Team:Paris Bettencourt/Modeling
From 2012.igem.org
Contents |
Overview
Safety is an important issue in synthetic biology, especially on environmental related projects. We already tried to answer the question, “how safe is safe enough?” by involving experts, publics and scientists, and also building biosafety devices. However, to really answer the question, actually we need first to ask ourselves a more basic question, “how do we measure safety?”. As we see synthetic biology as an engineering approach to biology, we could think about adaptation of safety engineering, a well studied engineering subset, that has been widely use to minimize risks on many fields of engineering, such as mechanical engineering, aircrafts, and manufactures. However, the risks they face are surely different from the risks of synthetic biology. Here we propose an approach to assess safety for environmental release of genetically engineered bacteria (GE bacteria).
Background
According to Dana et al1 there are four areas of risk research in environmental application of synthetic biology:
- Differences in the physiology of natural and synthetic organisms will affect how they interact with the surrounding environment,
- Escaped microorganisms have the potential to survive in receiving environments and to compete successfully with non-modified counterparts,
- Synthetic organisms might evolve and adapt quickly, perhaps filling new ecological niches, and
- Gene transfer.
Knowing that there are different areas of risk that we need to take into account, we need to design our safety containment with different parts to deal with each of them. Identifying the relationship between one part and the others will help us to see the reliability of the overall system.
Objectives
- Adapting existing safety assessment tools to synthetic biology.
- Proposing methods to assess safety in synthetic biology.
Methods and Results
We propose a method to assess hazards and risk2 in releasing genetically engineered bacteria to the environment. As a case study, we want to release GE bacteria which will perform some function in the environment and we want to apply 3 sets of bWARE safety containment (alginate beads, bWARE killswitch, semantic containment) to control the hazards and risks. Here we focus more on the reliability of the safety containment system rather than the functional part, although the similar assessment can also be applied to see the functional part reliability.
Hazards Identification
Identifying hazards means that we need to find and understand the possible harm that may happen in the application of our system. Generally there are two potential hazard in environmental application of synthetic biology1 which are the success escape of the GE bacteria followed by a success competition with the natural strain and the horizontal gene transfer.
Risk Assessment
Risk assessment will give an idea what kind of risk we face in releasing the GEO in the environment and help to design the safety containment. Here we adapted a risk management method in workplaces complying with health and safety law 3.
To-do-list to assess risk in 5 steps:
- Identify the hazards
- Decide who might be harmed and how
- Evaluate the risks and decide on precaution
- Record your findings and implement them
- Review your assessment and update if necessary
Worksheet example:
| What are the hazards? | Who might be harmed and how? | What are we already doing? | What further action is necessary? |
| GE bacteria outcompete natural strains | GE bacteria escaping the containment may outcompete natural strains if they have better fitness, creating natural imbalance | Using harmless strain or strain with low fitness compared to natural strain (standard E coli for lab) | Designing a safety containment to prevent the cells reproducing themselves at some point and/or separating them from natural strains |
| Horizontal gene transfer (HGT) | Other strain/species may uptake engineered gene, and if the gene gives advantage in the fitness, it may outcompete other strain and creating natural imbalance | - | Designing a safety containment to degrade DNA and/or separate the DNA from the environment and/or prevent natural strain having advantages from the DNA |
Control hazards and risks
In this step we decided what control elements we want to put to our system to reduce the risks. We should choose the best suitable control elements depending on the system function and application. As we already know that we want to apply bWARE safety containment, here we describe the role of each safety parts in the risks reduction.
- GE bacteria outcompete natural strains
- In order to prevent the success escape of GE bacteria and outcompete the natural strains, we will stop their reproduction by putting a self killing mechanism to kill the cells after they perform the function.
- In case of the failure of the killing mechanism, we will put the cells inside a physical containment so there will be no physical interaction between them and the surrounding cells.
- HGT
- We will use DNAse to degrade DNA after the cells perform the function so they won’t leave any genetic material behind.
- In case of the failure on inefficiency of the DNAse, the physical containment will still protect the DNA from the environment.
- In case of the leakiness of the physical containment, the DNA has special encryption system with the semantic containment so the receiver cell will not be able to read it.
