Team:Bielefeld-Germany/Labjournal/week19
From 2012.igem.org
(→weekly meeting) |
(→weekly meeting) |
||
| Line 49: | Line 49: | ||
==Week 19 (09/03 - 09/09/12)== | ==Week 19 (09/03 - 09/09/12)== | ||
=== weekly meeting === | === weekly meeting === | ||
| + | |||
| + | * We added a band for our sponsors on every wiki-page. | ||
| + | * On thursday we will meet with René Röspel, member of the german parliament. | ||
| + | * We are invited to the annual reception of the University Bielefeld on the 5th of october. Well that won't work for us. | ||
| + | * We still are working on a design for our team t-shirt. | ||
===Monday September 3rd=== | ===Monday September 3rd=== | ||
Revision as of 20:54, 25 September 2012
Contents |
Week 19 (09/03 - 09/09/12)
weekly meeting
- We added a band for our sponsors on every wiki-page.
- On thursday we will meet with René Röspel, member of the german parliament.
- We are invited to the annual reception of the University Bielefeld on the 5th of october. Well that won't work for us.
- We still are working on a design for our team t-shirt.
Monday September 3rd
- Team Cloning of Bacterial Laccases:
- We sent different plasmids for sequencing.
- We got another E.coli strain (Rosetta-gami 2) from our supervisor Dr. Christian Rückert. The cells carry a chloramphenicol-resistant plasmid, pRARE2, which supplies tRNAs for seven rare codons, AUA, AGG, AGA, CUA, CCC, GGA and CGG under the control of their native promoter. We want to use this strain for the expression of TTHL and BHAL hopeful that some rare codons are the reason that the laccases can’t be expressed by now. So we need the genes under control of a constitutive promoter because this strain has no T7 polymerase and we can’t use our inducible constructs. Further the plasmids need to have AMP resistance, because otherwise we can't select on positive transformed cells. Hopefully the plasmids, we sent for sequencing are ok, so we can use them.
- Team Cellulose Binding Domain:
- A autoinduced culture of KRX with the CBDcex(T7)+GFP_His-plasmid does not seem to produce the protein (no green fluorescence).
- Team Site Directed Mutagenesis:
- Clean-up of tvel10-t243g-PCR
- pfu-gradient-PCR of xccl-plasmid with xccl-primers with no product.
- Team Cultivation & Purification:
- Made one half of the SDS-Pages of the flask cultivation from 08/30.
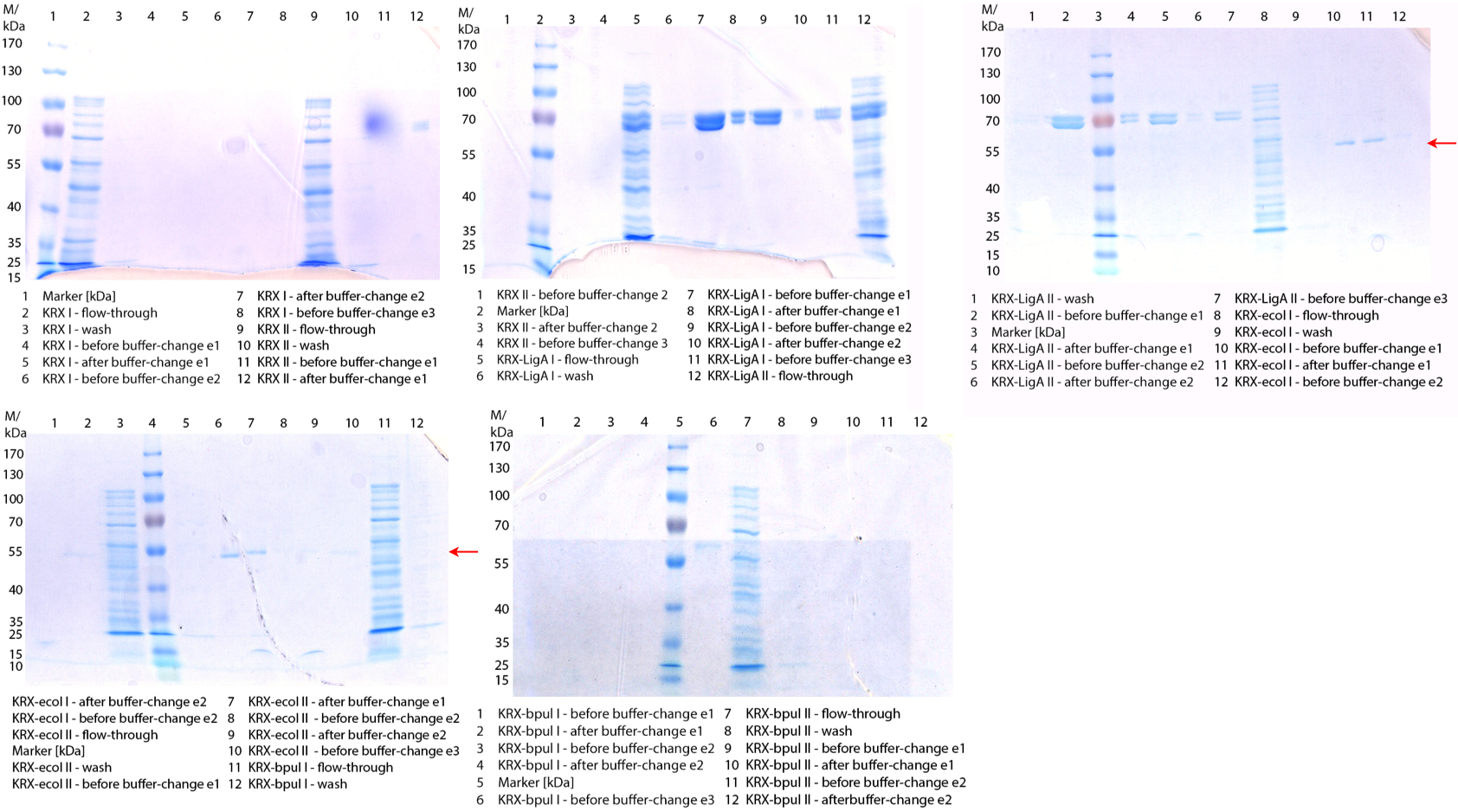 Figure 1: Cultivation from 08/28 (smaller scale but with a double determination) of E.coli KRX with BBa_K863000, BBa_K863005, BBa_K863010, BBa_K863015 or BBa_K863020 with positive (BBa_K525710) and negative control (E.coli KRX without plasmid). 60 mL in 300 mL flasks without baffles with autoinduction medium, 60 µg/mL chloramphenicol at 37 °C for 12 hours.
