Team:Bielefeld-Germany/Protocols
From 2012.igem.org
Contents |
Protocols - Overview
In this section we are going to note all protocols that the teams have used for their work.
| General Protocols | Team Cloning | Team Cultivation |
| Team Activity Test | Team Immobilization | Team Substrate |
| Team Yeast | Team Sequencing | Team Modelling |
Materials
This is where we are going to list all our materials, devices and equipment that we have used.
Devices
Tecan Infinite Microplate Reader
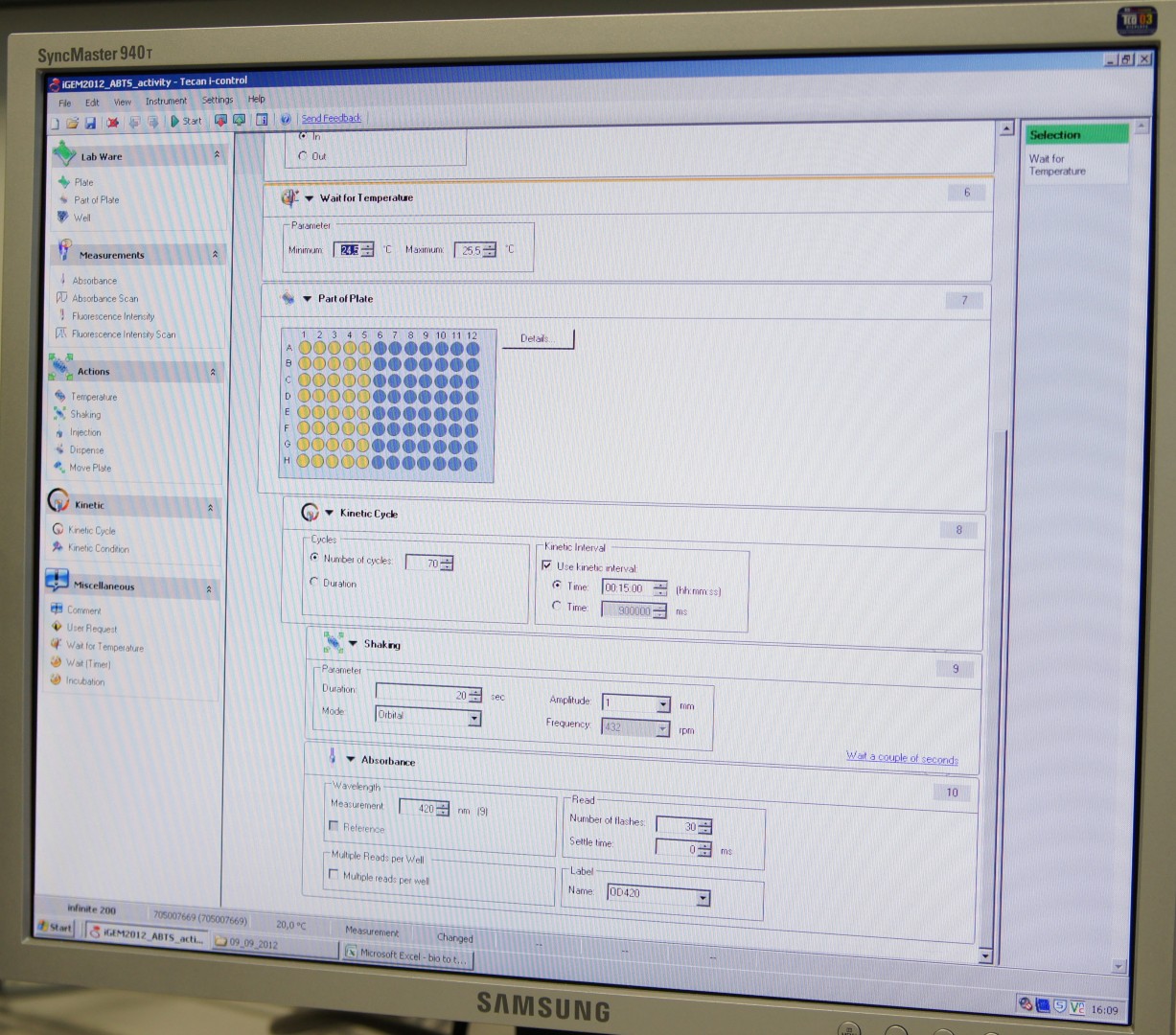
For measuring the Laccase activity we detected the level of oxidized ABTS via optical density at 420nm. The device we were able to use was a [http://www.tecan.com/platform/apps/product/index.asp?MenuID=1812&ID=1916&Menu=1&Item=21.2.10.1 Tecan Infinite Reader M200]. The program setup was in some parts adapted to the needs of our probes (like duration of the measurement) and in some parts standardized.
