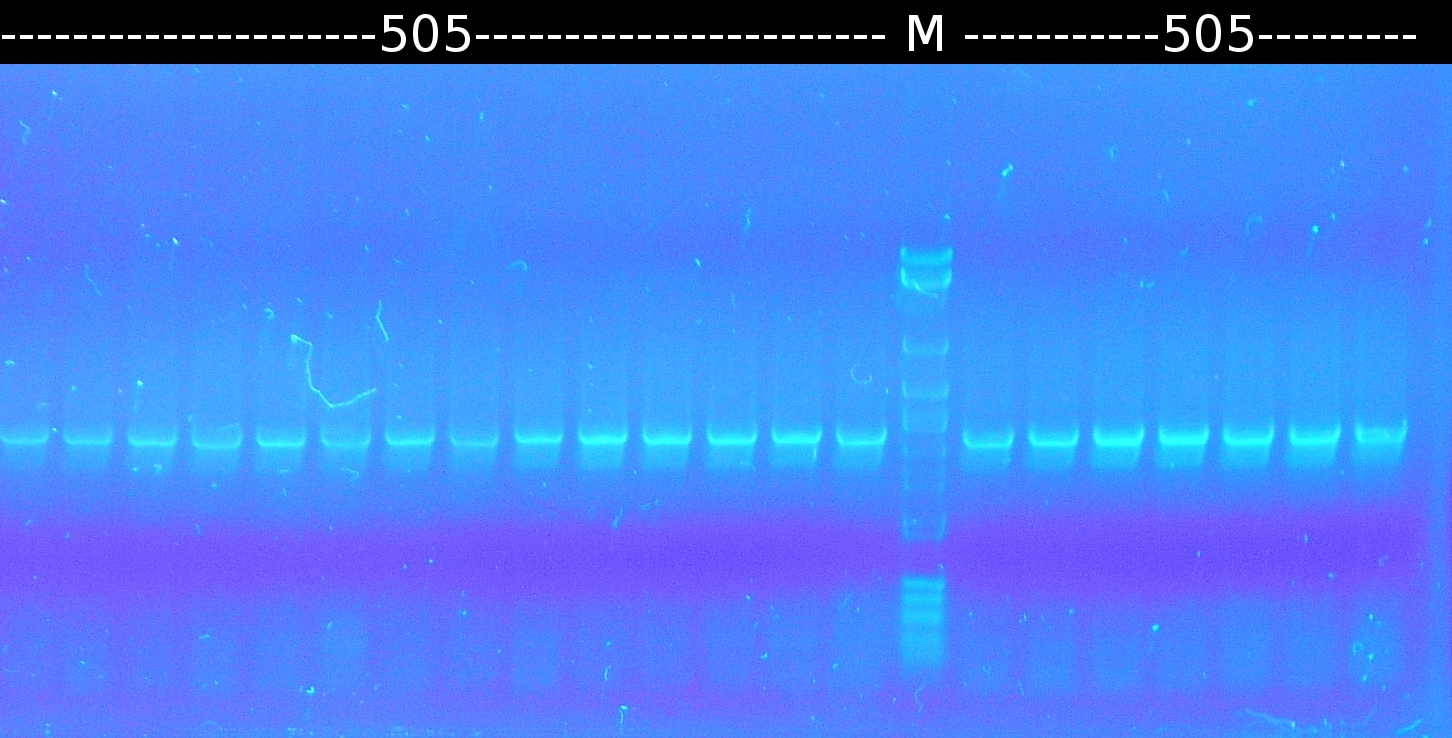
- Further colony-PCR of Comamonas t. (3 approaches) and [http://en.wikipedia.org/wiki/Polymerase_chain_reaction PCR] with genomic Comamonas t. DNA as template to obtain [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808004 tctA 505aa] (1), 2 approaches each (Primers: tctA_505 Biobrick). Subsequent analisys of the PCR products by Agarose gel electrophoresis. All PCR of [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808004 tctA 505aa] show only the band of the undesired smaller product.