Figure 1: Cultivation from 08/28 (smaller scale but with a double determination) of E.coli KRX with BBa_K863000, BBa_K863005, BBa_K863010, BBa_K863015 or BBa_K863020 with positive (BBa_K525710) and negative control (E.coli KRX without plasmid). 60 mL in 300 mL flasks without baffles with autoinduction medium, 60 µg/mL chloramphenicol at 37 °C for 12 hours. - Work on culture of E. coli KRX containing BBa_K863000 (08/31) was continued and we performed the cell disruption by high-pressure homogeniser in 3 cycles. Then it was purificated by using a TALON column. The product was given to the activity team for analysis.
- Fermentation of E. coli KRX with BBa_K863121 for the cellulose binding team.
- Settings: fermenter: Braun, autoinduction medium, 60 µg/mL chloramphenicol, 3 L, 37 °C, stirrer on cascade to hold a pO2 of 50 %, airflow: 2 NL/m. This time we cultivated only for 9 hours and then decreased the temperature at 20 °C for a better folding of the protein for 1 hour. We had some problems with cascade at the end of the fermentation.
- Cells of today's fermentation of E. coli KRX with BBa_K863121 were disrupted via high-pressure homogeniser and purified by using the Talon column. The flow-through fluoresced very strong, so that not that much of the protein had bound to the column. Conclusion: The column had to be reloaded.
- Fermentation of E. coli KRX containing BBa_K863000.
- Settings: fermenter: Braun, autoinduction medium, 60 µg/mL chloramphenicol, 3 L, 37 °C, stirrer on cascade to hold a pO2 of 50 %, airflow: 2 NL/m, 9 hours. We only cultivated 9 hours, because we saw that a great amount of our cells already died.
- Made precultures of E. coli KRX containing BBa_K863000, BBa_K863005 and one containing BBa_K863113.
- Made one half of the SDS-Pages of the flask cultivation from 08/30.
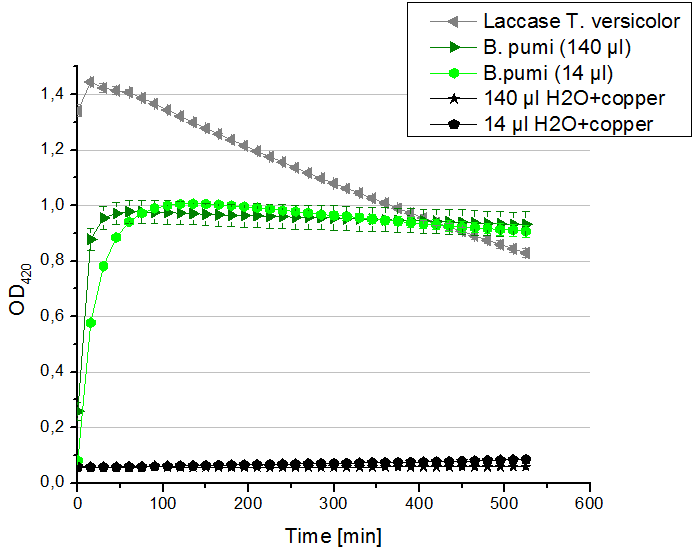
- Team Activity Tests:
- Today our hard-working Team Cultivation offered us some BPUL laccases. After fermentation and TALON purification the laccase was still hanging around in imidazole buffer, so we rebuffered it into nanopure water. We incubated the sample with copper and considered to measure the protein concentration before running the activity tests. Using the protocol for determining the protein concentration via Bradford (standards were 1 mg/ml, 1.5 mg/ml and 2 mg/ml BSA) we got a final BPUL laccase concentration of 0.22 mg/ml. With this information we designed the following activity test different as usual. As a positive control we took 140 µl of our bought laccase TVEL0 with a concentration of 0,021 mg/ml. Accordingly to this concentration we applied 14 µl of the BPUL laccase to get approximately the same amount of laccase into each well. Additionally we tested 140 µl of the BPUL laccase to make sure that we will definitely see any activity. After measuring the samples the whole night we got our results in the morning. It turned out, that our store bought laccase TVEL0 was rapidly active as seen before in our activity tests. The BPUL laccase was also active. Using 140 µl of it led to the maximal state of oxidation of ABTS after 45 minutes. With the reduced amount of 14 µl of laccase this optimum was reached after 90 minutes. For our following activity tests we want to include the protein concentration again.
Tuesday September 4th
- We signed in for our tracks today! Our first choice is the 'Environment' track since it is obvious that we want to clean up drinking water and therefore help our environment. The second choice is the 'New Application' track. We are hoping our filter system can be applied in wastewater treatment plants and establishes a new area of water purification.