Used setup for Laccase activity measurements: Temperature: 25°C; Orbital shaking before each measuring cycle (time depends on duration of each cycle); Number of flashes: 30
Media, buffer and other solutions
Ampicillin stock solution
- Solubilize 100 mg mL-1 Ampicillin
- Store at -20 °C
Chloramphenicol stock solution
- Solubilize 20 mg mL-1 Chloramphenicol in 100 % Ethanol
- Store at -20 °C
TAE buffer
For 1 L of 50 x TAE buffer you need:
- 242.48 g Tris
- 41.02 g Sodiumacetate
- 18.612 g EDTA
- Adjust pH to 7.8 with acetic acid
- Solve in dH2O
10 mL of the stock is diluted in 1 L dH2O for the gel electrophoresis (0.5 x TAE buffer).
DNA loading buffer
- 50 % (v/v) glycerol
- 1 mM EDTA
- 0.1 % (w/v) bromphenol blue
- Solve in ddH2O
LB media
For 1 L of LB media:
- 10 g Trypton
- 5 g Yeast extract
- 10 g NaCl
- 12 g Agar-Agar (for plates)
- Adjust pH to 7.4
YPD media
For 1 L of YPD media:
- 20 g Peptone
- 10 g Yeast extract
- 20 g Dextrose (add 50 mL sterile stock solution (40% dextrose))
- Adjust pH to 6.5
Primers
This is a list of primers we have used.
| primer name | length | sequence | |
|---|---|---|---|
| pSB1C3-5aox1-f | 60 | CGCTAAGGATGATTTCTGGAATTCGCGGCCGCTTCTAGAGAGATCTAACATCCAAAGACG | |
| pSB1C3-5aox1-r | 30 | GGTGGCGGCGGGCGTTTCGAATAATTAGTT | |
| 5aox1-mfalpha1-f | 68 | AGAAGATCAAAAAACAACTAATTATTCGAAACGCCCGCCGCCACCATGAGATTTCCTTCAATTTTTAC | |
| 5aox1-mfalpha1-r | 20 | AGCTTCAGCCTCTCTTTTCT | |
| mfalpha1-aarI-taox1-f | 80 | GTATCTCTCGAGAAAAGAGAGGCTGAAGCTACACGCAGGTGGTATGTATCACCTGCGTGTCTTGCTAGATTCTAATCAAG | |
| mfalpha1-aarI-taox1-r | 20 | TAAGCTTGCACAAACGAACT | |
| taox1-phis4-f | 60 | GTACAGAAGATTAAGTGAGAAGTTCGTTTGTGCAAGCTTATCATGCCATGGACAAGATTC | |
| taox1-phis4-r | 20 | GGCCGCTCGAGTATTCAGAA | |
| phis4-kozak-his4-f | 72 | AATAGTTTACAAAATTTTTTTTCTGAATACTCGAGCGGCCCCCGCCGCCACCATGACATTTCCCTTGCTACC | |
| phis4-kozak-his4-r | 30 | TTATTATTTCTCCATACGAACCTTAACAGC | |
| his4-3aox1-f | 60 | TCACCGCAATGCTGTTAAGGTTCGTATGGAGAAATAATAACGAGTATCTATGATTGGAAG | |
| his4-3aox1-r | 20 | AAAACAAGATAGTGCCCCTC | |
| 3aox1-pSB1C3-f | 60 | AGTCTGATCCTCATCAACTTGAGGGGCACTATCTTGTTTTTACTAGTAGCGGCCGCTGCA | |
| 3aox1-pSB1C3-r | 20 | CTCTAGAAGCGGCCGCGAAT | |
| taox-his4-f | 61 | GTACAGAAGATTAAGTGAGAAGTTCGTTTGTGCAAGCTTAAGATCTCCTGATGACTGACTC | |
| taox-his4-r | 27 | CTCGGATCTATCGAATCTAAATGTAAG | |
| his4-3aox1-f02 | 60 | TTATTTAGAGATTTTAACTTACATTTAGATTCGATAGATCCGAGTATCTATGATTGGAAG | |
| his4_gi537483_f | 46 | ACGTgaattcgcggccgcttctagagAGATCTCCTGATGACTGACT | |
| his4_gi537483_r | 41 | ctgcagcggccgctactagtaGATCTATCGAATCTAAATGT | |
| B.pumi_LAC_FW | ACGTGAATTCGCGGCCGCTTCTAGATGAACCTAGAAAAATTTGT | ||
| B.pumi_LAC_RV | CTGCAGCGGCCGCTACTAGTATTACTGGATGATATCCATCG | ||
| Xcc_LAC_FW_T7 | ACGTGAATTCGCGGCCGCTTCTAGAGtaatacgactcactatagggagagaggagaaaaATGTCATTCGATCCCTTGTC | ||
| Xcc_LAC_RV_HIS | CTGCAGCGGCCGCTACTAGTATTATTAGTGATGGTGATGGTGATGTGCCTCCACCCGCACTT | ||
