Team:UNAM Genomics Mexico/Results
From 2012.igem.org
| (29 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{:Template:Team:UNAM_Genomics_Mexico/webhtml| content= | {{:Template:Team:UNAM_Genomics_Mexico/webhtml| content= | ||
| + | __NOTOC__ | ||
| + | <br /> | ||
| + | <center><h1>'''GENERAL RESULTS'''</h1></center><br /> | ||
<br /> | <br /> | ||
| - | + | For the heavy metal AND we sent to the registry four Biological Parts: CI, and the three different ArsR and CzrA metal sensing repressors binding fused promoters (97, 98, and 99). We did not design promoter 98 (iGEM Newcastle 2009 did), however the part was not in the registry. <br /><br /> | |
| - | + | The constructs we have up to now are: <br /><br /> | |
| - | For the heavy metal AND we sent to the registry four Biological Parts: CI, and the three different | + | <center> |
| - | The constructs we have up to now are: <br /> | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
<table border="0" height="150" cellspacing="15" bgcolor="transparent" id="tablecontentbg" cellpadding="10"> | <table border="0" height="150" cellspacing="15" bgcolor="transparent" id="tablecontentbg" cellpadding="10"> | ||
<tr> | <tr> | ||
| - | <td id=" | + | <td id="rightcolumn2" align= "center"><br />[[ File:UnamgenomicsConstruccion1.png | 870px]]<br /><br /> <p>''' AmyE 5' +CzrA/ArsR (97)+ RBS P4 + RBS CI + RFP + TERMINATOR + Sp/Smr + TERMINATOR + AmyE 3' One complete AND'''</p></td> |
| + | </tr> | ||
| + | |||
| + | <tr> | ||
| + | <td id="rightcolumn2" align= "center"><br />[[File:UnamgenomicsAndmetals1.png | x90px]]<br /><br /> <p>'''AmyE 5' +CzrA/ArsR (98 and 99)+ RBS P4 + RBS CI + RFP + TERMINATOR'''</p></td> | ||
</tr> | </tr> | ||
<tr> | <tr> | ||
| - | |||
</tr> | </tr> | ||
</table> | </table> | ||
| + | </center> | ||
| + | AmyE 5' +CzrA/ArsR (97)+ RBS P4 + RBS CI + RFP + TERMINATOR + Sp/Smr + TERMINATOR + AmyE 3' <br /><br /> | ||
| + | also, | ||
| + | AmyE 5' +CzrA/ArsR (98-99)+ RBS P4 + RBS CI + RFP + TERMINATOR<br /><br /> | ||
| + | |||
| + | We still need to complete it with Sp/Smr + TERMINATOR + AmyE 3' (a part which we already have), do the same thing for promoter 99, and to add : <br /><br /> | ||
| + | |||
| + | <center> | ||
<table border="0" height="150" cellspacing="15" bgcolor="transparent" id="tablecontentbg" cellpadding="10"> | <table border="0" height="150" cellspacing="15" bgcolor="transparent" id="tablecontentbg" cellpadding="10"> | ||
<tr> | <tr> | ||
| - | <td id=" | + | <td id="contentcolumn" align= "center"><br />[[File:UnamgenomicsresultsOr3.png |x100px]] <p>'''Sp/Smr + TERMINATOR + AmyE 3''''</p></td> |