- Team Cloning of Bacterial Laccases:
- Digest of pSB1C3 + J23110 and pSB1C3+ J23103 for suffix insertion of the laccase genes.
- Digest of the different laccase ORFs for cloning in pSB1C3 backbone without any regulatory elements. Ligation of tthl, bhal, ecol and bpul in pSB1C3.
- Transformation of ligations from day before (J23100+J61101+pSB1A2+bhal_His).
- Team Site Directed Mutagenesis:
- plated six colonies of tvel-t243g for plasmid-isolation.
- Team Cultivation & Purification:
- Second half of the SDS-Pages of the flask cultivation from 08/30 was made.
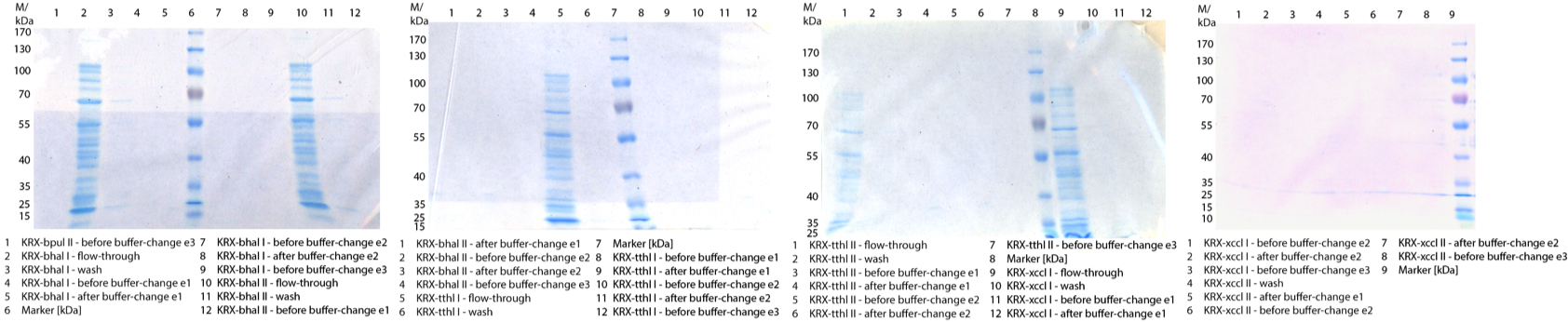 Figure 1: Cultivation from 08/28 (smaller scale but with a double determination) of E.coli KRX with BBa_K863000, BBa_K863005, BBa_K863010, BBa_K863015 or BBa_K863020 with positive (BBa_K525710) and negative control (E.coli KRX without plasmid). 60 mL in 300 mL flasks without baffles with autoinduction medium, 60 µg/mL chloramphenicol at 37 °C for 12 hours.
Figure 1: Cultivation from 08/28 (smaller scale but with a double determination) of E.coli KRX with BBa_K863000, BBa_K863005, BBa_K863010, BBa_K863015 or BBa_K863020 with positive (BBa_K525710) and negative control (E.coli KRX without plasmid). 60 mL in 300 mL flasks without baffles with autoinduction medium, 60 µg/mL chloramphenicol at 37 °C for 12 hours. - We reloaded our TALON column.
- After reloading we repeated the purification of BPUL produced by fermentation on 08/31. Therefore the flowthrough and the washing fraction were used. The different fractions should be tested by the activity team.
- A new fermentation of E. coli KRX containing BBa_K863005 was started
- Settings: fermenter: Infors, autoinduction medium, 60 µg/mL chloramphenicol, 3 L, 37 °C, stirrer on cascade to hold a pO2 of 50 %, airflow: 2 NL/m, 9 hours. We shorten the time, because the effect should be similar to the production of BPUL.
- The fermentation of E. coli KRX containing BBa_K863000 and the one containing BBa_K863005 were harvested and after centrifugation stored at 4 °C, because our high-pressure homogeniser is out of work.
- Made a flask cultivation for the cellulose binding team: cultivation of E. coli KRX containing the <partinfo>BBa_K863103</partinfo>-plasmid to express a cellulose binding domain (CBD) fused to a His-tagged GFP.
- Settings: 1 L flask with baffles, autoinduction medium, 60 µg/mL chloramphenicol, final volume: 250 mL, 37 °C, 120 rpm, 9 hours. We made a three-fold determination, but did not get fluorescing cultures. The problem is to purify this product, because it might be impossible to remove the protein from the column. If it binds via CBD and not via His-tag.
- Made precultures of E. coli KRX containing plasmids with laccases from E. coli, B. pumilus and BBa_K863012 behind a constitutive promoter as well as one containing BBa_K863000
- Second half of the SDS-Pages of the flask cultivation from 08/30 was made.
- Team Substrate Analytic:
- Today we let TVEL0 degrade Estradiol and Estrone. We prepared seven 2 mL Eppendorf tubes with: 425 µL BR-buffer, 50 µL TVEL0 and 25 µL Estradiol. To stop the reaction we added 1,5 mL MeOH. The Estradiol concentration should be 2 µg ml -1. We stopped at houre 0, 1, 2, 3, 4, 5, and over night, and mesured the Estradiol concentration with the HPLC and the LCMS.
Wednesday September 5th
- Team Cloning of Bacterial Laccases:
- Transformation of tthl, bhal, ecol and bpul in pSB1C3.
