Team:Groningen/Modeling
From 2012.igem.org
m |
|||
| Line 1: | Line 1: | ||
{{HeaderGroningen2012}} | {{HeaderGroningen2012}} | ||
| + | <br> | ||
| + | <br> | ||
| + | == Overview == | ||
| + | The modeling portion of the project focuses on three major initiatives: | ||
| + | # Estimation of the allowed initial medium composition which would: | ||
| + | ## Germinate the spores | ||
| + | ## Raise the population to reporter-required levels within the smallest possible spoilage time-frame | ||
| + | ## Maintaining an active population at that level for a prolonged period of time, avoiding the onset of dormancy or sporulation | ||
| + | # The creation of a dynamic model for ''B. subtilis'' suitable for flux balance analysis which responds to environmental cues. | ||
| + | # The creation of a succinct explanation of TnrA activation, according to current literature. | ||
| + | |||
| + | == Initial Medium Composition == | ||
| + | === Purpose === | ||
| + | The ''B. subtilis'' within the containment unit must go through distinct phases under defined time constraints. The only way to control the amount of biomass, and its behavior, is through the initial medium concentration. The containment unit does not contain any time-release capsules for providing a controlled level of nutrients. | ||
| + | |||
| + | === Result === | ||
| + | |||
| + | == Dynamic Model == | ||
| + | === Purpose === | ||
| + | For our purposes, the dynamic model seeks to provide an explanation for the gene expression observed in the microarray experiment. However, for the scientific community in general, a dynamic model would be enable phenotype prediction over time under varying environmental conditions. Such a model would also be able to predict phenotypes for knock-out mutant strains. | ||
| + | |||
| + | Probabilistic integrative modeling (PROM) applies gene expression data and environmental conditions to a transcriptional-regulatory network to constrain the fluxes through a stoichiometric model suitable for flux balance analysis. This method has been shown to accurately predict growth phenotypes in ''E. coli'' and in knock-out mutants of ''M. tuberculosis''. | ||
| + | |||
| + | === Result === | ||
| + | |||
| + | == TnrA Activation == | ||
| + | === Purpose === | ||
| + | Prior to the successful microarray experiment conducted by the wetwork team, the proposed volatile sensing mechanism tied reporter activation to the metabolism of ammonium/ammonia (NH4/NH3+). The nitrogen metabolism in ''B. subtilis'' is a convoluted mesh of reactions and (in)activation complexes mostly controlled by the TnrA transcription factor. In order to observe the effect of NH4 uptake on the TnrA, it was necessary to have a concise behavioral diagram. Unfortunately, no such diagram existed. The diagram for nitrogen metabolism on the KEGG database identified most of what was involved, but did so in an unclear manner. | ||
| + | |||
| + | === Result === | ||
| + | |||
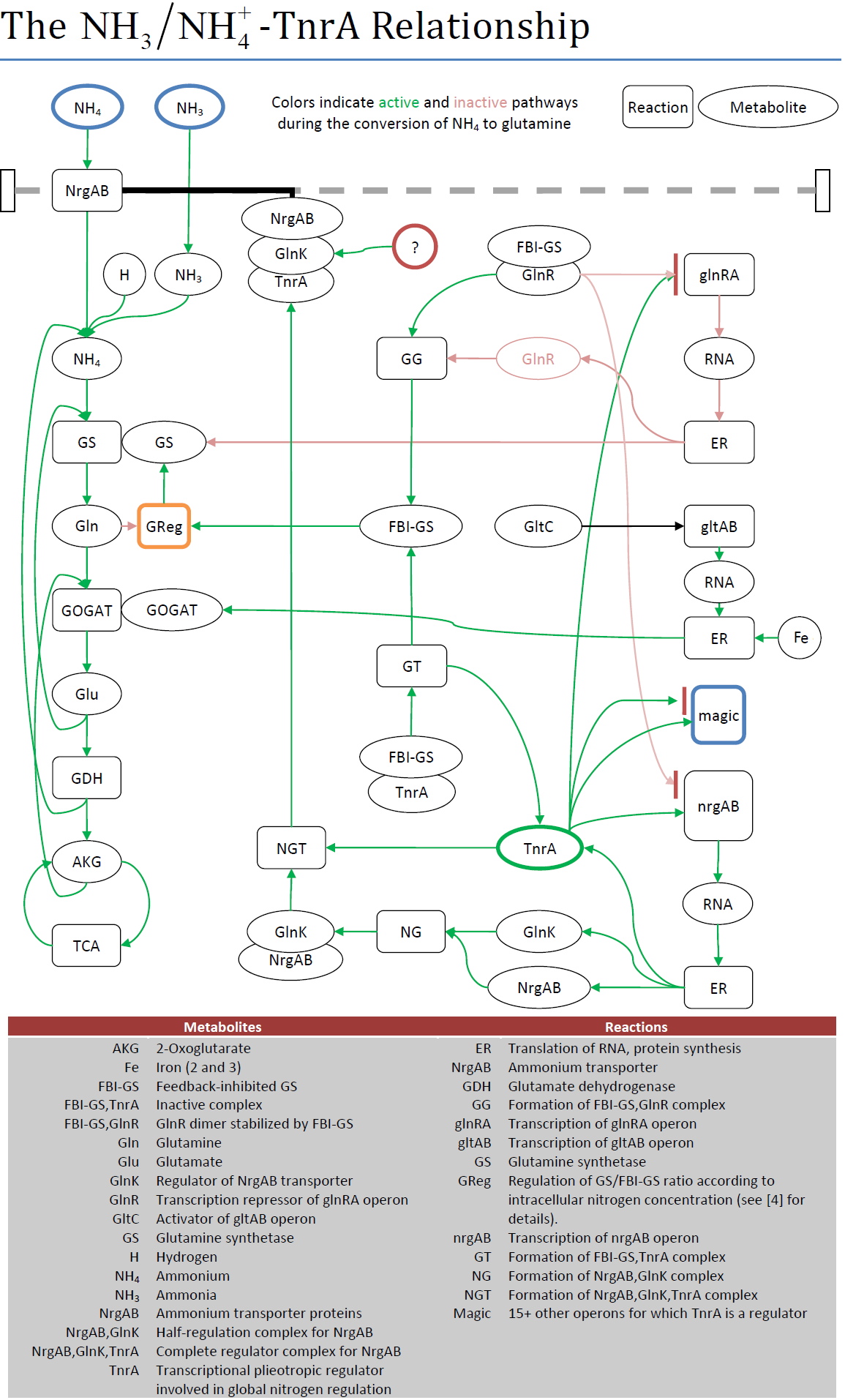