| E.coli_LAC_FW_T7 | ACGTGAATTCGCGGCCGCTTCTAGAGtaatacgactcactatagggagagaggagaaaaATGCAACGTCGTGATTTCTT | ||
| E.coli_LAC_RV_HIS | CTGCAGCGGCCGCTACTAGTATTATTAGTGATGGTGATGGTGATGTACCGTAAACCCTAACA | ||
| T.thermo_LAC_FW_T7 | ACGTGAATTCGCGGCCGCTTCTAGAGtaatacgactcactatagggagagaggagaaaaATGCTGGCGCGCAGGAGCTT | ||
| T.thermo_LAC_RV_HIS | CTGCAGCGGCCGCTACTAGTATTATTAGTGATGGTGATGGTGATGACCCACCTCGAGGACTC |
This page lists all molecular genetics protocols we use in our project
Yeat: Complete genome isolation
The complete genome isolation was done with the [http://www.promega.com/resources/protocols/technical-manuals/0/wizard-genomic-dna-purification-kit-protocol/ Promega Wizard genomic DNA purification system kit].
- Pellet 10 mL of over-night liquid culture grown in YPD broth in a 1.5 mL tube by centrifugation at 14,000 x g for 2 minutes.
- Remove the supernatant.
- Resuspend the cells in 90 μL of 50 mM EDTA.
- Add 10 μL of 1000u lyticase and pipet 4 times to mix.
- Incubate the sample at 37°C for 60 minutes to digest the cell wall.
- Centrifuge the sample at 14,000 × g for 2 minutes and then remove the supernatant.
- Add 300 μl of Nuclei Lysis Solution to the cell pellet and pipet to mix.
- Add 100 μl of Protein Precipitation Solution and vortex at high speed for 20 seconds.
- Let the sample sit on ice for 5 minutes.
- Centrifuge at 14,000 × g for 3 minutes.
- Transfer the supernatant containing the DNA to a clean 1.5 ml tube containing 300 μl of room temperature isopropanol.
- Gently mix by inversion until the DNA is visible.
- Centrifuge at 14,000 × g for 2 minutes.
- Carefully decant the supernatant and drain the tube on clean absorbent paper.
- Add 300 μl of room temperature 70% ethanol and invert the tube several times to wash the DNA pellet.
- Centrifuge at 14,000 × g for 2 minutes.
- Drain the tube on clean absorbent paper and allow the pellet to air-dry for 15 minutes.
- Add 50 μl of DNA Rehydration Solution.
- Add 1.5μl of RNase Solution to the purified DNA sample. Vortex the sample for 1 second and incubate at 37°C for 15 minutes.
- Rehydrate the DNA by incubating at 65°C for 1 hour. Periodically mix the solution by gently tapping the tube.
- Store the DNA at 2–8°C.
Arabidopsis thaliana: Growth Conditions and Plant Material
Six weeks old A. thaliana plants, ecotype Columbia 0 (wildtype), have been gratefully offered by Patrick Treffon and Thorsten Seidel. They have been cultivated under normal day conditions (14 hours light [100 µmol ⁄ quanta m-2s-1] at 21°C, 10 hours darkness at 18°C). For induction of the formation of siliques the plants were shifted into long day conditions (16 hours light [100 µmol ⁄ quanta m-2s-1] at 21°C, 18 hours darkness at 18°C). After two weeks in long day conditions the plants have developed 2 cm long siliques. The siliques were harvested and frozen in liquid nitrogen for further use.