- Team Cultivation & Purification:
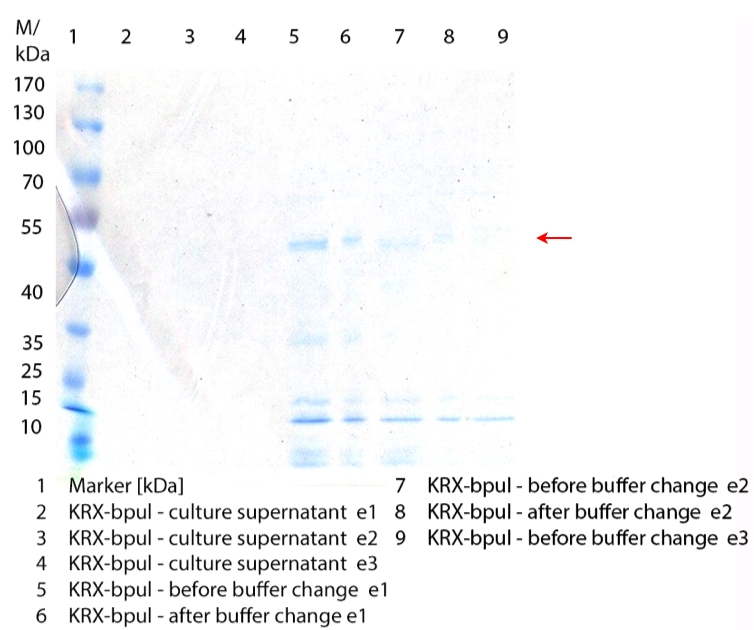 Figure 1: First fermentation of E. coli KRX containing BBa_K863000 (08/31). 3 L fermenter (Infors), 37 °C, pO2 of 50 % for 12 hours. Purification of the supernatant via HisTrap column. The red arrow shows BPUL in lane 5-8 (elution 1 and 2).
Figure 1: First fermentation of E. coli KRX containing BBa_K863000 (08/31). 3 L fermenter (Infors), 37 °C, pO2 of 50 % for 12 hours. Purification of the supernatant via HisTrap column. The red arrow shows BPUL in lane 5-8 (elution 1 and 2).- Performed the cell disruption of yesterday's flask cultivation to produce CBD-GFP+His fusion protein via sonification and made a SDS-Page of it. The problem is to purify this product, because it might be impossible to remove the protein from the column. If it binds via CBD and not via His-tag.
- Made a flask cultivation of E. coli KRX containing new BioBricks: Laccase from E. coli, B. pumilus and BBa_K863012 behind a constitutive promoter.
- Settings: 100 mL flask without baffles, autoinduction medium, 60 µg/mL chloramphenicol, final volume: 60 mL, 37 °C, 120 rpm, 12 hours, double determination.
- We repeated the purification of His-tagged GFP by using the flow-through and the washing fraction, but it did not work. Then we got to know that it could not work, because the sequencing showed us that the GFP was not tagged to His.
- Made a SDS-Page of the fractions of purified B. pumilus laccase produced at 08/31.
- Starting of another fermentation of E. coli KRX with BBa_K863000.
- Settings: fermenter: Braun, autoinduction medium, 60 µg/mL chloramphenicol, 3 L, 37 °C, stirrer on cascade to hold a pO2 of 50 %, airflow: 2 NL/m, 6 hours. Found out that we got a maximal optical density after 4 hours, after which the cells start to die. Therefore we harvested the cells after 6 hours.
- Aim: Getting a growth kinetics.
- Made preculture of E. coli KRX containing BBa_K863005
- Team Activity Tests:
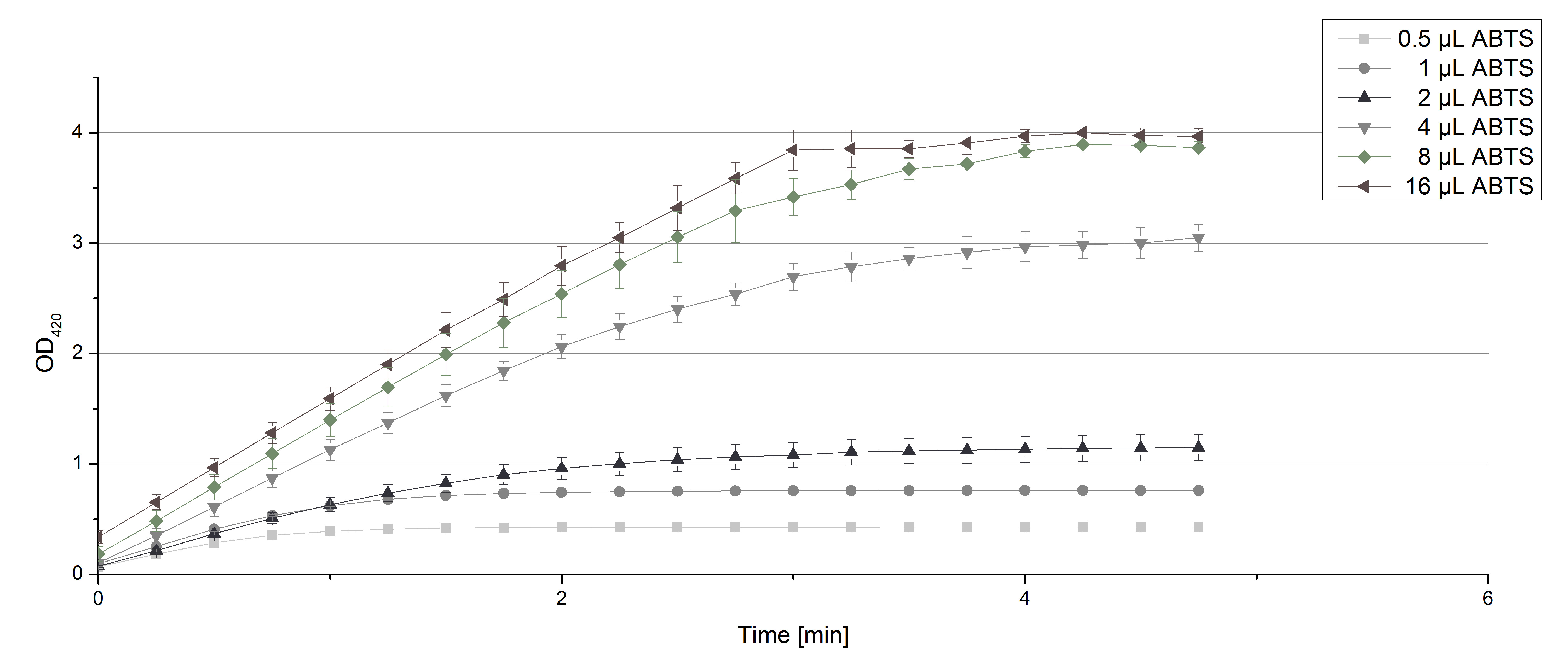
- Today we had to deliver some data for Team Modeling. To calculate enzyme kinetics of our laccases the initial reaction speed has to be determined. With our usual approach using 0.1 U TVEL0 Laccase we couldn't accurately measure the first minute of reaction with ABTS because everything went to fast. So we cut down the volume of TVEL0 laccase from 140 µL to 35 µL (0.025 U) and measured the oxidizing potential of TVEL0 with different ABTS concentrations. The results are shown in Fig. 1. There you go, Team Modeling! Make the best out of it!
