Team:University College London/LabBook/Week13
From 2012.igem.org



Contents |
Monday 03/09
Aim:
To repeat the DNAse/Nuclease Halo experiment in order to check whether results are repeatable. This included collecting data of halo diameters and colony diameters over 28 hour, and to determine how the nuclease production increases overtime.
Methods:
In order to be able to characterise cells, the following protocol was used:
1. Prepare 11-16 plates (10ml DNAse - Agar + 10ul AMP + 10ul 1M IPTG). IPTG induces the lac promoter which in turn activates the transcription of nuclease.
2. Streak cells onto all plates at the same time
3. Incubate at 37°C
4. Apply hydrochloric acid (HCl) to the first plate before putting in the incubator (set time as zero)
5. Take a second reading after four hours, followed by six readings every 3 hours, and a final three readings every two hours.
6. When the reading is taken, observe the following:
a) Diameter of the colony (once the diameter of the colony is measured, pick the colony and put it to grow in LB for nine hours)
b) Diameter of the halo that is achieved once HCl is applied
c) OD from a)
d) Estimate of the depth of the colony on the agar plate
For BL21 cell line that has been modified to contain nuclease:
1. Prepare 11-16 plates (DNAse - Agar + CMP)
2. Streak cells onto all plates at the same time
3. Incubate at 37°C
4. Apply HCl to the first plate before putting in the incubator (set time as zero)
5. Take a second reading after four hours, followed by six readings every 3 hours, and a final three readings
every two hours.
6. When the reading is taken, observe the following:
a. Diameter of the colony (once the diameter of the colony is measured, pick the colony and put it to grow in LB for nine hours)
b. Diameter of the halo that is achieved once HCL is applied
c. OD from a)
d. Estimate of the depth of the colony on the agar plate
Results:
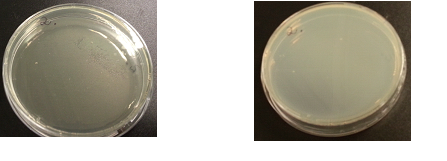
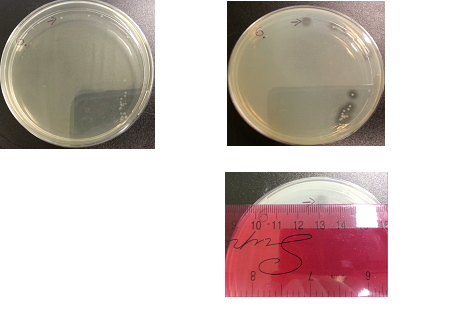
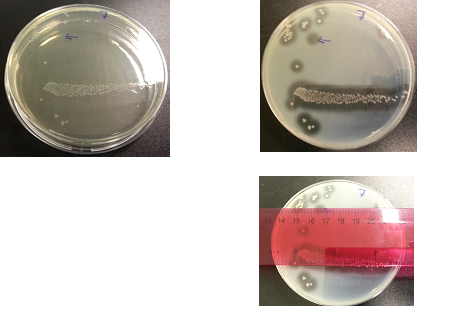
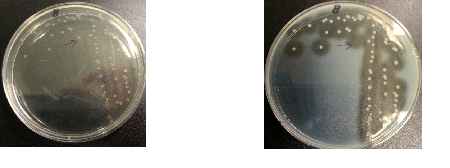
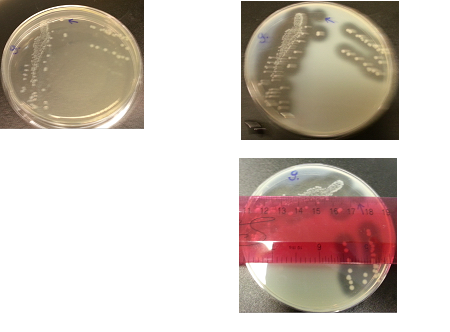
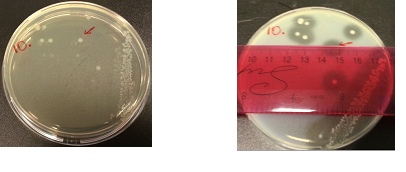
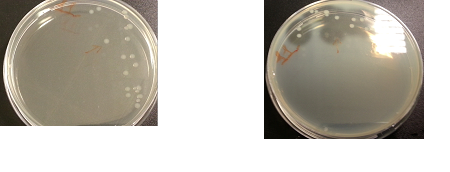
The following images show the results of the DNase assay with the left column showing the plates before HCL and the right column showing growth on plates after HCL:
The following table shows the colony and halo diameters, plus the OD of one plate at different sampling times.
