Team:NTU-Taida/Project/Sensor
From 2012.igem.org
Sensor
Contents |
Fatty Acid Biosensor
In order to regulate the release of GLP-1, we decide to use a fatty acid biosensor which will switch on when the intestinal environment is filled with fatty nutrient. Our team chose fatty acid as a sensor of metabolic states for two main reasons. First, fatty acid is considered as the major cause to obesity because of their highly calorigenic effect. Second, GLP-1 level increases once the blood glucose concentration rises, but there is no significant GLP-1 secretion when the free fatty acid concentration increases in blood stream. Therefore, we hope to compensate this weakness in the appetite regulating mechanisms. By designing the device that secrets GLP-1 on sensing fatty acid around, we aim to make a dietary bacteria.
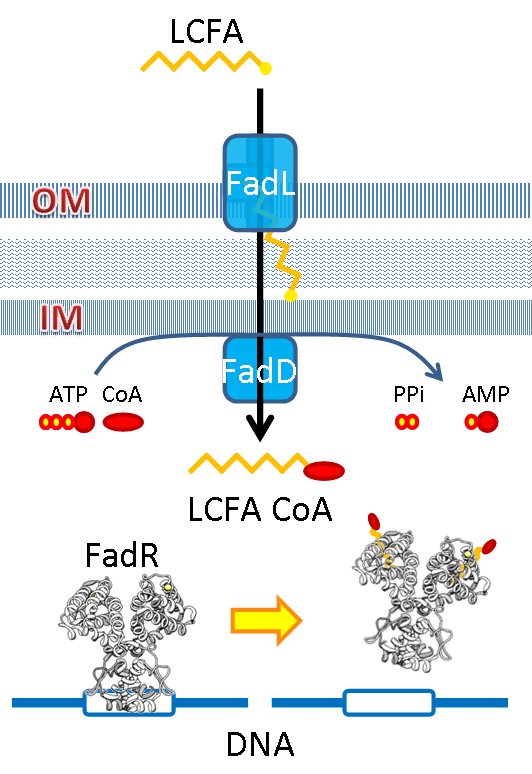
We use the natural fatty acyl-CoA biosensor, the fadBA promoter (PfadBA) from E. coli as the sensor in our design. fadBA is one of the β-oxidation gene which E. coli turns on when the fatty acyl-CoA concentration in the surrounding rises. Therefore, we plan to clone the PfadBA as the promoter in our device, which enables the bacteria to detect fatty acid in the environment.
However, the baseline expression of the reportor gene (mRFP) downstream of PfadBA is so high that we cannot see significant signal rise after induction of fatty acid (oleic acid). Therefore, we decide to lower the baseline expression by overexpression of FadR, an endogenous repressor of PfadBA, whose repressive function is antagonized by fatty acyl-CoA. By co-transforming constructs with PfadBA and FadR into the bacterial platform, it is made capable of changing gene expression in response to environmental fatty acid concentration and produce GLP-1 when the host is in taking a meal.
Phs Thermal Promoter
E. coli is pretty sensitive and responsive to temperature changes. We brought a thermal sensitive promoter Phs, located within dnaG, and transient induced to temperature upshift, to our circuit. As described by Wayne E. Talyor et al., Phs can forward upregulation of a series of proteins. (Taylor, 1984) The increase activity of Phs promoter is partially compatible with the increase synthesis of sigma factor. The quick respsonse of Phs is suitable since its the sigma factor synthesis peaks 10 minutes after the temperature upshift, and so does Phs RNA levels. The ratio of increase before and after the temperature rise can be more than 20 folds. On particular note, Phs is lack of consensus region over -10 region, which explains its poor activity in 30℃. Thus, Phs is marked as a candidate in our circuit, which can only be functional after ingestion into human body. We use this thermal promoter in the suicide device. See the Safety section for more information.
Modified cI Promoter with Temperature-Sensitive cI Repressor (CIts)
For the constitutive delivery of GLP-1 peptide in the case of prevention of neurodegenerative diseases, we utilize the composite BioBrick BBa_K098988 designed by Harvard iGEM team 2008 as the backbone of our constitutive delivery module, which gives raised expression of peptide under normal body temperature, 37℃. The original circuit involves a thermosensitive cI repressor (CIts) under the expression of a strong promoter, J23119, and is followed by a CI promoter (PcI) region and reporter of desire. Under conditions demonstrated before, its increase of reporter expression after temperature upshift is minimal (1.8 fold). And thus, we design novel cI promoter regions with altered binding affinity to CIts repressor in search of better efficiency. The original cI promoter has two binding regions: OR2 followed by OR1. Here we change these regions into new ones: OR1 and OL1 in order. The binding affinity between OL1 and cI repressor are larger than OR1 and OR2. Therefore, by enhancing the strength of baseline repression level of cI, we can improve the sensitivity to temperature difference of this device. See the Circuit section for further application of this device.
| OL1 | TACCACTGGCGGTGATA |
| OR1 | TACCTCTGGCGGTGATA |
| OR2 | TAACACCGTGCGTGTTG |
Reference
- Zhang F, et al. (2012) Design of a dynamic sensor-regulator system for production of chemicals and fuels derived from fatty acids. Nat Biotechnol 30(4):354-9.
- Iram SH, et al. (2005) Unexpected Functional Diversity among FadR Fatty Acid Transcriptional Regulatory Proteins. J Biol Chem 280(37):32148-56.
- van Aalten DM, et al. (2000) Crystal structure of FadR, a fatty acid-responsive transcription factor with a novel acyl coenzyme A-binding fold. EMBO J 19(19):5167-77.
- Taylor, W. E., D. B. Straus, et al. (1984). "Transcription from a heat-inducible promoter causes heat shock regulation of the sigma subunit of E. coli RNA polymerase." Cell 38(2): 371-381.
- Wedler, H. and R. Wambutt (1995). "A temperature-sensitive lambda cI repressor functions on a modified operator in yeast cells by masking the TATA element." Mol Gen Genet 248(4): 499-505
- Maeda YT, et al. (2006) Regulatory dynamics of synthetic gene networks with positive feedback. J Mol Biol 359:1107–1124.
 "
"