- Team Substrate Analytic:
- The degradations were stopped with methanol. The samples were evaporated for six hours and diluted in 50 % - 50 % acetonitril - water. Then we used the C18E-columns for purifying the samples. The measuring with LC-MS was done from Marcus Persicke.
- It´s time to regenerate the column. Dominik showed us the protokol.
Thursday September 6th
- Team Cloning of Bacterial Laccases:
- Colony PCR on J23100 + J61101 + bhal_HIS showed four positive colonies which were plated on LB with CM + AMP.
- Plating colonies from colony PCR on bpul+ pSB1C3 and tthl + pSB1C3 on new plates.
- We still have to wait for the sequencing results but control restrictions suggests that we have tthl_HIS, bhal_HIS, ecol_HIS and bpul_HIS in a vector with a constitutive promoter. So we transformed the plasmids in E.coli Rosetta Gami 2. The cells carry a chloramphenicol-resistant plasmid, pRARE2, which supplies tRNAs for seven rare codons, AUA, AGG, AGA, CUA, CCC, GGA and CGG under the control of their native promoter.
- Team Cellulose Binding Domain:
- gradient-PCRs to generate CBDcex_Freiburg and CBDclos_Freiburg on the orignial plasmids with the CBDcex_Freiburg + CBDclos_Freiburg and 2AS_Freiburg-Reverseprimers. Only the CBDcex_Freiburg-product could be generated (68,5°C is best).
- two colonies of const.GFP_His plated on selection-agar for plasmid-isolation.
- Team Cultivation & Purification:
- Made a SDS-Page of cultivation from 08/31.
- Fermentation of E. coli KRX containing BBa_K863005 to get a growth kinetics
- Settings: fermenter: Infors, autoinduction medium, 60 µg/mL chloramphenicol, 3 L, 37 °C, stirrer on cascade to hold a pO2 of 50 %, airflow: 2 NL/m.
- Preparation of the bigger fermenter (7 L) for the first time
- Performing the cell disruption via sonification and the purification via HisTrap column supported by Äkta of the flask cultivation from 09/05.
- We got to know that the working group of physical chemistry has another homogeniser, that we could use. We performed the cell disruption via high-pressure homogeniser of fermentations from 09/03, 04 and 05.
- Made preculture of E. coli KRX containing BBa_K863005
- Team Substrate Analytic:
- The fluorescent lamp is broken. So it was not possible to mesure today.
Friday September 7th
- Team Cloning of Bacterial Laccases:
- Plasmid isolation of different plated colonies and control digest. The restriction showed positive results for the plasmids for bpul+pSB1C3, tthl + pSB1C3 and bhal_His + J23110 + RBS + pSB1C3. So we sent the plasmids for sequencing.
- Transformation of <partinfo>BBa_K863012</partinfo> and <partinfo>BBa_K863022</partinfo> in E. coli Rosetta-Gami 2.
- Team Shuttle Vector:
- Designed and ordered sequencing primers for the shuttle vector. We set a primer every 700 to 750 bases.
- Team Cultivation & Purification:
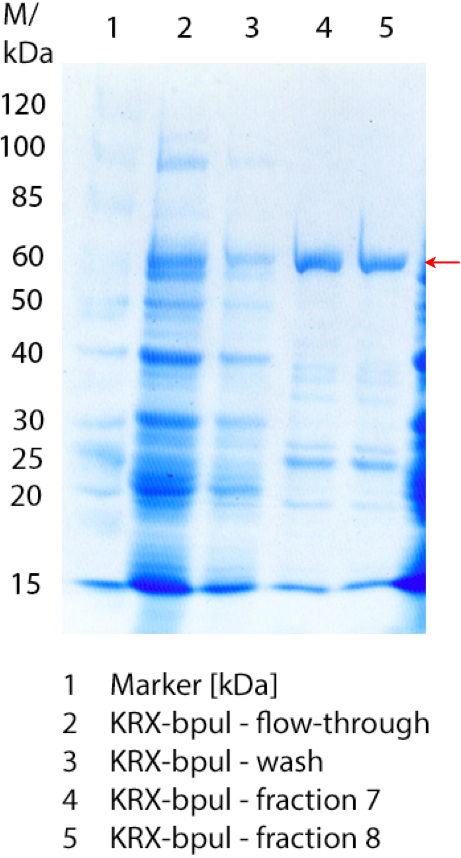
- Today we performed lots of purifications. This time we used a Ni-NTA column, because it seemed to work better than the TALON column. Therefore it was reloaded newly before starting the purifications. The purified samples were BPUL produced on 09/03 and 09/05 as well as ECOL produced on 09/04. The results were not that promising, so maybe we harvested the cells too early. They start dying more or less when we expect them to produce our (toxic) protein. So we think we had to let some of them die, so that the rest will produce our protein. The next fermentations should endure at least 12 hours.
