Team:Paris Bettencourt/Encapsulation
From 2012.igem.org
|
Aim : Harness bacteria-containing gel beads to assure cell entrapment and complement activity of genetic safety systems. Experimental system: Bacterial cells are encapsulated in alginate beads. We used a cell containment assay based on plating to assess the release of cells from alginate beads. In addition, we aimed at improving the entrapment of cells through stabilization by polyethyleneimine and covalent cross-linkage by glutaraldehyde. Achievements :
|
Contents |
Overview
Polymer gels have found a place in microbial biotechnology by providing a means of spatial organization. The micro-environments within gel beads can grant the microbes within protection, nutrients, and selective agents/chemicals. Given this, gel beads are already attractive for environmental applications of genetically modified bacteria. Synthetic bacterial systems may benefit from (or require) nutrients and agents added to gel beads. Many other practical reasons for use of beads exist, such as transportation and analysis.
Our interest is to take the concept of cell encapsulation further by implementing total cell entrapment. While normal gel beads protect the bacteria within from the surroundings, we are here to ensure the surroundings are protected from the bacteria within the beads. A method was found for encapsulation and entrapment of yeast [2] and was adapted to synthetic bacterial systems.
The above beads show the visual contrast between treated and untreated alginate beads. The pink ones are treated, and the white ones are not.
Objectives
Our goal is to design a live-bacteria entrapment system. More than just encapsulating bacteria, we want to fully prevent their escape from the bead body into the surroundings.
Alginate and other gel-based beads have been used successfully to prolong enzymatic activity in bioreactors[3], but systems such as these are usually designed to allow steady release of microbes. This is not acceptable when microbes containing potentially dangerous synthetic genes are being used in the environment, so we aim to prevent release of bacteria entirely.
Covalent Stabilization of Alginate Beads
A detailed protocol for creation of stabilized alginate beads can be found [http://openwetware.org/wiki/IGEM:Paris_Bettencourt_2012/Protocols/stabilized_beads here]
Cell-containing 2% alginate can be created by mixing 4% alginate in equal parts with cell suspension. Alginate beads are created by extruding 2% alginate beads through a needle into 1.5% CaCl2 solution and curing for 1 hour. The beads are then equilibrated at 50mM CaCl2 overnight.
Covalent Stabilization procedure: Polyethyleneimine (0.5% w/v with 50mM CaCl2) diffuses slowly into alginate beads over 2-24 hours to vary thickness. The beads are washed with water. The covalent cross-linkages are formed between polyethyleneimine molecules by incubating in glutaraldehyde (1% with 10mM sodium phosphate pH=7) for 1 minute followed by washing.
Experiments and results
White-Blue Screening Assay
We had some concerns regarding the toxicity of chemicals used for the covalent stabilization of alginate. Glutaraldehyde, specifically, is used to fix and kill cells. We found, however, that cells are at least partially protected within the beads from death by glutaraldehyde. We demonstrated this by a white-blue screening assay. BL21 strain (LacZα fragment inducible by IPTG) was used.
- 2% Alginate beads containing cells were prepared (10mL saturated culture mixed with 10 mL 4% Alginate).
- 4g beads were washed with MgSO4 once and set aside at 4°C (untreated alginate beads).
- 4g beads were treated as described above with polyethyleneimine and glutaraldehyde.
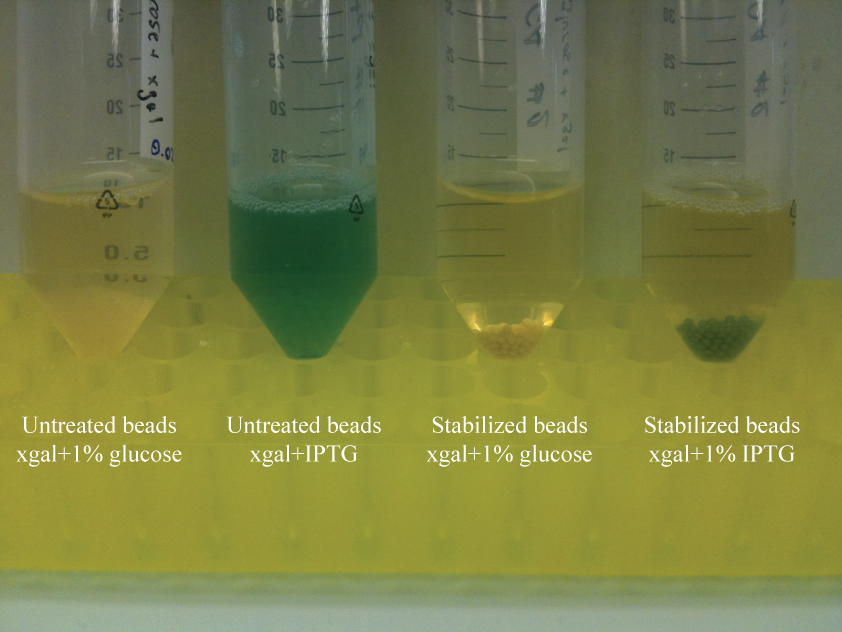
- Treated and untreated beads were separated into tubes of LB containing either xgal and 1% glucose (for efficient repression of LacZα) or xgal and IPTG.