- Ligation off the digested genes into pSB1A2 and transformation into DH5a. Overnight incubation at 37°C
- Colony-PCR of Comamonas t. and [http://en.wikipedia.org/wiki/Polymerase_chain_reaction PCR] with genomic Comamonas t. DNA as template to obtain [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808004 tctA 505aa] (1). [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808005 tctB 197aa] (2), [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808002 tctA_503aa] (3), [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808003 tctB 162aa] (4), [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808001 tphC 322aa] (5) used as control (Primers: tctA_503 Biobrick, tctA_505 Biobrick, tctB_162 Biobrick, tctB_197 Biobrick, tphC Biobrick)
- Miniprep and Nanodrop measurement of:
- A3.1 : 111.6 ng/µl 260/280= 1.72
- A3.4 : 79.5 ng/µl 260/280= 1.66
- C4.6 : 69.6 ng/µl 260/280= 1.76
- A4.2 : 96.9 ng/µl 260/280= 1.62
- A2.4 : 109.7 ng/µl 260/280= 1.72
- A2.5 : 160.8 ng/µl 260/280= 1.79
- A5.13 : 188.7 ng/µl 260/280= 1.85
- A5.12 : 114.5 ng/µl 260/280= 1.81
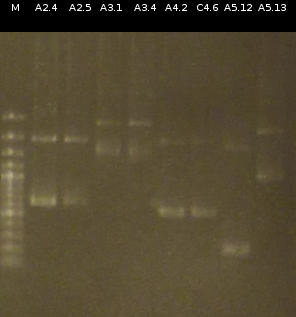
- Checking the Miniprep with [http://en.wikipedia.org/wiki/Polymerase_chain_reaction PCR] (Primers: tctA_503 Biobrick, tctB_162 Biobrick, tctB_197 Biobrick, tphC Biobrick) and Agarose gel electrophoresis
- Colony-PCR-Screen of the transformants (Primers: tctA_505 Biobrick) and analisys of the PCR products by Agarose gel electrophoresis
- No bands visible
- Ligation of [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808004 tctA 505aa] into pSB1C3 and subsequently transformation into DH5a. No colonies
- The following vectors were sequenced: pSB1A2_197.4, pSB1A2_197.5, psSB1A2_503.1, pSB1A2_503.4, pSB1A2_162.2, pSB1C3_162.6, pSB1A2_322.12, pSB1A2_322.13
- Restriction digest of [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808004 tctA 505aa] using EcoRI/SpeI. Ligation into pSB1C3 and subsequently transformation into DH5a.
- Preperations for pure culture: colony-PCR of A2.4, A2.5, A3.1, A3.4, A4.2, C4.6, A5.12, A5.13 (2 colonies each, Primers: tctA_503 Biobrick, tctB_162 Biobrick, tctB_197 Biobrick, tphC Biobrick) and afterwards checking the PCR products by Agarose gel electrophoresis. Positive colonies were picked, streaked onto fresh media and incubated again.
- As A5.12 and A5.13 are missing the right insert the colony-PCR was repeated (5 and 6 colonies were used, Primers: tphC Biobrick). To provide additional monitoring the colony-PCR was repeated for A2.5_2, A3.1_2 und A4.2_2. 2 positive colonies were again streaked onto fresh media.
- The Arabinose-Promotor ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K808000 Ara]) was isolated from pbad_turbo_gfp and colonies containing this plasmid (3 approaches each). Approach T2 is further in use.
week 7 (02.-06.07.12)
- Restriction digest of [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808000 Ara] and pXylE using EcoRI and SpeI. Ligation of the restricted products and transformation into DH5a. No colonies
- Restriction digest of pSB1A2 using EcoRI and SpeI
- DH5a-cells containing the vectors A4.2_2, A3.1_2, A2.5_2, A5.13_5 were diluted in glycerol (20%) and freezed (-80°C).
- Mutagenic PCR of A4.2, A5.13, A3.1 to eliminate the PstI restriction site (Primers: tctA_503_PstI, tctB_162_PstI, tphC_PstI). Transformation into DH5a
- Checking the DH5a-cells for correct transformation with the vectors after the mutagenic PCR by colony-PCR (checking 7 colonies each, Primers: tctA_503 Biobrick, tctB_162 Biobrick, tphC Biobrick). The PCR products were digested with PstI. The gel was alternately loaded with digested and uncut PCR products.
