Team:Bielefeld-Germany/Protocols/Materials
From 2012.igem.org
(→Primers) |
(→Materials) |
||
| Line 65: | Line 65: | ||
* 20 g Dextrose (add 50 mL sterile stock solution (40% dextrose)) | * 20 g Dextrose (add 50 mL sterile stock solution (40% dextrose)) | ||
* Adjust pH to 6.5 | * Adjust pH to 6.5 | ||
| + | |||
| + | |||
| + | ===SDS-PAGE gel=== | ||
| + | |||
| + | The following amouts are for one gel. | ||
| + | Stacking gel 5 %: | ||
| + | *775 μL H<sub>2</sub>O | ||
| + | *1.25 mL 0,25 M Tris (pH 6,8) | ||
| + | *425 μL Bis/Acrylamide (0,8 %, 30 %) | ||
| + | *50 μL 5 % SDS | ||
| + | *25 μL 10 % Ammonium persulfate | ||
| + | *3 μL TEMED | ||
| + | |||
| + | Separating gel 12 %: | ||
| + | *1.5 mL H<sub>2</sub>O | ||
| + | *2.8 mL 1 M Tris (pH 8,8) | ||
| + | *3.0 mL Bis/ Acrylamide (0,8%, 30%) | ||
| + | *150 μL 5% SDS | ||
| + | *37.5 μL 10% Ammonium persulfate | ||
| + | *5 μL TEMED | ||
| + | |||
| + | |||
| + | ===SDS running buffer=== | ||
| + | *25 mM Tris [pH 8,3] | ||
| + | *192 mM Glycerol | ||
| + | *0.1 % SDS | ||
| + | |||
| + | |||
| + | ===4x Laemmli-buffer=== | ||
| + | *250 mM Tris-HCl | ||
| + | *40 % [v/v] Glycerol | ||
| + | *20 % [v/v] 2-Mercapthoethanol | ||
| + | *80 g L<sup>-1</sup> SDS | ||
| + | *0.04 g L<sup>-1</sup> BPB | ||
Revision as of 14:04, 19 September 2012
Contents |
Materials
</center> This is where we are going to list all our materials, devices and equipment that we have used.
Devices
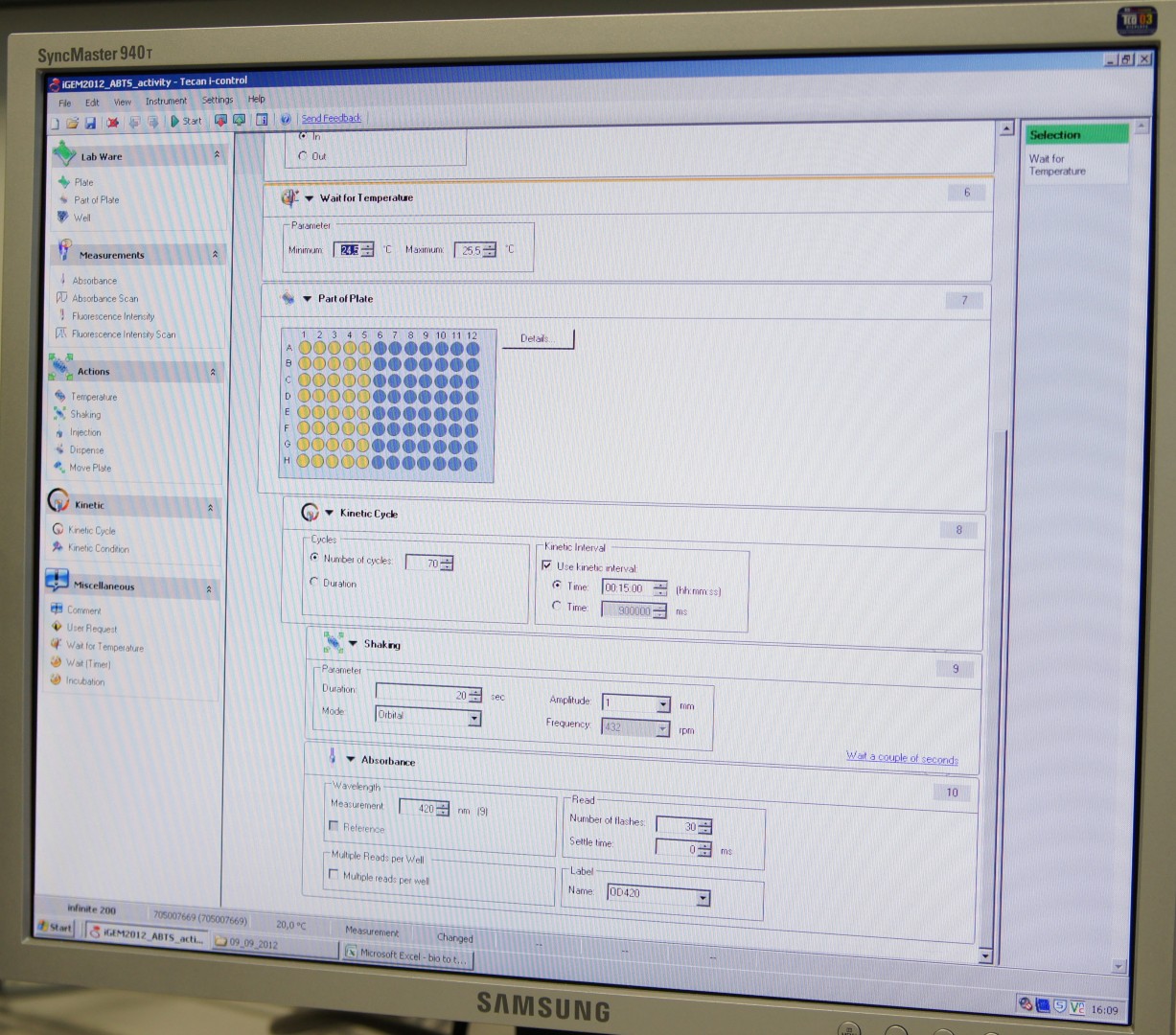
Tecan Infinite Microplate Reader
For measuring the Laccase activity we detected the level of oxidized ABTS via optical density at 420nm. The device we were able to use was a Tecan Infinite Reader M200. The program setup was in some parts adapted to the needs of our probes (like duration of the measurement) and in some parts standardized.