Arabidopsis thaliana: Total RNA Isolation
The frozen plant material has to be grinded in a precooled mortar in liquid nitrogen. About 120 mg of pulverized plant material are transfered into a precooled 2 ml Eppendorf tube and kept frozen until the following steps:
- Add 0.5 ml lysis buffer and immediately homogenize through rough shaking.
- Add 0.5 ml of saturated phenol and mix strongly.
- Add 0.5 ml of chloroform isoamyl alcohol (24:1) and vortex again at high speed for at least 30 seconds.
- Centrifugate for 5 min at 13,000 rpm.
- The lower phase contains now lipids and lipophilic compounds. The upper phase contains nucleic acids (~ 550 µl) and has to be carefully transferred into a new 2 ml Eppendorf tube. This tube has to be filled with 0.5 ml saturated phenol and 0.5 ml chloroform isoamyl alcohol (24:1). Mix immediately.
- Centrifugate at 13,000 rpm for 3 minutes.
- Prepare a new 2 ml Eppendorf tube with 1 ml of chloroform isoamyl alcohol (24:1). Transfer the upper aqueous phase (~ 540 µl) containing the protein purified nucelic acids into the new tube and vortex strongly.
- Centrifugate at 13,000 rpm for 3 minutes.
- Prepare a new 1.5 ml Eppendorf tube with 0.5 ml of pure isopropanol. For the last time transfer the upper phase (~ 400 µl) into the new tube and mix gently.
- Incubate the mixture over night at -20°C. The nucleic acids will precipitate.
- Centrifugate the samples at 13,000 rpm for 15 minutes at 4°C.
- Discard the supernatant and resuspend the pellet in 375 µl sterile H2O.
- Add 125 µl 8 M lithium chloride and incubate for 2 hours on ice at 4°C. At this point most of the RNA is going to be precipitated.
- Centrifugate at 13,000 rpm at 4°C and discard the supernatant.
- Wash the pellet with 100 µl 70% (v/v) ethanol and discard it after centrifugation.
- Dry the pellet at room temperature.
- Dissolve the pellet in sterile H2O (~ 25 µl, depending on the size of the pellet).
- Check the quantity and quality of the RNA with a Nanodrop spectrophotometer before starting with a cDNA synthesis.
Arabidopsis thaliana: cDNA Synthesis
After a successful total RNA isolation the RNA has to be translated in cDNA through RT-PCR:
- Take 3 µg/µl of total RNA and add sterile H2 to 8 µl.
Additionally add
| 1,1 mM | Oligo-d(T)-Primer |
| 0,83 mM | dNTPs |
| 3,5 µl | H2O |
- Vortex and centrifugate shortly.
- Incubate the samples for 10 minutes at 70°C.
- Immediately transfer the samples into ice water for 5 minutes.
- After cooling the samples centrifugate shortly.
- To start the synthesis add
| 6 µl | 5xMMLV-Puffer |
| 4,5 µl | H2O |
| 1 µl | MMLV-reverse Transkriptase [200 U/µl] |
| 0,5 µl | RNasin RNase-Inhibitor [40 U/µl] |
- Mix the samples and centrifugate shortly.
- Incubate for 1 hour at 42°C to translate the RNA into cDNA.
- Transfer the samples to 70°C for 15 minutes to stop the reaction.
- The new synthesized cDNA can be used for PCR after diluting 1:10 with water. Store the cDNA at -20°C.
Ethanol precipitation to clean DNA
To get rid of distracting salts the DNA has to be cleaned. For this we used the following protocol:
- If the volume of the sample containing the DNA is less than 200 µl bring the volume up to 200 µl.
- Add 1/10th volume of 3M sodium acetate and mix.
- Now add 2 volumes of -20°C cold 100% ethanol and vortex for 10 seconds.
- The sample can now be placed in a -20°C freezer overnight or incubated for 30 minutes at -80°C.
- Centrifugate for 10 minutes at 4°C.
- Discard the supernatant containing the ethanol.
- Wash the pellet with 500 µl 4°C cold 70% ethanol by rolling the sample gently.
- Discard the supernatant.
- Let the pellet dry at room temperature or speedvac the pellet.
- Resuspend the Pellet in water (amount is depending on the size of the pellet).
| 55px | | | | | | | | | | |
 "
"