- Fermentation of E. coli KRX containing BBa_K863005 within the Bioengineering NFL22 fermenter.
- Settings: fermenter: Biostat NFL22 (7 L), autoinduction medium, 60 µg/mL chloramphenicol, final volume: 6 L, 37 °C, stirrer increased 2 % if the pO2 got below 30 %, airflow: 5 NL/m, 12 hours.
Saturday September 8th
- Team Cultivation & Purification:
- Harvesting and centrifugation of fermentation of 09/07. Pellet stored at 4 °C.
- Made precultures of E. coli KRX containing BBa_K863005, BBa_K863000 and of E. coli Rosetta-Gami 2 containing BBa_K863012 and laccase from B.halodurans <partinfo>BBa_K863022</partinfo>.
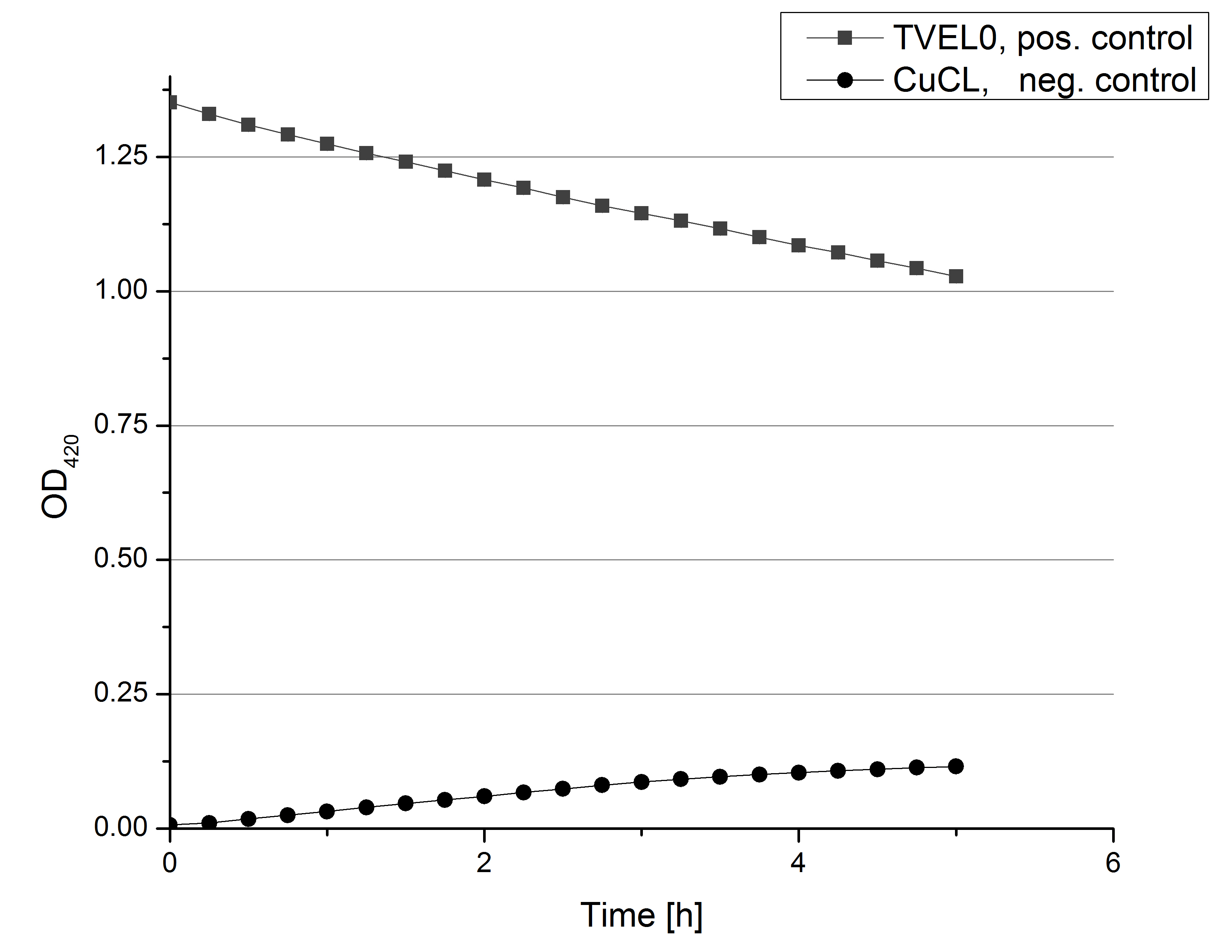
- Team Activity Tests: Today we managed to rebuffer the BPUL samples given by Team Cultivation. We got fraction 3 to 7 and 15 to 19 of the purification, because they have shown a nice peak. After bringing the BPUL laccases into water, we incubated with 0.5 mM CuCl2 and measured the protein concentration via Bradford. The offered fractions contained the following protein concentrations after rebuffering: 0.0087 mg/mL (fr. 3), 0.0110 fr. 4), 0.0632 (fr. 5), 0.1672 (fr. 6), 0.1229 (fr. 7), 0.0060 (fr. 15), 0.0034 (fr. 16), 