Used setup for Laccase activity measurements: Temperature: 25°C; Orbital shaking before each measuring cycle (time depends on duration of each cycle); Number of flashes: 30
Media, buffer and other solutions
Ampicillin stock solution
- Solubilize 100 mg mL-1 Ampicillin
- Store at -20 °C
Chloramphenicol stock solution
- Solubilize 20 mg mL-1 Chloramphenicol in 100 % Ethanol
- Store at -20 °C
TAE buffer
For 1 L of 50 x TAE buffer you need:
- 242.48 g Tris
- 41.02 g Sodiumacetate
- 18.612 g EDTA
- Adjust pH to 7.8 with acetic acid
- Solve in dH2O
10 mL of the stock is diluted in 1 L dH2O for the gel electrophoresis (0.5 x TAE buffer).
Briton Robinson Buffer
- 0,1 mM acetic acid
- 0,1 mM boric acid
- 0,1 mM phosphoric acid
- adjust to pH 5 with sodium hydroxide
DNA loading buffer
- 50 % (v/v) glycerol
- 1 mM EDTA
- 0.1 % (w/v) bromphenol blue
- Solve in ddH2O
LB media
For 1 L of LB media:
- 10 g Trypton
- 5 g Yeast extract
- 10 g NaCl
- 12 g Agar-Agar (for plates)
- Adjust pH to 7.4
YPD media
For 1 L of YPD media:
- 20 g Peptone
- 10 g Yeast extract
- 20 g Dextrose (add 50 mL sterile stock solution (40% dextrose))
- Adjust pH to 6.5
SDS-PAGE gel
The following amouts are for one gel. Stacking gel 5 %:
- 775 μL H2O
- 1.25 mL 0,25 M Tris (pH 6,8)
- 425 μL Bis/Acrylamide (0,8 %, 30 %)
- 50 μL 5 % SDS
- 25 μL 10 % Ammonium persulfate
- 3 μL TEMED
Separating gel 12 %:
- 1.5 mL H2O
- 2.8 mL 1 M Tris (pH 8,8)
- 3.0 mL Bis/ Acrylamide (0,8%, 30%)
- 150 μL 5% SDS
- 37.5 μL 10% Ammonium persulfate
- 5 μL TEMED
SDS running buffer
- 25 mM Tris [pH 8,3]
- 192 mM Glycerol
- 0.1 % SDS
4x Laemmli-buffer
- 250 mM Tris-HCl
- 40 % [v/v] Glycerol
- 20 % [v/v] 2-Mercapthoethanol
- 80 g L-1 SDS
- 0.04 g L-1 BPB
Primers
This is a list of primers we have used.