Results
Both treated and untreated beads turn blue in the assay. Because the cells within beads were not induced prior to encapsulation, this indicated that cells are able to grow and express proteins within the beads. Interestingly, the supernatant of the induced untreated beads is blue, indicating release of cells. The supernatant of the induced stabilized beads is not blue. This suggests efficient entrapment of cells in the stabilized beads, but another assay is required to validate this result.
Cell Containment Assay
Our objective is to entrap cells that are still viable and able to perform metabolism. To asses this, beads were suspended in buffer and allowed to incubate at room temperature over several days. Presuming that treated beads could result in total cell containment, we wished to see if more viable cells would be released by physically destroying the beads.
Experimental setup
- 2% Alginate beads containing cells were prepared (10mL saturated culture mixed with 10 mL 4% Alginate).
- 2g beads were washed with MgSO4 once and set aside at 4°C (untreated alginate beads).
- 2g beads were treated as described above with polyethyleneimine and glutaraldehyde.
- Untreated and Stabilized beads were added to 10 mL PBS buffer and left at room temperature.
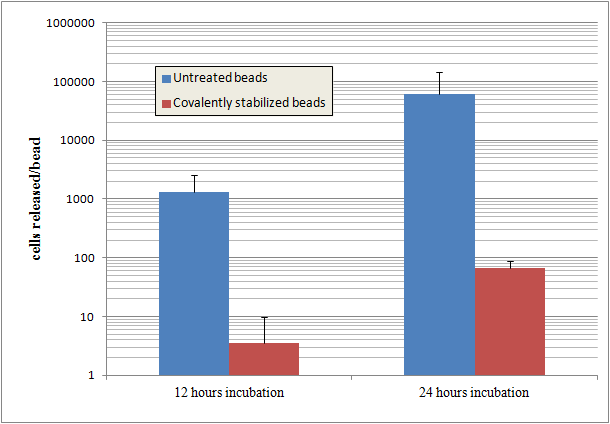
- Supernatant was plated at 12 hours and 24 hours incubation at different dilutions to quantify release of cells.
Results
Beads that are covalently stabilized entrap cells much more effectively than untreated alginate beads. The untreated beads release cells at a rate up to 1000 times more than the stabilized beads.
Alginate beads are disrupted by the phosphate in PBS buffer. Despite this, stabilized beads still show a high degree of cell entrapment after 24 hours.
Preliminary Colicin system test
A mixture of 50% RPF expressing and 50% colicin producing cells was encapsulated in alginate beads. As a control, RFP producing cells were encapsulated with wild type cells. After incubating the beads in LB for about 12 hours, the two different mixtures were compared. The number of cells that had grown in the supernatant of the colicin-bearing mixture was 2.5 times less than in the supernatant of the mixture. This indicates that the combination of colicin-toxin system is effective with the use of alginate beads.
Further Experiments
This method for stabilization of beads was adapted from a method designed for yeast cells. Although the system appears effective at preventing cell release, we are concerned that the treatment method is lethal for cells. We need to follow up the cell containment assay with cell activity assays to assure that we have living cells capable of performing their intended function. This may include a white-blue screening assay to see if cells are capable of protein production within.
We intend to find a covalent cross-linkage agent that is less lethal to cells than glutaraldehyde and then demonstrate better performance using this.
Lastly, it is a high priority to get alginate bead systems to work with our other biosafety systems such as the colicin toxin system. We can analyze the viability of cells using propidium iodide stain with the beads. In the end, the beads are supposed to work with the complete biosafety system. Particularly the arabinose-based delay system requires beads to retain arabinose physically and modulate
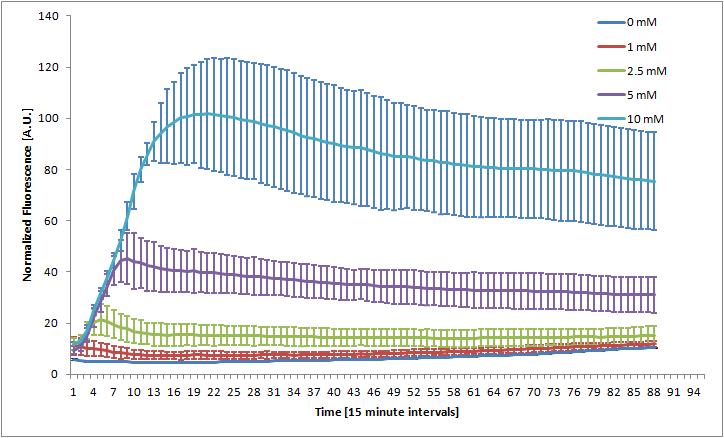
Bristol 2010 Nitrate Reporter
The Bristol 2010 team probably did not have access to an automated kinetic fluorescence and OD measurement device. We monitored GFP expression level at different KNO3 induction levels in order to better characterize the part.