| Date | Time | Colony diameter | Halo diameter | Absorbance at OD600 |
|---|---|---|---|---|
| 03.09.2012 | 12.30pm | 0 | 0 | 0 |
| 03.09.2012 | 16.30pm | 0 | 0 | 0 |
| 03.09.2012 | 19.30 pm | 0 | 0 | 0 |
| 03.09.2012 | 22.30 pm | 0.5mm | 1mm | 0.002 |
| 04.09.2012 | 01.30 am | 1mm | 3.5mm | 0.182 |
| 04.09.2012 | 04.30 am | 1.5mm | 7mm | 0.238 |
| 04.09.2012 | 07.30 am | 2mm | 9mm | 0.297 |
| 04.09.2012 | 10.30am | 2.5mm | 11mm | 0.518 |
| 04.09.2012 | 12.30pm | 3mm | 12.5mm | 0.701 |
| 04.09.2012 | 14.30pm | 3.5mm | 14mm | 0.811 |
| 04.09.2012 | 16.30pm | 4mm | 15mm | 0.906 |
Conclusion:
We have obtained data consistent with the previous DNase assay conducted, and thereby conclude that a linear correlation exists between colony size and halo diameter.

Tuesday 04/09
Aim:
To pick colonies from all the ligation transformation plates, prepared in Week 12. This included the ligations of curli, laccase and nuclease to the linker and the pb. This was done in order to purify the DNA and use it in subsequent ligations. This was done because gels from Week 12 showed unexpected bands. Hence, it was necessary to check whether this was due to unsuccessful purification and analytical digest, or due to unsuccessful ligation.
Methods:
Step 2 - Inoculating Colonies into a Selective Broth:: Add Yul of antibiotic to reach desired antibiotic concentration.
(For Ampicillin this is 50ug/ml, For Kanamycin it is 25ug/ml, for Tetracycline it is 15ug/ml, and for Chloramphenicol it is 25ug/ml)
Step 4 – Selecting a Colony: Select a clear, isolated colony and using an inoculation hoop scoop up a colony onto the tip. Deposit in the falcon tube
Step 5 - Culture: Culture your falcon tubes overnight at a temperature of 37 oC. Leave for no longer than 16 hours.
Wednesday 05/09
Step 1 - Aim:
To carry out a miniprep of the inoculations from the ligation transformation plates. After this, an analytical digest is prepared and run on a gel in order to confirm DNA identity
Method:
In order to carry out a miniprep, the following protocol was used:
Step 1 - Pellet Cells: Pellet 1.5-5ml bacterial culture containing the plasmid by centrifugation g = 6000
Time = 2 min
Temperature = (15-25oC)
Step 2 - Resuspend Cells: Add 250ul S1 to the pellet and resuspend the cells completely by vortexing or pipetting. Transfer the suspension to a clean 1.5ml microcentrifuge tube.
Step 3 - Puncturing Cell Membrane: Add 250ul S2 gently mix by inverting the tube 4-6 times to obtain a clear lysate. Incubate on ice or at room temperature for NOT longer than 5 min.
Step 4 - Neutralising S2: Add 400ul Buffer NB and gently mix by inverting the tube 6-10 times, until a white precipitate forms.
Step 5 - Centrifuge:
g: 14000
Time:10 minutes
Temperature: (15-25oC)
Step 6 - Centrifuge: Transfer the supernatant into a column assembled in a clean collection tube (provided. Centrifuge:
g = 10,000
Time: 1 minute
Temperature: (15-25oC)
Step 7 - Wash Column: Discard the flow-through and wash the spin column by adding 700ul of Wash Buffer. Centrifuge:
g - 10,000
Time - 1 minute
Temperature: (15-25oC)
Step 8 - Remove residual ethanol: Centrifuge:
g - 10,000
Time - 1 minute
Temperature: (15-25oC)
Step 9 - Elute DNA: Place the column into a clean microcentrifuge tube. Add 50-100ul of Elution Buffer, TE buffer or sterile water directly onto column membrane and stand for 1 min. Centrifuge:
g - 10,000
Time - 1 minute
Temperature: (15-25oC)
Step 10 - Storage: Store DNA at 4oC or -20oC
In order to carry out an analytical digest, the following protocol was used:
Step 2 - Adding Ingredient: Add the following ingredients to autoclaved/sterile eppendorf tubes
Insert Table
Step 3 - Addition of BioBrick: Flick contents gently and centrifuge.