- The transformation of DH5a with pSB1C3_tctA_505aa was verified by colony-PCR (Primers: tctA_505 Biobrick). The positive colonies were used for Miniprep
- Miniprep (Fermentas Kit) and Nanodrop measurement:
- A5.13m_1 : 105.2 ng/µl 260/280= 1.84
- A5.13m_2 : 223.0 ng/µl 260/280= 1.87
- A5.13m_3 : 110.6 ng/µl 260/280= 1.87
- A5.13m_4 : 99.6 ng/µl 260/280= 1.90
- A5.13m_6 : 326.6 ng/µl 260/280= 1.88
- A5.13m_7 : 145.0 ng/µl 260/280= 1.84
- A3.1m_1 : 241.7 ng/µl 260/280= 1.87
- A3.1m_2 : 188.4 ng/µl 260/280= 1.89
- A3.1m_3 : 112.4 ng/µl 260/280= 1.89
- A3.1m_4 : 173.1 ng/µl 260/280= 1.85
- A3.1m_5 : 111.6 ng/µl 260/280= 1.85
- A3.1m_6 : 138.1 ng/µl 260/280= 1.87
- A3.1m_7 : 178.9 ng/µl 260/280= 1.86
- A4.2m_1 : 57.0 ng/µl 260/280= 1.84
- A4.2m_2 : 46.5 ng/µl 260/280= 1.77
- A4.2m_3 : 55.7 ng/µl 260/280= 1.88
- A4.2m_4 : 111.3 ng/µl 260/280= 1.82
- A4.2m_5 : 236.9 ng/µl 260/280= 1.89
- A4.2m_6 : 69.3 ng/µl 260/280= 1.89
- A4.2m_7 : 72.3 ng/µl 260/280= 1.85
- C1.2 : 132.9 ng/µl 260/280= 1.72
- new transformation of DH5a with pXylE_Ara
week 8 (09.-13.07.12)
- Checking the transformation of DH5a with pSB1C3_tctA_505aa again using colony-PCR (Primers: tctA_505 Biobrick) [*]. The colonies Nr. 2, 8 and 10 were picked and streaked onto fresh media.
- colony-PCR of DH5a_pXylE_Ara (Primers: Ara_RBS, no positve colonies, data not shown)
- Sequencing the following vectors: pSB1A2_4.2m_5, pSB1A2_4.2m_4, pSB1A2_5.13m_3, pSB1A2_5.13m_2, pSB1A2_3.1m_1, pSB1A2_3.1m_5, pSB1A2_3.1m_7
- Checking the transformation of DH5a with A1 (5 colonies) and C1 (7 colonies) using colony-PCR (Primers: tctA_505 Biobrick, data not shown)
- Checking the transformation of DH5a with pSB1C3_tctA_505aa again by colony-PCR (Primers: tctA_505 Biobrick). Only the colonies Nr. 2, 8 and 10 (see above) were checked (2 colonies each).
- Another colony-PCR of DH5a_pXy19_Ara (Primers: Ara_RBS)
- Miniprep (Fermentas Kit) and Nanodrop measurement of:
- A5.13m_2 : 191.2 ng/µl 260/280= 1.84
- A5.13m_3 : 168.1 ng/µl 260/280= 1.78
- A2.5 : 193.0 ng/µl 260/280= 1.83
- A3.1m_5 : 146.1 ng/µl 260/280= 1.82
- A3.1m_1 : 250.1 ng/µl 260/280= 1.84
- A4.2m_4 : 124.8 ng/µl 260/280= 1.85
- Restriction digest of A5.13m_2, A5.13m_3, A3.1m_1, A3.1m_5, A2.5, A4.2m_4 with EcoRI and SpeI. The restriction was done to enable the ligation of the mutated genes into pSB1C3. Afterwards DH5a was transformed with the ligation product.
- Miniprep (Fermentas Kit) and Nanodrop measurement of:
- C505_2_2 : 253.6 ng/µl 260/280= 1.60
- C505_2_3 : 274.7 ng/µl 260/280= 1.88
- C505_8_2 : 462.1 ng/µl 260/280= 1.38
- C505_8_3 : 149.1 ng/µl 260/280= 1.84
- C505_10_3 : 160.4 ng/µl 260/280= 1.98
- Restriction digest of the Miniprep products with EcoRI and EcoRI/SpeI. Afterwards the restriction products were analyzed by Agarose gel electrophoresis. Gel loading: uncut, cut once, cut twice
- C505_2_3 und C505_8_2 were selected for mutagenic PCR.