| + | </tr> | ||
| + | </table> | ||
| + | </center> | ||
| + | AmyE 5' +CzrA/ArsR (98)+ RBS LasR + RBS CI to RFP + TERMINATOR and Sp/Smr + TERMINATOR + AmyE 3' as well. | ||
| + | We are also trying to ligate our fused promoters to GFP in order to characterize them. <br /><br /> | ||
| + | For the arabinose-xylose AND gate we sent to the registry the omega cassette, pBad/pXyl, pVeg, and XylR. | ||
| + | (XylR already existed in the registry, but ours is optimized for Bacillus subtilis). We characterized the omega cassette. <br /><br /> | ||
| + | The constructs we have up to now are:<br /><br /> | ||
| + | |||
| + | <center> | ||
| + | <table border="0" height="150" cellspacing="15" bgcolor="transparent" id="tablecontentbg" cellpadding="10"> | ||
| + | <tr> | ||
| + | <td id="leftcolumn3" align= "center"><br />[[File:UnAMgenomicsResultados andsugartbueno.png| 870px]] <p>'''AmyE 5' + pBad/pXyl +RBS + P4 +RBS + CI + RBS + RFP + TT''''</p></td> | ||
</tr> | </tr> | ||
</table> | </table> | ||
| + | </center> | ||
| - | + | AmyE 5' + pBad/pXyl +RBS + P4 +RBS + CI + RBS + RFP + TT<br /><br /> | |
| - | + | ||
| - | AmyE 5' + | + | |
| - | + | <center> | |
| + | <table border="0" height="150" cellspacing="15" bgcolor="transparent" id="tablecontentbg" cellpadding="10"> | ||
| + | <tr> | ||
| + | <td id="leftcolumn3" align= "center"><br />[[File:UnamgenomcismexicoArac omega.png| 870px]] <p>'''RBS AraC + omega cassette + AmyE 3''''</p></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | </center> | ||
| - | |||
| - | |||
| - | + | RBS AraC + omega cassette + AmyE 3'<br /><br /> | |
| + | <center> | ||
| + | <table border="0" height="150" cellspacing="15" bgcolor="transparent" id="tablecontentbg" cellpadding="10"> | ||
| + | <tr> | ||
| + | <td id="leftcolumn3" align= "center"><br />[[File:UnamgenomicsSweetand 2.png| 870px]] <p>'''Pveg + RBS XylR'''</p></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | </center> | ||
| - | |||
| - | |||
| - | |||
| - | + | Pveg + RBS XylR (we are working hard to obtain the final construct which joins these tree parts.<br /><br /> | |
| - | For the A3-LasB ( | + | For the A3-LasB (GFP reporter) OR gate we sent to the registry A3.<br /><br /> |
| - | + | ||
| - | + | ||
| - | + | ||
| - | [[File: | + | We have:<br /><br /> |
| + | <center> | ||
| + | <table border="0" height="150" cellspacing="15" bgcolor="transparent" id="tablecontentbg" cellpadding="10"> | ||
| + | <tr> | ||