| + | The figure below used the behavioral information from literature to highlight the active and inactive pathways during ammonium uptake. In terms of the project, the creation of this diagram brought to light a critical problem for using TnrA to sense extracellular NH4/NH3+; TnrA is only active when glutamine synthetase (GS) is actively converting NH4 to glutamine. This in turn is regulated by cell growth, not the amount of NH4/NH3+ present. However, in one article it was observed that GS is also active under conditions of nitrogen limitation. If the medium contained the precise amount of glutamine necessary to support the initial growth phase, then the glutamine would be depleted at the sensing phase. GS would activate to try and create more glutamine. GS would only deactivate when the amount of extracellular NH4/NH3+ reached a level to remove the lack of nitrogen as factor limiting further growth. | ||
| + | |||
| + | The real result? If this sensing pathway was selected, then the wetwork team would have to find the necessary levels of glutamine to exploit the edge of nitrogen limitation. Possible, but not exactly the easiest mechanism to implement for the consumer market. | ||
{|align="center" | {|align="center" | ||
| Line 9: | Line 43: | ||
|} | |} | ||
| - | References | + | === References === |
| + | # KEGG Database, "Nitrogen Metabolism - Bacillus subtilis", Kanehisa Laboratories, Last Modified: July 23, 2010. http://www.kegg.jp/kegg-bin/show_pathway?org_name=bsu&mapno=00910 | ||
# SubtiWiki, http://subtiwiki.uni-goettingen.de/wiki/index.php | # SubtiWiki, http://subtiwiki.uni-goettingen.de/wiki/index.php | ||
# K. Gunka, F.M. Commichau, “Control of glutamate homeostasis in Bacillus subtilis: a complex interplay between ammonium assimilation, glutamate biosynthesis and degradation,” Molecular Biology, under review (2012). | # K. Gunka, F.M. Commichau, “Control of glutamate homeostasis in Bacillus subtilis: a complex interplay between ammonium assimilation, glutamate biosynthesis and degradation,” Molecular Biology, under review (2012). | ||
# N.A. Doroshchuk, M.S. Gelfand, D.A. Rodlanov, “Regulation of Nitrogen Metabolism in Gram-Positive Bacteria,” Molecular Biology, vol. 40(5), pp. 829-836, (2006). | # N.A. Doroshchuk, M.S. Gelfand, D.A. Rodlanov, “Regulation of Nitrogen Metabolism in Gram-Positive Bacteria,” Molecular Biology, vol. 40(5), pp. 829-836, (2006). | ||
# Study Guide, Chem153C, University of California, Los Angeles. http://vohweb.chem.ucla.edu/voh/classes%5Cspring10%5C153CID28%5C11AminoAcidBiosynthesisSQA.pdf | # Study Guide, Chem153C, University of California, Los Angeles. http://vohweb.chem.ucla.edu/voh/classes%5Cspring10%5C153CID28%5C11AminoAcidBiosynthesisSQA.pdf | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
Revision as of 10:09, 3 August 2012








Contents |
Overview
The modeling portion of the project focuses on three major initiatives:
- Estimation of the allowed initial medium composition which would:
- Germinate the spores
- Raise the population to reporter-required levels within the smallest possible spoilage time-frame
- Maintaining an active population at that level for a prolonged period of time, avoiding the onset of dormancy or sporulation
- The creation of a dynamic model for B. subtilis suitable for flux balance analysis which responds to environmental cues.