- Restriction digest of pSB1C3 with EcoRI and SpeI
- Restriction digest of pXylE with EcoRI and XbaI and of [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808000 Ara] with EcoRI and SpeI. Dephosphorylation of pXylE. Ligation of pXylE and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808000 Ara]
week 9 (16.-20.07.12)
- mutagenic PCR of C505_2_3 and C505_8_2. (Primers: tctA_505_PstI, did not work, data not shown)
- [http://en.wikipedia.org/wiki/Polymerase_chain_reaction PCR] of [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808003 tct_B162m] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808005 tct_B197m] with primer to add a RBS in front and behind the gene (Primers: tctB_162_RBS, tctB_197_RBS)
- Restriction digest of the PCR products with EcoRI and SpeI for ligation into pSB1C3_5.13m (tphC)
- Adding a RBS to one site of [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808000 Ara] using [http://en.wikipedia.org/wiki/Polymerase_chain_reaction PCR] (Primers: Ara_RBS)
- Miniprep (Fermentas Kit) and Nanodrop measurement of:
- C3.1m_1 : 278.9 ng/µl 260/280= 1.88
- C4.2m_4 : 214.2 ng/µl 260/280= 1.86
- 5.13m_3 : 256.3 ng/µl 260/280= 1.88
- C2.5_2 : 44.8 ng/µl 260/280= 2.00
- C5.13m_2 : 60.8 ng/µl 260/280= 1.89
week 10 (23.-27.07.12)
- Restriction digest of pSB1C3_5.13m_3_2 and pSB1C3_5.13m_3_3 with EcoRI and XbaI. Afterwards the products are ligated with RBS_tctB162m_RBS and RBS_tctB197_RBS each.
- Restriction digest of A3.1m_1 for Ligation with Ara
- Restriction of pXylE for ligation with Ara_RBS
- [http://en.wikipedia.org/wiki/Polymerase_chain_reaction PCR] of Ara_RBS and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808000 Ara] for ligation into pXylE and pSB1C3 (Primers: Ara_RBS). Afterwards restriction digest with restriction enzymes: Ara_RBS is cut with with EcoRI and SpeI and Ara is cut with EcoRI and XbaI
- Ligation of the restricted PCR-products to: pSB1A2_tctB197_tphCm, pSB1C3_tctB162m_tphCm, pSB1C3_tct197_tphCm, pSB1A2_tctB162m_tphCm, pSB1A2_Ara_tctA503m, pSB1C3_Ara_tctA503m, pSB1A2_Ara_XylE and transformation into DH5a
- mutagenic PCR of 505_8_2 (Primers: tctA_505_PstI)
week 11 (30.07.-03.08.12)
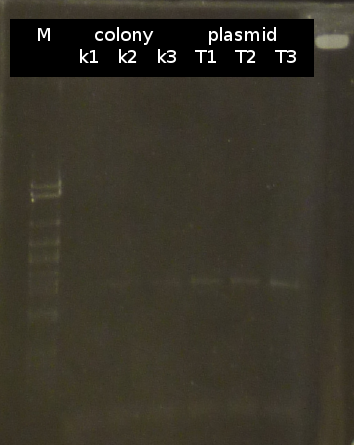
- Checking transformation of DH5a by colony-PCR: pSB1A2_tctB197_tphCm, pSB1C3_tctB162m_tphCm, pSB1C3_tct197_tphCm, pSB1A2_tctB162m_tphCm, pSB1A2_Ara_tctA503m, pSB1C3_Ara_tctA503m, pSB1A2_Ara_XylE (Primers: [http://partsregistry.org/Part:BBa_G00100 VF2], [http://partsregistry.org/Part:BBa_G00101 VR]). Positive colonies are picked and used for inoculation of fluid media.