| primer name | length | sequence | |
|---|---|---|---|
| F | 28 | GTTTCTTCGAATTCGCGGCCGCTTCTAG | |
| R | 29 | GTTTCTTCCTGCAGCGGCCGCTACTAGTA | |
| pSB1C3-5aox1-f | 60 | CGCTAAGGATGATTTCTGGAATTCGCGGCCGCTTCTAGAGAGATCTAACATCCAAAGACG | |
| pSB1C3-5aox1-r | 30 | GGTGGCGGCGGGCGTTTCGAATAATTAGTT | |
| 5aox1-mfalpha1-f | 68 | AGAAGATCAAAAAACAACTAATTATTCGAAACGCCCGCCGCCACCATGAGATTTCCTTCAATTTTTAC | |
| 5aox1-mfalpha1-r | 20 | AGCTTCAGCCTCTCTTTTCT | |
| mfalpha1-aarI-taox1-f | 80 | GTATCTCTCGAGAAAAGAGAGGCTGAAGCTACACGCAGGTGGTATGTATCACCTGCGTGTCTTGCTAGAT TCTAATCAAG | |
| mfalpha1-aarI-taox1-r | 20 | TAAGCTTGCACAAACGAACT | |
| taox1-phis4-f | 60 | GTACAGAAGATTAAGTGAGAAGTTCGTTTGTGCAAGCTTATCATGCCATGGACAAGATTC | |
| taox1-phis4-r | 20 | GGCCGCTCGAGTATTCAGAA | |
| phis4-kozak-his4-f | 72 | AATAGTTTACAAAATTTTTTTTCTGAATACTCGAGCGGCCCCCGCCGCCACCATGACATTTCCCTTGCTACC | |
| phis4-kozak-his4-r | 30 | TTATTATTTCTCCATACGAACCTTAACAGC | |
| his4-3aox1-f | 60 | TCACCGCAATGCTGTTAAGGTTCGTATGGAGAAATAATAACGAGTATCTATGATTGGAAG | |
| his4-3aox1-r | 20 | AAAACAAGATAGTGCCCCTC | |
| 3aox1-pSB1C3-f | 60 | AGTCTGATCCTCATCAACTTGAGGGGCACTATCTTGTTTTTACTAGTAGCGGCCGCTGCA | |
| 3aox1-pSB1C3-r | 20 | CTCTAGAAGCGGCCGCGAAT | |
| taox-his4-f | 61 | GTACAGAAGATTAAGTGAGAAGTTCGTTTGTGCAAGCTTAAGATCTCCTGATGACTGACTC | |
| taox-his4-r | 27 | CTCGGATCTATCGAATCTAAATGTAAG | |
| his4-3aox1-f02 | 60 | TTATTTAGAGATTTTAACTTACATTTAGATTCGATAGATCCGAGTATCTATGATTGGAAG | |
| his4_gi537483_f | 46 | ACGTgaattcgcggccgcttctagagAGATCTCCTGATGACTGACT | |
| his4_gi537483_r | 41 | ctgcagcggccgctactagtaGATCTATCGAATCTAAATGT | |
| B.pumi_LAC_FW | 44 | ACGTGAATTCGCGGCCGCTTCTAGATGAACCTAGAAAAATTTGT | |
| B.pumi_LAC_RV | 41 | CTGCAGCGGCCGCTACTAGTATTACTGGATGATATCCATCG | |
| B.halo_FW | 44 | ACGTGAATTCGCGGCCGCTTCTAGATGAAAAAATCATATGGAGT | |
| B.halo_RV | 41 | CTGCAGCGGCCGCTACTAGTATTACTCAGGCATATTTGGAA | |
| T.thermo_LAC_FW | 44 | ACGTGAATTCGCGGCCGCTTCTAGATGCTGGCGCGCAGGAGCTT | |
| T.thermo_LAC_RV | 41 | CTGCAGCGGCCGCTACTAGTACTAACCCACCTCGAGGACTC | |
| E.coli_LAC_FW_T7 | 79 | ACGTGAATTCGCGGCCGCTTCTAGAGtaatacgactcactatagggagagaggagaaaaATGCAACGTCGTGATTTCTT | |
| E.coli_LAC_RV_HIS | 62 | CTGCAGCGGCCGCTACTAGTATTATTAGTGATGGTGATGGTGATGTACCGTAAACCCTAACA | |