Step 4 - Centrifuge:
RPM: 14000
Time: 1 minute
Temperature: 18oC
Step 5 - Digest Program: Place the samples on a thermocycler under the following conditions:
RPM: 550
Time: 1 hour
Temperature: 37oC
Step 6 - Denaturing Enzymes: If you are not running the samples on a gel immediately, denature the restriction enzymes by running the samples on a thermocycler under the following conditions:
RPM: 550
Time: 25 minutes
Temperature: 65oC
Temperature: 18oC
In order to run digest samples on a gel, the following protocol was used:
Preparing the Gel
Step 1: Within a conical flask, add 3ml 50X TAE, 1.5g Agarose, and 150ml RO water
Step 2: Heat for 1 min in microwave. Swirl. Heat again for 30s. If solution is clear stop. Else repeat.
Step 3: Cool solution under running cold water.
Step 4: Add 20ul ethidium bromide (normal concentration of EB solution is 500ug/ul)
Step 5: Pour into a sealed casting tray in a slow steady stream, ensuring there are no bubbles
Running a gel
Step 6: Add 1 part loading buffer to five parts of loading sample
Step 7: Position the gel in the tank and add TAE buffer, enough to cover the gel by several mm
Step 8: Add 5ul of DNA ladder to lane 1
Step 9: Add samples to the remaining wells
Step 10: Run at 100 volts for 1hour and 15 minutes
Imaging the Gel
Step 11: Place gel in GelDoc 2000 chamber
Step 12: Turn GelDoc 2000 chamber on
Step 13: From computer: Quantity One > Scanner > Gel_Doc_Xr>Manuqal Acquire
Step 14: Alter the exposure/settings to give a clear image.
TAE - Tris-acetate-EDTA
EDTA - ethylenediamine tetraacetic acid
Results:
Gel not shown
Conclusion:
Since band sizes were as expected, the DNA was taken forward and used in a preparative digest in preparation of ligation.
Step 2 - Aim:
To carry out a preparative digest on the purified DNA and then carry out a ligation.
Methods:
The following table shows the DNA part and the enzymes used to digest it.
| No | Part | Enzymes |
|---|---|---|
| 1 | laccase | X+P |
| 2 | curli | X+P |
| 3 | irrE | X+P |
| 4 | irrE | E+P |
| 5 | nuclease | X+P |
| 6 | nuclease | X+P |
| 7 | plasmid backbone | E+P |
| 8 | Plasmid backbone | E+P |
| 9 | Linker | E+S |
Digest Upstream Part with EcoRI-HF™ and SpeI
| Ingredient | Amount |
|---|---|
| Constitutive promoter – rbs construct | 500 ng |
| EcoRI-HF | 1 µl |
| Spel | 1 µl |
| 10X NEBuffer 2 | 5 µl |
| 100X BSA | 0.5 µl |
| H2O | to 50 µl |
Digest Downstream Part with Xbal and Pstl
| Ingredient | Amount |
|---|---|
| Curli (from pcr product)
Laccase (from pcr product) irrE (from pcr) Nuclease (in puc57) | 500 ng |
| Xbal | 1 µl |
| Pstl | 1 µl |
| 10X NEBuffer 2 | 5 µl |
| 100X BSA | 0.5 µl |
| H2O | to 50 µl |
| Ingredients | Amounts |
|---|---|
| EcoRI-HF | 1 µl |
| Pstl | 1 µl |
| 10X NEBuffer 2 | 5 µl |
| 100X BSA | 0.5 µl |
| H2O | to 50 µl |
Incubate all three restriction digest reactions at 37°C for 10 minutes and then heat inactivate at 80°C for 20 minutes.
Ligate the Upstream and Downstream Parts into the digested Destination Plasmid.
| Ingredients | Amounts |
|---|---|
| Upstream Part digestion | 2 µl |
| Downstream Part digestion | 2 µl |
| Destination Plasmid digestion | 2 µl |
| 10X T4 DNA Ligase Buffer | 2 µl |
| T4 DNA Ligase | 1 µl |
| H2O | 11 µl |
Incubate at room temperature for 10 minutes and then heat inactivate at 80°C for 20 minutes.