- Miniprep (Fermentas Kit) of fluid culture and Nanodrop measurement of:
- pSB1C3_503 : 110.0 ng/µl 260/280= 1.82
- pSB1A2_tctB197_tphCm : 32.0 ng/µl 260/280= 2.10
- pSB1A2_Ara_tctA503m : 202.0 ng/µl 260/280= 1.78
- pSB1A2_tctB162m_tphCm : 36.5 ng/µl 260/280= 1.91
- pSB1C3_Ara_tctA503m : 133.2 ng/µl 260/280= 1.83
- pSB1C3_tctB162m_tphCm : 35.2 ng/µl 260/280= 1.93
- pSB1C3_tct197_tphC : 30.6 ng/µl 260/280= 1.88
- Restriction digest of pSB1A2_Ara_tctA503m, pSB1C3_Ara_tctA503m, pSB1A2_tctB162m_tphCm and pSB1C3_tctB162m_tphCm and building the final constructs by ligation: pSB1A2_tctA503m_tctB162m_tphCm und pSB1C3_tctA503m_tctB162m_tphCm. Transformation of the final constructs into DH5a.
- Transformation of pSB1C3_503 into DH5a
- Checking the mutagenic PCR product of 505_8_2 and the transformation of DH5a with pSB1C3_503 by [http://en.wikipedia.org/wiki/Polymerase_chain_reaction PCR] (Primers: tctA_505 Biobrick, tctA_503 Biobrick)
- Restriction digest of [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808000 Ara] und C4.2_4_1 using EcoRI and PstI, building pSB1C3_Ara by ligation and transformation of the vector into DH5a
- Miniprep (Fermentas Kit) and Nanodrop measurement of:
- pSB1C3_Ara : 86.7 ng/µl 260/280= 1.83
- pSB1C3_505m : 113.8 ng/µl 260/280= 1.85
- Restriction digest of pSB1C3_505m with PstI. Checking if the restriction site has been successfully removed
- restriction site hasn't been removed
- Screnning for transformation of DH5a with pSB1A2_tctA503m_tctB162m_tphCm and pSB1C3_tctA503m_tctB162m_tphCm using colony-PCR (Primers: [http://partsregistry.org/Part:BBa_G00100 VF2], [http://partsregistry.org/Part:BBa_G00101 VR]) [*]
- Checking the constructs pSB1A2/pSB1C3_Ara_503m and pSB1A2/pSB1C3_tctB162m_tphCm by [http://en.wikipedia.org/wiki/Polymerase_chain_reaction PCR] (Primers: [http://partsregistry.org/Part:BBa_G00100 VF2], [http://partsregistry.org/Part:BBa_G00101 VR]) [*]
- Ligation of [http://partsregistry.org/wiki/index.php?title=Part:BBa_K808000 Ara] into pSB1C3
- Checking if pSB1A2_tctB162m_tphCm and pSB1C3_tctB162m_tphCm have the right Prefix/Suffix by digesting both with EcoRI and XbaI and with EcoRI and PstI [*]
- Ligation of pSB1A2_tctB162m_tphCm and pSB1C3_tctB162m_tphCm with Ara_503
- Miniprep (Fermentas Kit) and Nanodrop measurement of:
- pSB1A2_tctB162m_tphCm_1 : 45.3 ng/µl 260/280= 1.90
- pSB1A2_tctB162m_tphCm_2 : 74.5 ng/µl 260/280= 1.82
- pSB1C3_tctB162m_tphCm_1 : 95.8 ng/µl 260/280= 1.86
- pSB1C3_tctB162m_tphCm_2 : 85.7 ng/µl 260/280= 1.81
- pSB1C3_tctB162m_tphCm_3 : 100.1 ng/µl 260/280= 1.84
- pSB1C3_tctB162m_tphCm_4 : 79.9 ng/µl 260/280= 1.88
- pSB1C3_tctB162m_tphCm_5 : 89.8 ng/µl 260/280= 1.85
- Sequencing pSB1A2_tctB162m_tphCm_2 and pSB1C3_tctB162m_tphCm_5
week 12 (06.-10.08.12)
- Miniprep (Fermentas Kit) and Nanodrop measurement of:
- pSB1A2_tctB162m_tphCm_2 : 55.7 ng/µl 260/280= 1.81