| Xcc_LAC_FW_T7 | 79 | ACGTGAATTCGCGGCCGCTTCTAGAGtaatacgactcactatagggagagaggagaaaaATGTCATTCGATCCCTTGTC | |
| Xcc_LAC_RV_HIS | 62 | CTGCAGCGGCCGCTACTAGTATTATTAGTGATGGTGATGGTGATGTGCCTCCACCCGCACTT | |
| B.pumi_LAC_FW_T7 | 79 | ACGTGAATTCGCGGCCGCTTCTAGAGtaatacgactcactatagggagagaggagaaaaATGAACCTAGAAAAATTTGT | |
| B. pumi_LAC_RV_HIS | 62 | CTGCAGCGGCCGCTACTAGTATTATTAGTGATGGTGATGGTGATGCTGGATGATATCCATCG | |
| E.coli_LAC_FW_T7 | 79 | ACGTGAATTCGCGGCCGCTTCTAGAGtaatacgactcactatagggagagaggagaaaaATGCAACGTCGTGATTTCTT | |
| E.coli_LAC_RV_HIS | 62 | CTGCAGCGGCCGCTACTAGTATTATTAGTGATGGTGATGGTGATGTACCGTAAACCCTAACA | |
| T.thermo_LAC_FW_T7 | 79 | ACGTGAATTCGCGGCCGCTTCTAGAGtaatacgactcactatagggagagaggagaaaaATGCTGGCGCGCAGGAGCTT | |
| T.thermo_LAC_RV_HIS | 62 | CTGCAGCGGCCGCTACTAGTATTATTAGTGATGGTGATGGTGATGACCCACCTCGAGGACTC | |
| Pc_lac35_FW_oS | 38 | acgtcacctgcgtgtagctgccatagggcctgtggcgg | |
| Pc_lac35_RV | 39 | acgtcacctgcgtgtcaagTCAGAGGTCGCTGGGGTCAA | |
| Tc_lac5_FW_oS | 38 | acgtcacctgcgtgtagctggtatcggtcctgtcgccg | |
| Tc_lac5_RV | 39 | acgtcacctgcgtgtcaagTTACTGGTCGCTCGGGTCGC | |
| Tv_lac5.P.FW | 43 | gaattcgcggccgcttctagATGTCGAGGTTTCACTCTCTTCT | |
| Tv_lac5.S.RV | 41 | CTGCAGCGGCCGCTACTAGTATTACTGGTCGCTCGGGTCGC | |
| Pc_lac35.P.FW | 43 | gaattcgcggccgcttctagATGTCGAGGTTCCAGTCCCTCTT | |
| Pc_lac5.S.RV | 41 | CTGCAGCGGCCGCTACTAGTATTACTGGTCGCTCGGGTCGC | |
| J23117_RBS_FW | 70 | aattcgcggccgcttctagagttgacagctagctcagtcctagggattgtgctagcaaagaggagaaata | |
| J23117_RBS_RV | 70 | ctagtatttctcctctttgctagcacaatccctaggactgagctagctgtcaactctagaagcggccgcg | |
| J23103_RBS_FW | 70 | aattcgcggccgcttctagagctgatagctagctcagtcctagggattatgctagcaaagaggagaaata | |
| J23103_RBS_RV | 70 | ctagtatttctcctctttGCTAGCATAATCCCTAGGACTGAGCTAGCTATCAGctctagaagcggccgcg | |
| J23110_RBS_FW | 70 | aattcgcggccgcttctagagtttacggctagctcagtcctaggtacaatgctagcaaagaggagaaata | |
| J23110_RBS_RV | 70 | ctagtatttctcctctttgctagcattgtacctaggactgagctagccgtaaactctagaagcggccgcg | |
| J23103_K_FW | 25 | CTGACAGCTAGCTCAGTCCTAGGTA | |
| J23110/117_K_FW | 25 | TTTACGGCTAGCTCAGTCCTAGGTA | |
| T7_K_FW | 26 | TAATACGACTCACTATAGGGAAAGAG |
| 55px | | | | | | | | | | |
 "
"