Check controls
To check and predict the reliability of our system, we used fault tree analysis to assess multiple safety containment in one biodevice. This top-down approach allows us to see the relationship between each containment element and predict the overall failure probability. With this method we can see what basic events are the key elements in our system.
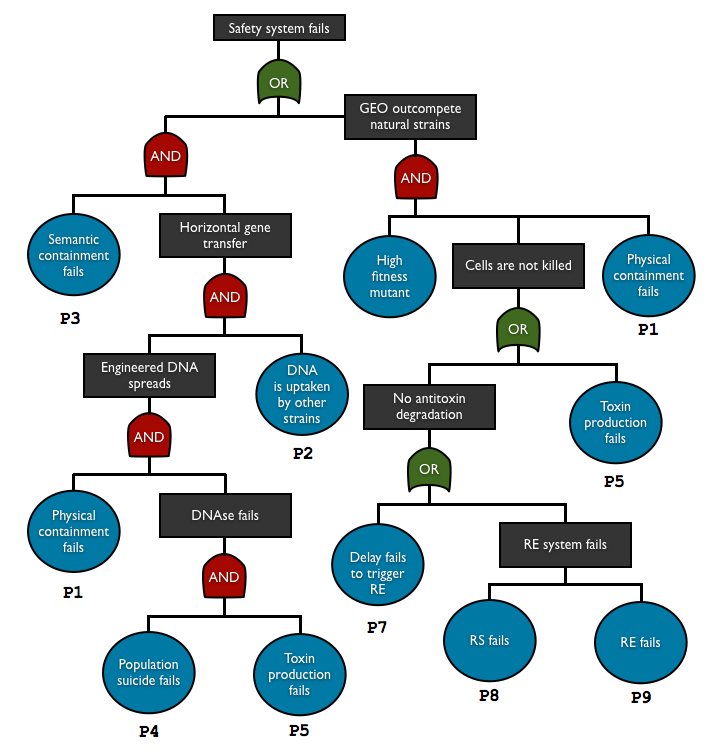
AND gate represents independent events
P(A and B) = P(A)P(B)
OR gate corresponds to set of union
P(A or B) = P(A) + P(B)
so the total failure probability of the system is
Ptotal = P1.P2.P3.P4.P5 + P1.P6.(P5+P7+P8+P9)
List of the basic events
| Notation | Failure component | Failure mode | Consequence | Method for determining the probability |
| P1 | physical containment | leakage | DNA/cell escape | experiment |
| P2 | DNA uptake | DNA transferred into natural strain | HGT | transconjugant to donor ratio in HGT is typically <10-5 [12] |
| P3 | semantic containment | mutation/recombination/plasmid loss | HGT | estimation* |
| P4 | population suicide | no surrounding cells with enough toxin | no death | based on the cell density and the expectation number of surrounding non-mutant |
| P5 | toxin production | mutation/recombination/plasmid loss | no DNA degradation (w/o antitoxin), no death (with antitoxin) | estimation* |
| P6 | mutant fitness | beneficial mutation | outcompetition of the mutant | estimation* |
| P7 | delay system | mutation/recombination/plasmid loss | no antitoxin degradation | estimation* |
| P8 | restriction enzyme | mutation/recombination/plasmid loss | no antitoxin degradation | estimation* |
| P9 | restriction site | mutation/recombination/plasmid loss | no antitoxin degradation | estimation* |
References
1 - Genya V Dana et al. Four steps to avoid a synthetic biology disaster. 2012. Nature vol 483
2 - WorkSafe Victoria. A handbook for workplaces, controlling OHS hazards and risks. 2007. Edition No.1
3 - “Five steps to risk assessment” from [http://www.hse.gov.uk/risk/fivesteps.htm http://www.hse.gov.uk/risk/fivesteps.htm]. - risk management method in workplaces complying with health and safety law
 "
"


 Overview
Overview Delay system
Delay system Semantic containment
Semantic containment Restriction enzyme system
Restriction enzyme system MAGE
MAGE Encapsulation
Encapsulation Synthetic import domain
Synthetic import domain Safety Questions
Safety Questions Safety Assessment
Safety Assessment