- pSB1C3_tctB162m_tphCm_5 : 101.2 ng/µl 260/280= 1.92
- Mutagenic PCR of pSB1C3_505_8_2 (Primers: tctA_505_PstI) and subsequent transformation into DH5a
- Ligation of Ara_503m with pSB1A2_tctB162m_tphCm/pSB1C3_tctB162m_tphCm and subsequent transformation into DH5a. Screening the transformants using colony-PCR (Primers: [http://partsregistry.org/Part:BBa_G00100 VF2], [http://partsregistry.org/Part:BBa_G00101 VR])
- only colony 1, 2 and 3 have the right insert
- Miniprep (Fermentas Kit) and Nanodrop measurement of:
- pSB1A2_Ara_tctA503m : 278.0 ng/µl 260/280= 1.86
- pSB1C3_Ara_tctA503m : 169.0 ng/µl 260/280= 1.87
- pSB1C3_Ara_1 : 59.0 ng/µl 260/280= 1.90
- pSB1C3_Ara_4 : 55.5 ng/µl 260/280= 1.80
- pSB1C3_Ara_tctA503m_tctB162m_tphCm : 188.8 ng/µl 260/280= 1.86
- pSB1C3_Ara_tctA503m_tctB162m_tphCm :168.8 ng/µl 260/280= 1.84
- pSB1C3_Ara_tctA503m_tctB162m_tphCm :271.3 ng/µl 260/280= 1.85
- pSB1C3_Ara_XylE : 149.8 ng/µl 260/280= 1.86
- pSB1C3_tctB197_tphC : 95.8 ng/µl 260/280= 1.84
- pSB1A2_tctB197_tphC : 69.0 ng/µl 260/280= 1.88
- Sequencing of 3 different pSB1C3_Ara_tctA503m_tctB162m_tphC
- Analysis of mutagenic PCR products of 505_8_2 and restriction digest with PstI
- Restriction digest of Ara_RBS with EcoRI and SpeI and of pSB1C3_505m with EcoRI and XbaI. Building pSB1C3_Ara_505m by ligation and subsequent transformation into DH5a.
week 13 (13.-17.08.12)
- Restriction digest of pSB1C3_197_tphCm with EcoRI and XbaI
- Checking the transformation of DH5a with pSB1C3_Ara_505m by using colony-PCR (Primers: [http://partsregistry.org/Part:BBa_G00100 VF2], [http://partsregistry.org/Part:BBa_G00101 VR])
- Sequencing pSB1C3_Ara_XylE, pSB1C3_Ara_tctA503m_tctB162m_tphCm (=Operon1), pSB1C3_505m
- Restriction digest of Ara_RBS with EcoRI and SpeI and of pSB1C3_tphCm with EcoRI and XbaI. Building pSB1C3_Ara_tphCm by ligation and subsequent transformation into DH5a
- Restriction digest of pSB1C3_Ara_tctA505m with EcoRI and SpeI. Ligation of Ara_tctA505m and tctB197_tphCm into pSB1C3
week 14 (20.-24.08.12)
- Checking the transformation of DH5a with pSB1C3_Ara_tphCm using colony-PCR (Primers: [http://partsregistry.org/Part:BBa_G00100 VF2], [http://partsregistry.org/Part:BBa_G00101 VR])
Week 15-19 charakterisation and uptake tests
- Restriction digest of pSB1C3_Ara_tphCm and pSB1C3_Ara_tctA505m with PstI and SpeI
- Restriction digest of pSB1C3_197_tphCm with XbaI and PstI.
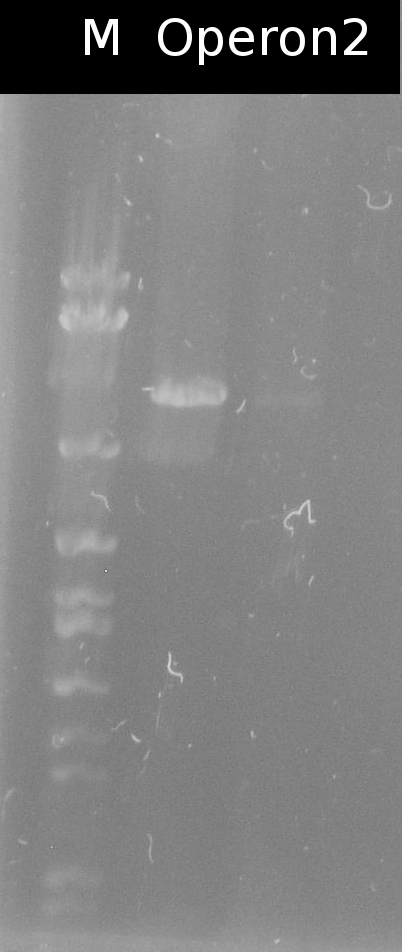
- Building the construct pSB1C3_Ara_tctA505m_197_tphCm (= Operon2) by ligation. Subsequent transformation into DH5a
- Checking the transformation of DH5a with Operon2 using colony-PCR (Primers: [http://partsregistry.org/Part:BBa_G00100 VF2], [http://partsregistry.org/Part:BBa_G00101 VR])
- Charakterisation of AraC-Pbad
(link zu dem biobrick) was done. (link zu dem part und protocol)?