- The creation of a succinct explanation of TnrA activation, according to current literature.
Initial Medium Composition
Purpose
The B. subtilis within the containment unit must go through distinct phases under defined time constraints. The only way to control the amount of biomass, and its behavior, is through the initial medium concentration. The containment unit does not contain any time-release capsules for providing a controlled level of nutrients.
Result
Dynamic Model
Purpose
For our purposes, the dynamic model seeks to provide an explanation for the gene expression observed in the microarray experiment. However, for the scientific community in general, a dynamic model would be enable phenotype prediction over time under varying environmental conditions. Such a model would also be able to predict phenotypes for knock-out mutant strains.
Probabilistic integrative modeling (PROM) applies gene expression data and environmental conditions to a transcriptional-regulatory network to constrain the fluxes through a stoichiometric model suitable for flux balance analysis. This method has been shown to accurately predict growth phenotypes in E. coli and in knock-out mutants of M. tuberculosis.
Result
TnrA Activation
Purpose
Prior to the successful microarray experiment conducted by the wetwork team, the proposed volatile sensing mechanism tied reporter activation to the metabolism of ammonium/ammonia (NH4/NH3+). The nitrogen metabolism in B. subtilis is a convoluted mesh of reactions and (in)activation complexes mostly controlled by the TnrA transcription factor. In order to observe the effect of NH4 uptake on the TnrA, it was necessary to have a concise behavioral diagram. Unfortunately, no such diagram existed. The diagram for nitrogen metabolism on the KEGG database identified most of what was involved, but did so in an unclear manner.
Result
The figure below used the behavioral information from literature to highlight the active and inactive pathways during ammonium uptake. In terms of the project, the creation of this diagram brought to light a critical problem for using TnrA to sense extracellular NH4/NH3+; TnrA is only active when glutamine synthetase (GS) is actively converting NH4 to glutamine. This in turn is regulated by cell growth, not the amount of NH4/NH3+ present. However, in one article it was observed that GS is also active under conditions of nitrogen limitation. If the medium contained the precise amount of glutamine necessary to support the initial growth phase, then the glutamine would be depleted at the sensing phase. GS would activate to try and create more glutamine. GS would only deactivate when the amount of extracellular NH4/NH3+ reached a level to remove the lack of nitrogen as factor limiting further growth.
The real result? If this sensing pathway was selected, then the wetwork team would have to find the necessary levels of glutamine to exploit the edge of nitrogen limitation. Possible, but not exactly the easiest mechanism to implement for the consumer market.
| Fig. m1. Reactions involved between ammonium uptake and TnrA. |
References
- KEGG Database, "Nitrogen Metabolism - Bacillus subtilis", Kanehisa Laboratories, Last Modified: July 23, 2010. http://www.kegg.jp/kegg-bin/show_pathway?org_name=bsu&mapno=00910
- SubtiWiki, http://subtiwiki.uni-goettingen.de/wiki/index.php
- K. Gunka, F.M. Commichau, “Control of glutamate homeostasis in Bacillus subtilis: a complex interplay between ammonium assimilation, glutamate biosynthesis and degradation,” Molecular Biology, under review (2012).
- N.A. Doroshchuk, M.S. Gelfand, D.A. Rodlanov, “Regulation of Nitrogen Metabolism in Gram-Positive Bacteria,” Molecular Biology, vol. 40(5), pp. 829-836, (2006).
- Study Guide, Chem153C, University of California, Los Angeles. http://vohweb.chem.ucla.edu/voh/classes%5Cspring10%5C153CID28%5C11AminoAcidBiosynthesisSQA.pdf
 "
"