Summary of the cuts:
| No. | Part | Enzymes |
|---|---|---|
| 1 | laccase | X+P |
| 2 | curli | X+P |
| 3 | irrE | X+P |
| 4 | irrE | E+P |
| 5 | nuclease | X+P |
| 6 | nuclease | X+P |
| 7 | plasmid backbone | E+P |
| 8 | Plasmid backbone | E+P |
| 9 | Linker | E+S |
Step 3 - Aim:
To carry out a transformation of the ligation samples.
Methods:
Step 1 - Addition of BioBrick: To the still frozen competent cells, add 1 - 5 µL of the resuspended DNA to the 2ml tube.
Step 4 - Incubation: Close tube and incubate the cells on ice for 45 minutes.
Step 5 - Heat Shock: Heat shock the cells by immersion in a pre-heated water bath at 37ºC for 10 minutes.
Step 6 - Incubation: Incubate the cells on ice for 2 minutes.
Step 7 - Add media: Add 1.5ml of Lysogeny Broth and transfer to a falcon tube.
Step 8 - Incubation: Incubate the cells at 37ºC for 1 hour at RPM 550.
Step 9 - Transfer: transfer the solution back into a 1.5ml Eppendorf and centrifuge
RPM: 14000
Time: 2 minutes
Temperature (18-25oC)
Step 10 - Resuspend:Remove supernatant and resuspend in 100ul LB
Step 11 - Plating: Spread the resuspended cell solution onto a selective nutrient agar plate. Place the plates in a 37°C static incubator, leave overnight (alternatively a 30°C static incubator over the weekend)
Thursday 06/09
Aim:
To check whether transformations using ligation product on the previous day were successful.
Results & Conclusion:
No colonies were visible on the plates. Hence, the transformation was considered to be unsuccessful and needs to be repeated.

Wednesday 05/09
Aim:
To characterise the two strains of laccase obtained from the laccase ligation.
Method:
1) W3110 cells transformed with laccase and control W3110 cells are inoculated in 10mL of LB media, and incubated overnight at 37˚C in a 200rpm shaker
2) The O/N culture is then centrifuged at 6100g for 20 minutes, in order to extract the media containing laccase. The supernatant is retained.
3) In order to take readings, cuvettes are prepared with 2.2mL KH2PO4 buffer + 0.5mL Laccase supernatant + 0.3mL Syringaldazine, which is added immediately before readings are begun (as explained in step 4). A blank is created using 0.5mL of LB media instead of laccase supernatant.
4) The optical density at 530nm of each of the samples is measured at five minute intervals over 30 minutes.
Result:
Not shown
Conclusion:
We have determined that laccase strain 2 is a better laccase producing strain. Furthermore, we have proven the function of our BioBrick, as the oxidation activity of both laccase stratins was significantly higher than that of the control W3110 strain.

Thursday 06/09
Aim:
To compare the difference in growth of Indoli and E. coli growth in LB & MB media
Methods:
1. Inoculate 10ul of W3110 glycerol stock in 10ml of LB and another 10ul of W3110 glycerol stock in 10ml of MB
2. Repeat this with an indolifex glycerol stock
3. Measure the absorbence of all four samples every hour for 12 hours. Then take a final measurement after another 12 hours. Thus, measurements span 24 hours.
Results:
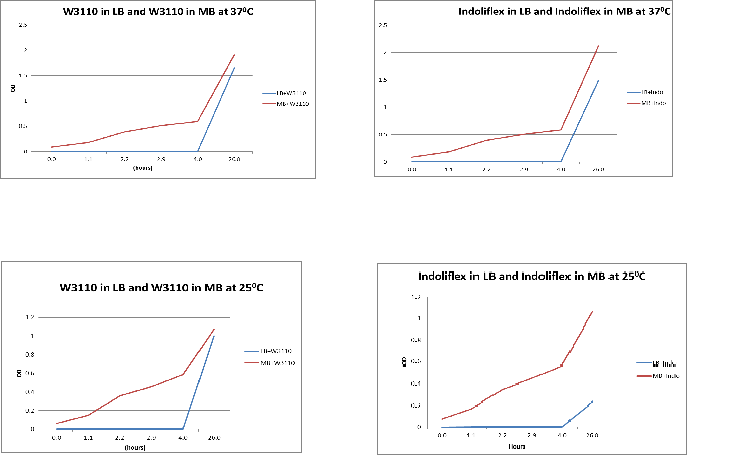
The following graphs show growth of the different strains and different media. The flasks were placed in a 370C shaker at 200rpm.
Conclusion:
From the above graphs, it can be seen that marine bacteria grows better in MB, irrespective of the temperature used.
 "
"