| + | <td id="contentcolumn" align= "center"><br />[[File:UnamgenomicsmexicoOrtenemos.png | 870px]] <p>'''AmyE 5' + LasB + RBS + GFP + TT + A3 and need to ligate it with RBS + GFP + Terminator + omega cassette terminator + AmyE 3''''</p></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | </center> | ||
| - | |||
| + | AmyE 5' + LasB + RBS + GFP + TT + A3 and need to ligate it with RBS + GFP + Terminator + omega cassette terminator + AmyE 3' which we also already have. <br /><br /> | ||
| - | Regarding our work with Bacillus subtilis, we designed a new standard for working with it. This way future iGEM teams that wish to work with Bacillus subtilis will not have trouble transforming it. We designed a series of videos to explain how this is done. <br /> | + | |
| - | [[Team:UNAM_Genomics_Mexico/Results/ | + | == '''''Bacillus subtilis''''' == |
| + | <br /> | ||
| + | Regarding our work with ''Bacillus subtilis'', we designed a new standard protocol for working with it. | ||
| + | This way future iGEM teams that wish to work with Bacillus subtilis will not have trouble transforming it. | ||
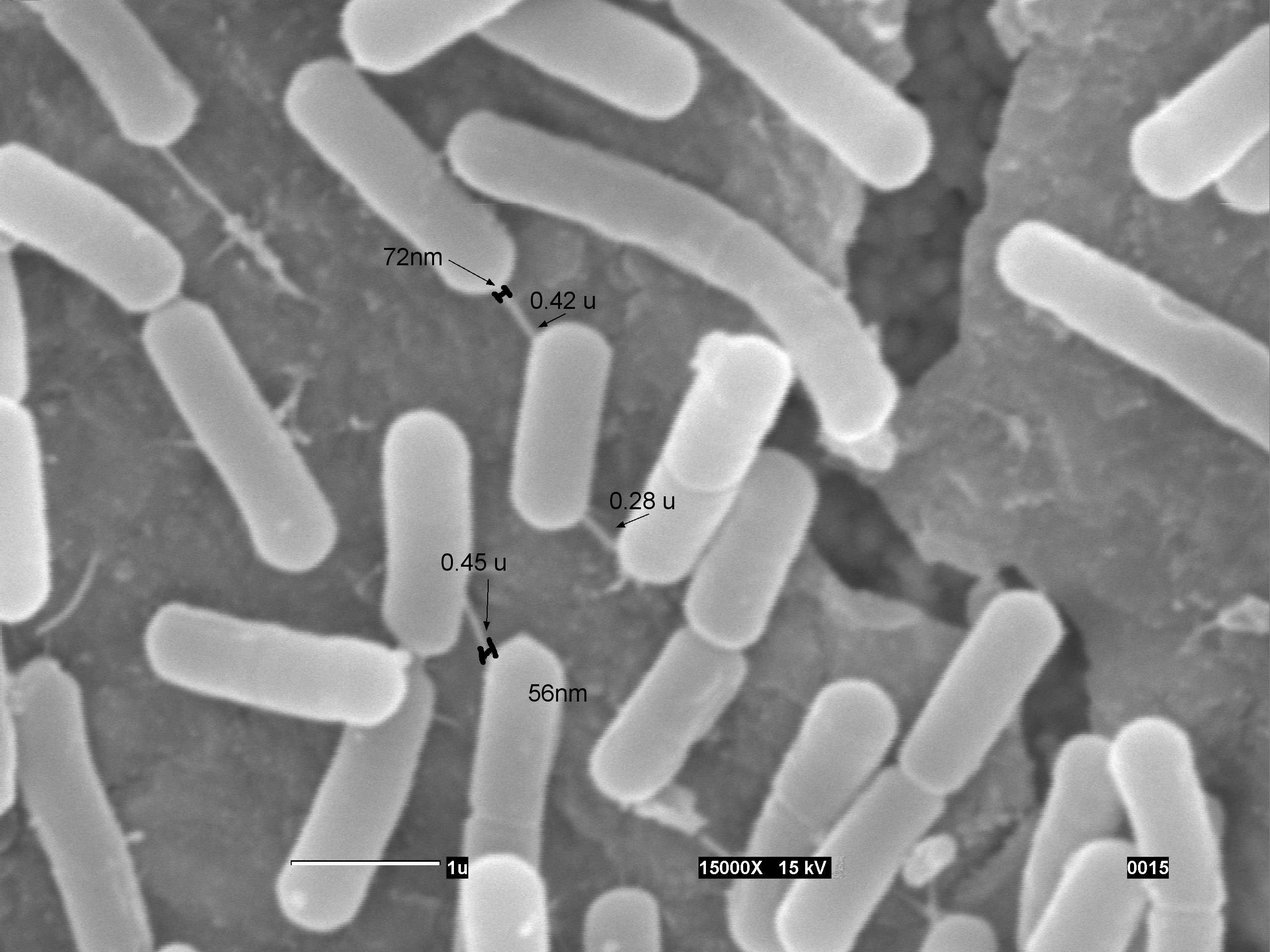
| + | We designed a series of videos to explain how this is done [[Team:UNAM_Genomics_Mexico/Results/Bacillus | Video with the protocol ]]; also PY79 cells were grown to midexponential phase, plated on LB agar, incubated for 6 hr at 37°C, and visualized by SEM, in which we were able to observe the NANOTUBES, just as Ben Yehuda et. al. reported. | ||
| + | <br /><br /> | ||
| + | ::::::::::::::::::::::::::::::::::::::[[File:UnamgenomcisUp.png|right | 120px |link=Team:UNAM_Genomics_Mexico/Results#GENERAL_RESULTS]] | ||
| + | |||
| + | <center> | ||
| + | <table border="0" height="150" cellspacing="15" bgcolor="transparent" id="tablecontentbg" cellpadding="10"> | ||
| + | <tr> | ||
| + | <td id="rightcolumn2" align= "center"><br />[[File:UnamgenomicsREJ35 valores.jpg|800px| link=Team:UNAM_Genomics_Mexico/Notebook/Bacillus]]<p>'''PY79 cells were grown to midexponential phase, plated on LB agar, incubated for 6 hr at 37°C, and visualized by SEM Rej35_valores:(A)Measurment of the length and width of the nanotubes connecting the (PY79) cells grown on solid LB medium. The scale bar represent 1 micrometer'''</p></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | </center> | ||
| + | <br /><br /> | ||
Latest revision as of 00:20, 26 October 2012

GENERAL RESULTS
For the heavy metal AND we sent to the registry four Biological Parts: CI, and the three different ArsR and CzrA metal sensing repressors binding fused promoters (97, 98, and 99). We did not design promoter 98 (iGEM Newcastle 2009 did), however the part was not in the registry.