- Photometric measurement of TPA uptake in M9 minimal media at 240 nm. (did not work)
- Preparation of E. coli strain C43(DE3) carrying putative terephtalate Operon1 ( BBa_K808006 ), putative terephtalate Operon2 (BBa_K808007 ) and AraC-Pbad plasmids (BBa_K808000 ) Gas chromatography–mass spectrometry (GC-MS)
Three different overnight culture of Escherichia Coli strain C43(DE3) carrying putative terephtalate Operon1 ( BBa_K808006 ), putative terephtalate Operon2 (BBa_K808007 ) and AraC-Pbad plasmids (BBa_K808000 ) was inoculated 1:100 into fresh 100 mL LB media. 0,001% L-(+)-Arabinose (inducer) was added at the optical density (OD600) of 1.3-1.4 and incubated at 30°C for 2 hour, shaking at 230 rpm. Afterwards 2 mL of each cultures was transferred to a 2 mL Epi with each 50,-100,-150, -200,-250,-300,- µM TPA and incubated on a heating-block at 30 °C, shaked at 1000 rpm for 5 hours. After the incubation the cultures were centrifuged for 15 min at 17.000xg. 100 µM cultures were chosed for GC-MC measurment.The supernatan was transferred into a new Eppi and the protocol sample preparation for GC-MS was accomplished.
- Preparation of Escherichia Coli for the energy dispersive X-ray spectroscopy (EDX).
For the EDX measument the cells have to be fixed, embedded in a SPURR-embedding medium (resin) and cut in to small rectangles smaller than 100 nm. For more details: EDX Preparation of Escherichia Coli for the energy dispersive X-ray spectroscopy (EDX) The embedded E. Colis was checked under a electron microscope (Fig..yyy)
Figure xxyx. Electron microscope picture of embedded E. Colis carrying putative terephtalate Operon1 ( BBa_K808006 ), putative terephtalate Operon2 (BBa_K808007 ) and AraC-Pbad plasmids (BBa_K808000 ).
Preparation of Escherichia Coli strain C43(DE3) carrying putative terephtalate Operon1 ( BBa_K808006 ), putative terephtalate Operon2 (BBa_K808007 ) and AraC-Pbad plasmids (BBa_K808000 ) Gas chromatography–mass spectrometry (GC-MS)
Three different overnight culture of Escherichia Coli strain C43(DE3) carrying putative terephtalate Operon1 ( BBa_K808006 ), putative terephtalate Operon2 (BBa_K808007 ) and AraC-Pbad plasmids (BBa_K808000 ) was inoculated 1:100 into fresh 100 mL LB media. 0,001% L-(+)-Arabinose (inducer) was added at the optical density (OD600) of 1,3-1,4 and incubated at 30°C for 2 hour, shaking at 230 rpm. Afterwards 2 mL of each cultures was transferred to a 2 mL Epi with each 50,-100,-150, -200,-250,-300,- µM TPA and incubated on a heating-block at 30 °C, shaked at 1000 rpm for 5 hours. After the incubation the cultures were centrifuged for 15 min at 17.000xg. 100 µM cultures were chosed for GC-MC measurment.The supernatan was transferred into a new Eppi and the protocol sample preparation for GC-MS was accomplished.
Preparation of Escherichia Coli for the energy dispersive X-ray spectroscopy (EDX)
For the EDX measument the cells have to be fixed, embedded in a SPURR-embedding medium (resin) and cut in to small rectangles smaller than 100 nm. For more details: EDX Preparation of Escherichia Coli for the energy dispersive X-ray spectroscopy (EDX) The embedded E. Colis was checked under a electron microscope (Fig..yyy)
 "
"