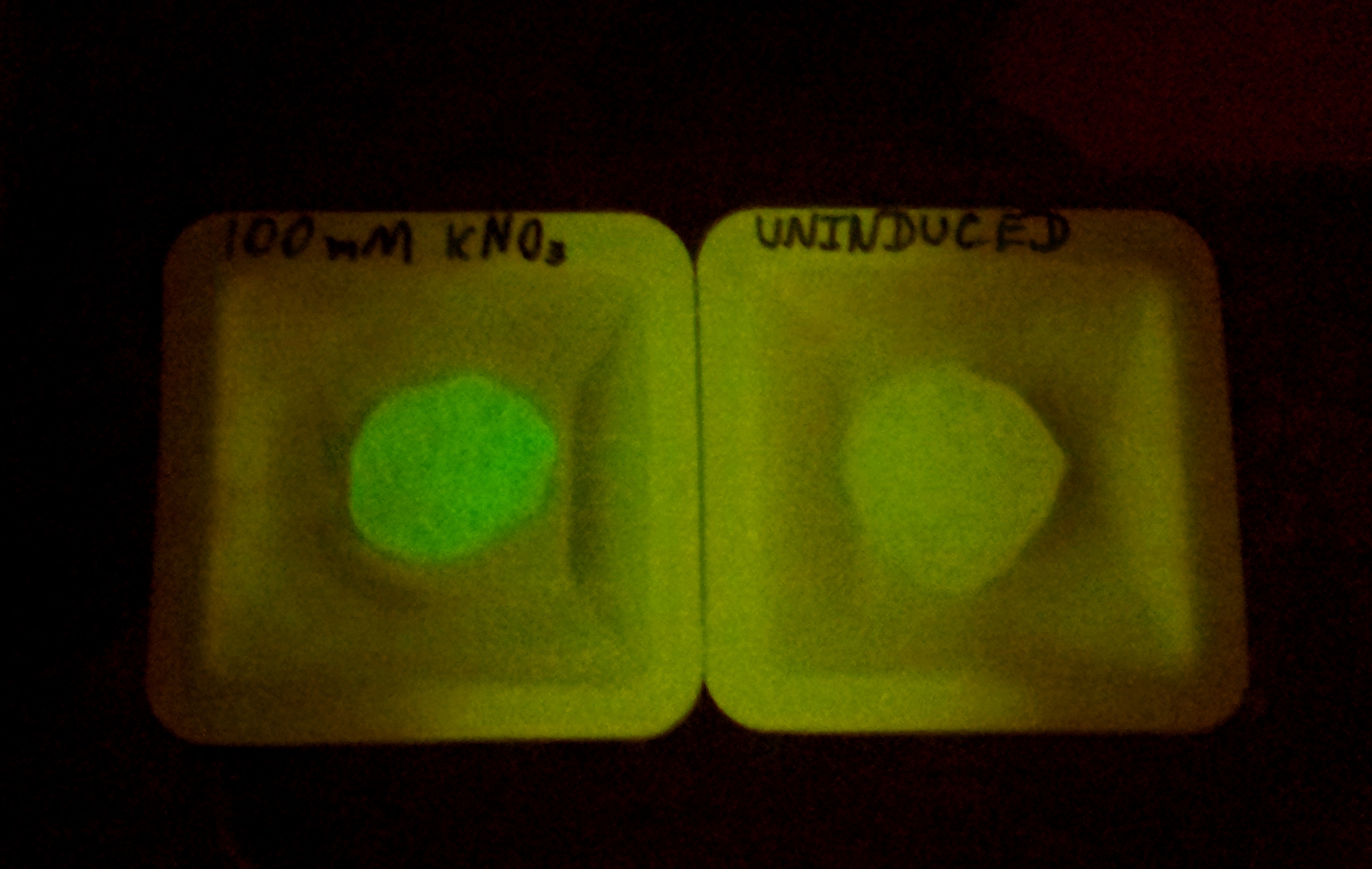
Alginate beads were created (uninduced) and then incubated in plain LB and LB with 100mM KNO3. The difference in green fluorescence was visible by eye under the Illumatool.
References
1. agrEcoli, Bristol 2010 igem team wiki
2. Birnbaum, S., Pendleton, R., Larsson, P.-O. & Mosbach, K. Covalent stabilization of alginate gel for the entrapment of living whole cells. Biotechnology Letters 3, 393-400 (1981). [http://dx.doi.org/10.1007/BF01134097 Paper]
3. Boon, N. et al. Bioaugmenting bioreactors for the continuous removal of 3-chloroaniline by a slow release approach. Environmental science & technology 36, 4698-4704 (2002). [http://view.ncbi.nlm.nih.gov/pubmed/12433184 Paper]
 "
"


 Overview
Overview Delay system
Delay system Semantic containment
Semantic containment Restriction enzyme system
Restriction enzyme system MAGE
MAGE Encapsulation
Encapsulation Synthetic import domain
Synthetic import domain Safety Questions
Safety Questions Safety Assessment
Safety Assessment