The constructs we have up to now are:
 AmyE 5' +CzrA/ArsR (97)+ RBS P4 + RBS CI + RFP + TERMINATOR + Sp/Smr + TERMINATOR + AmyE 3' One complete AND |
 AmyE 5' +CzrA/ArsR (98 and 99)+ RBS P4 + RBS CI + RFP + TERMINATOR |
AmyE 5' +CzrA/ArsR (97)+ RBS P4 + RBS CI + RFP + TERMINATOR + Sp/Smr + TERMINATOR + AmyE 3'
also,
AmyE 5' +CzrA/ArsR (98-99)+ RBS P4 + RBS CI + RFP + TERMINATOR
We still need to complete it with Sp/Smr + TERMINATOR + AmyE 3' (a part which we already have), do the same thing for promoter 99, and to add :
 Sp/Smr + TERMINATOR + AmyE 3' |
AmyE 5' +CzrA/ArsR (98)+ RBS LasR + RBS CI to RFP + TERMINATOR and Sp/Smr + TERMINATOR + AmyE 3' as well.
We are also trying to ligate our fused promoters to GFP in order to characterize them.
For the arabinose-xylose AND gate we sent to the registry the omega cassette, pBad/pXyl, pVeg, and XylR.
(XylR already existed in the registry, but ours is optimized for Bacillus subtilis). We characterized the omega cassette.
The constructs we have up to now are:
 AmyE 5' + pBad/pXyl +RBS + P4 +RBS + CI + RBS + RFP + TT' |
AmyE 5' + pBad/pXyl +RBS + P4 +RBS + CI + RBS + RFP + TT
 RBS AraC + omega cassette + AmyE 3' |
RBS AraC + omega cassette + AmyE 3'
 Pveg + RBS XylR |
Pveg + RBS XylR (we are working hard to obtain the final construct which joins these tree parts.
For the A3-LasB (GFP reporter) OR gate we sent to the registry A3.
We have:
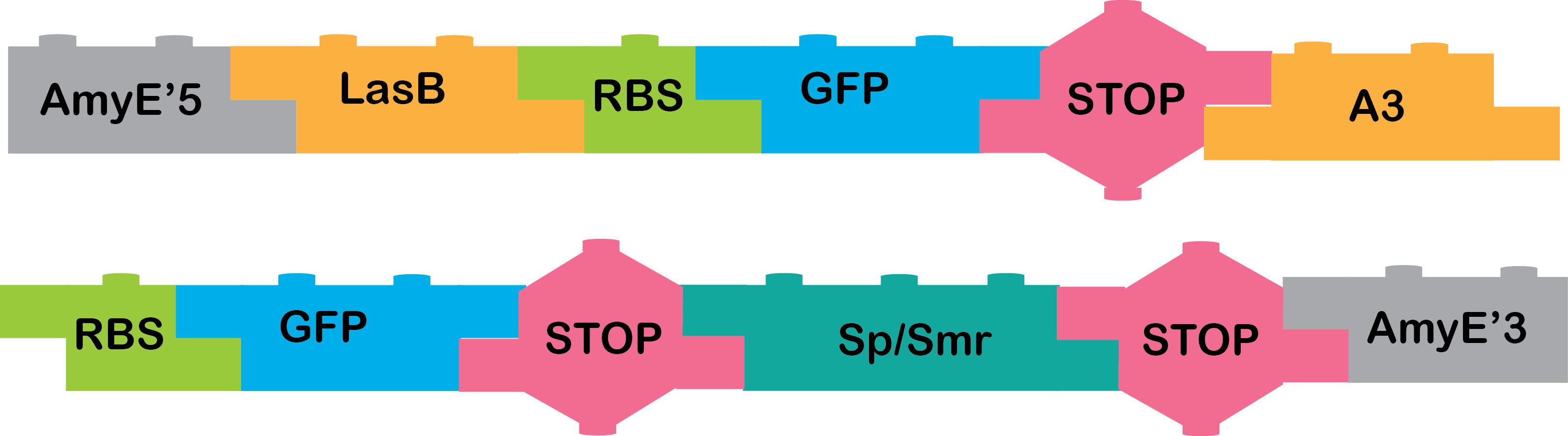 AmyE 5' + LasB + RBS + GFP + TT + A3 and need to ligate it with RBS + GFP + Terminator + omega cassette terminator + AmyE 3' |
AmyE 5' + LasB + RBS + GFP + TT + A3 and need to ligate it with RBS + GFP + Terminator + omega cassette terminator + AmyE 3' which we also already have.
Bacillus subtilis
Regarding our work with Bacillus subtilis, we designed a new standard protocol for working with it.
This way future iGEM teams that wish to work with Bacillus subtilis will not have trouble transforming it.
We designed a series of videos to explain how this is done Video with the protocol ; also PY79 cells were grown to midexponential phase, plated on LB agar, incubated for 6 hr at 37°C, and visualized by SEM, in which we were able to observe the NANOTUBES, just as Ben Yehuda et. al. reported.
 "
"