Team:HokkaidoU Japan/Notebook/aggregation Week 11
From 2012.igem.org
Contents |
September 10th
Digestion of eCFP-RBS-pSB1A2 and pBAD-RBS-pSB1A2
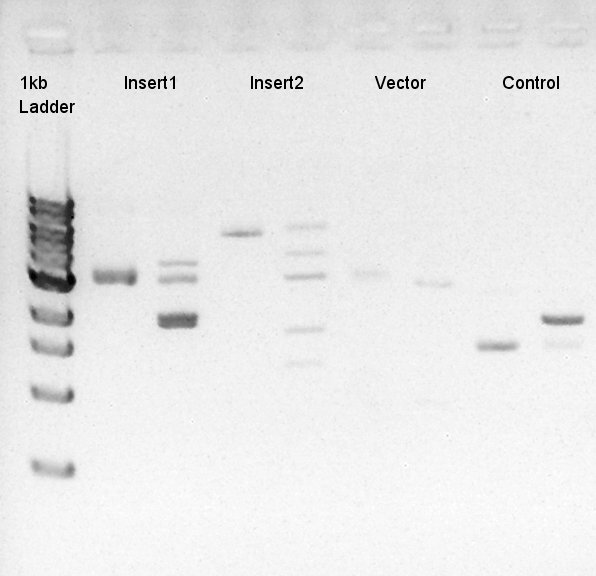
To make a construct of pBAD-RBS-eCFP-RBS-Ag43-dT-pSTV28 by 3piece ligation, we digested pBAD-RBS-eCFP-RBS-pSB1A2 with EcoRI & SpeI, Ag43-dT-pSB1AK3 (previously digested with HindIII) with XbaI & NotI and pSTV28 with EcoRI and NotI. Then we digested pT7-RBS-pSB1C3 with XbaI & SpeI as a control for confirmation of the ability of restriction enzyme. Insert1 (pBAD-RBS-eCFP-RBS-pSB1A2)
| DNA solution ( 35ng/ul) | 17 ul |
| EcoRI | 1 ul |
| SpeI | 1 ul |
| 10xH buffer | 3 ul |
| DW | 8 ul |
| Total | 30 ul |
Insert2 (Ag43-dT-pSB1AK3)
| DNA solution ( 35ng/ul) | 25 ul |
| XbaI | 1 ul |
| NotI | 1 ul |
| 10xK buffer | 1.5 ul |
| 100xBSA | 0.3ul |
| DW | 1.5 ul |
| Total | 30 ul |
Vector(pSTV28)
| DNA solution ( 15ng/ul) | 9 ul |
| EcoRI | 1 ul |
| NotI | 1 ul |
| 10xH buffer | 2 ul |
| 10xBSA | 0.2ul |
| DW | 7 ul |
| Total | 20 ul |
control (pT7-RBS-pSB1C3)
| DNA solution (30~40 ng/ul) | 10 ul |
| XbaI | 1 ul |
| SpeI | 1 ul |
| 10xM buffer | 2 ul |
| DW | 6 ul |
| Total | 20 ul |
| Number | Degree | Minute |
| 1 | 37 | 120 |
| 2 | 70 | 20 |
| 3 | 4 | HOLD |
Ethanol precipitation of ptet-RBS-eYFP-dT-pSB1A2 and pSTV28
- Added 5 ul of NaoAc, 1.5 ul of glycogen and 125 ul of 100% ethanol.
- Centrifuged in 14000 rpm, 30 min at 4C.
- Remove supernatant and added 220 ul of 70% ethanol.
- Centrifuged in 15000 rpm, 15 min at 4C.
- Remove supernatant and air drying in room temperature then added 5 ul of DW.
We confirmed that the concentration of Insert1 DNA solution is 50 ng/ul, Insert2 DNA solution is 10~15 ng/ul and Vector DNA solution is 10 ng/ul.
Ligation of pBAD-RBS-eCFP-RBS, Ag43-dT and pSTV28
| Vector DNA (10 ng/ul) | 4 ul |
| Insert1 DNA (50 ng/ul) | 2 ul |
| Insert2 DNA (10~15 ng/ul) | 4 ul |
| Ligation Mighty Mix | 10 ul |
| Total | 20 ul |
Ligation reaction time was in detail below.
| Degree | Minute |
| 16 | 30 |
| 65 | 10 |
| 4 | Hold |
Transformation of pBAD-RBS-eCFP-RBS-Ag43-dT-pSTV28
- Mixed 2 ul pBAD-RBS-eCFP-RBS-Ag43-dT-pSTV28 ligation product to 50 ul of thawed competent cells on ice.
- Incubated on ice for 30 min.
- Mixed 350 ul of LB.
- Incubated for 2 hrs to get the resistance to Chloramphenicol.
- Prepared and Labeled two plastic plates with LB plate medium which contained appropriate antibiotics (LBC).
- Plated 300 ul of the culture onto first dish and spread.
- Mixed 450 ul of LB to 50 ul of the culture and plated 300 ul of it onto second dish and spread.
- Incubated the plates at 37C for 14 hours.
September 11th
Colony PCR of pBAD-RBS-eCFP-RBS-pSB1A2
| DNA solution | 4 ul |
| Kapa-Taq(Taq polymerase) | 5 ul |
| Forward Primer(pbad-f2 primer) | 0.5 ul |
| Reverse Primer(PS-R down primer) | 0.5 ul |
| Total | 10 ul |
| Number | Degree | Second |
| 1 | 95 | 120 |
| 2 | 95 | 30 |
| 3 | 53.3 | 30 |
| 4 | 72 | 60 |
| 5 | 72 | 60 |
| 6 | 4 | HOLD |
Cycle:2~4 x 35
- We noticed that this step was actually 68.9 degree after reaction.
We used N1 (DW only) and N2(pBAD-RBS-Ag43-dT-pSB1A2)as controls. Desired product is about 542bp.
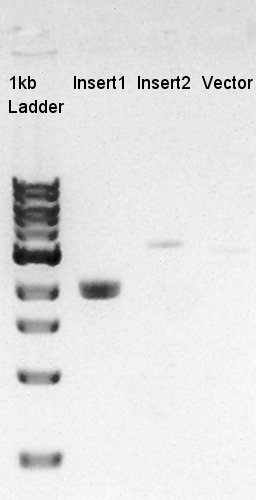
[[image:|thumb|Colony PCR result]]
Because of mis-setting the PCR program, we failed to amplify desired DNA fragment so we retried the colony PCR in same reagent, correct PCR machine program.
We noticed that the ligated DNA contains at least Ag43-dT-BioBrick Suffix
 "
"