0.0326 (fr. 18), 0.0225 (fr. 19). The protein amount in fraction 17 couldn't be measured but we didn't leave it behind. We adjusted the protein concentrations to the concentration of TVEL0, but did also measurements using 140 µL of each sample, because the protein concentrations do not only represent gained laccases. The results of our plain activity measurement of the samples are demonstrated in Fig. 1 and Fig. 2 (positive and negative control shown in Fig. 3):
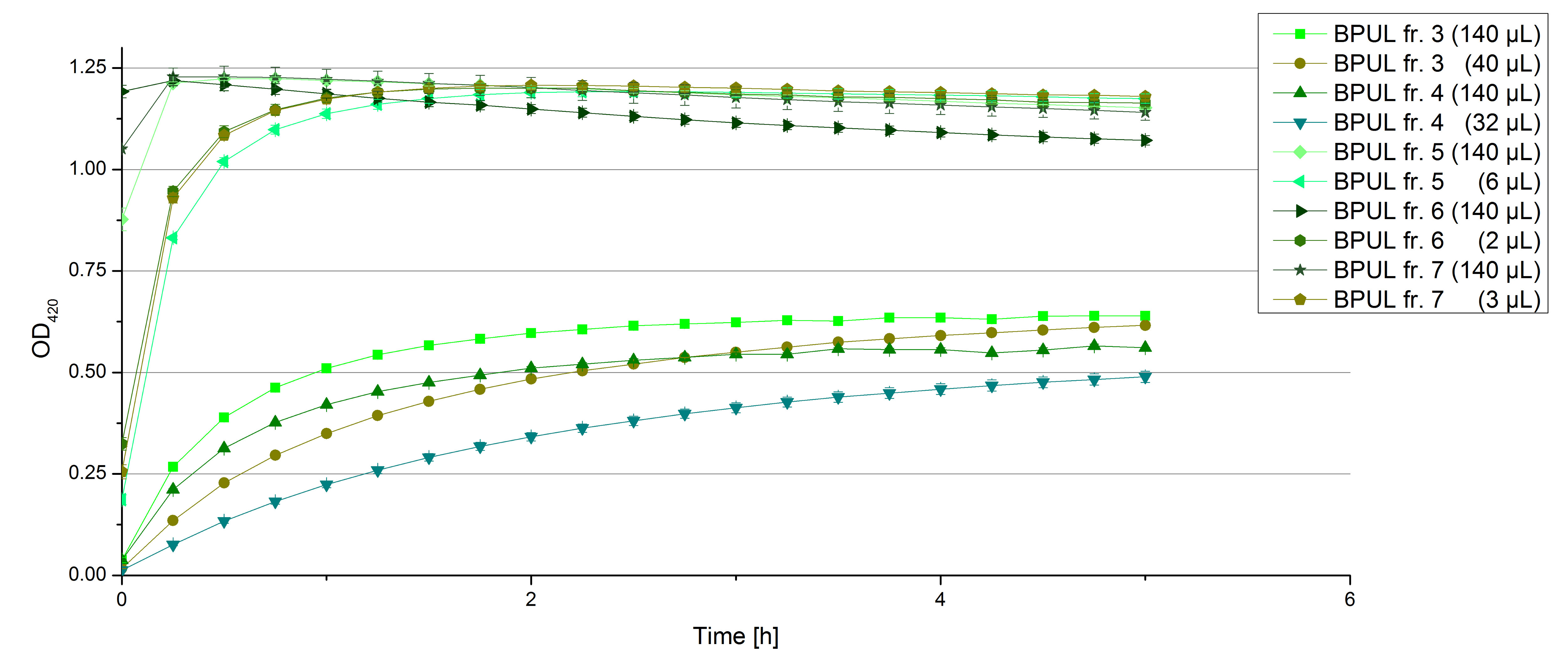
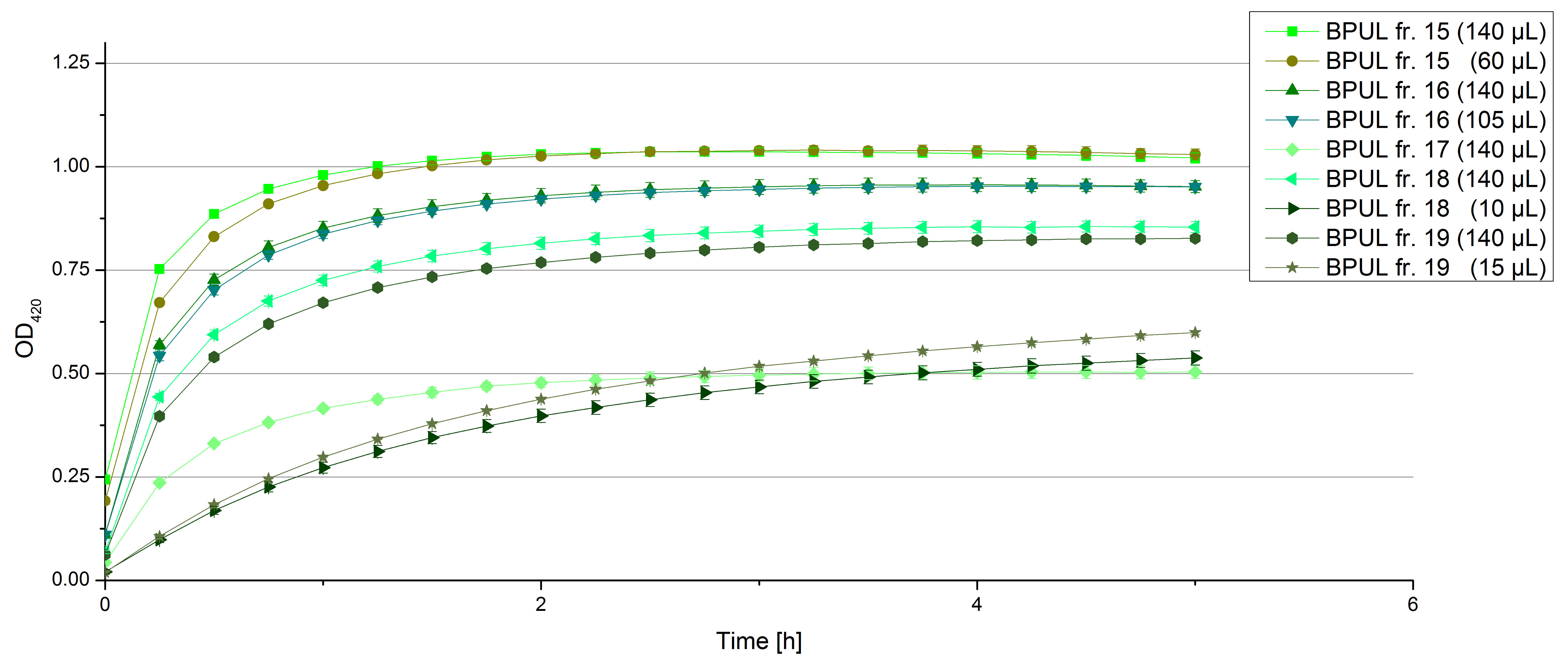
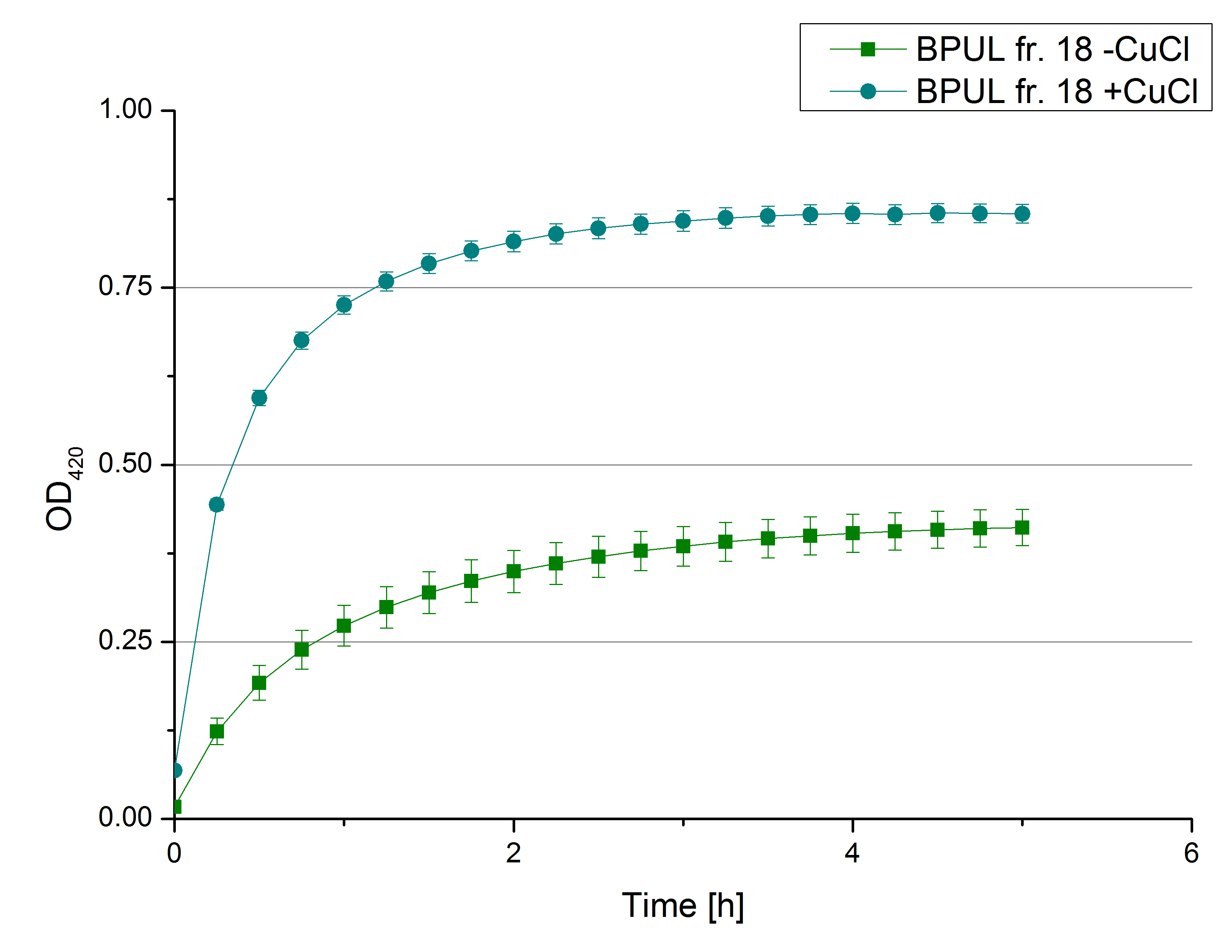
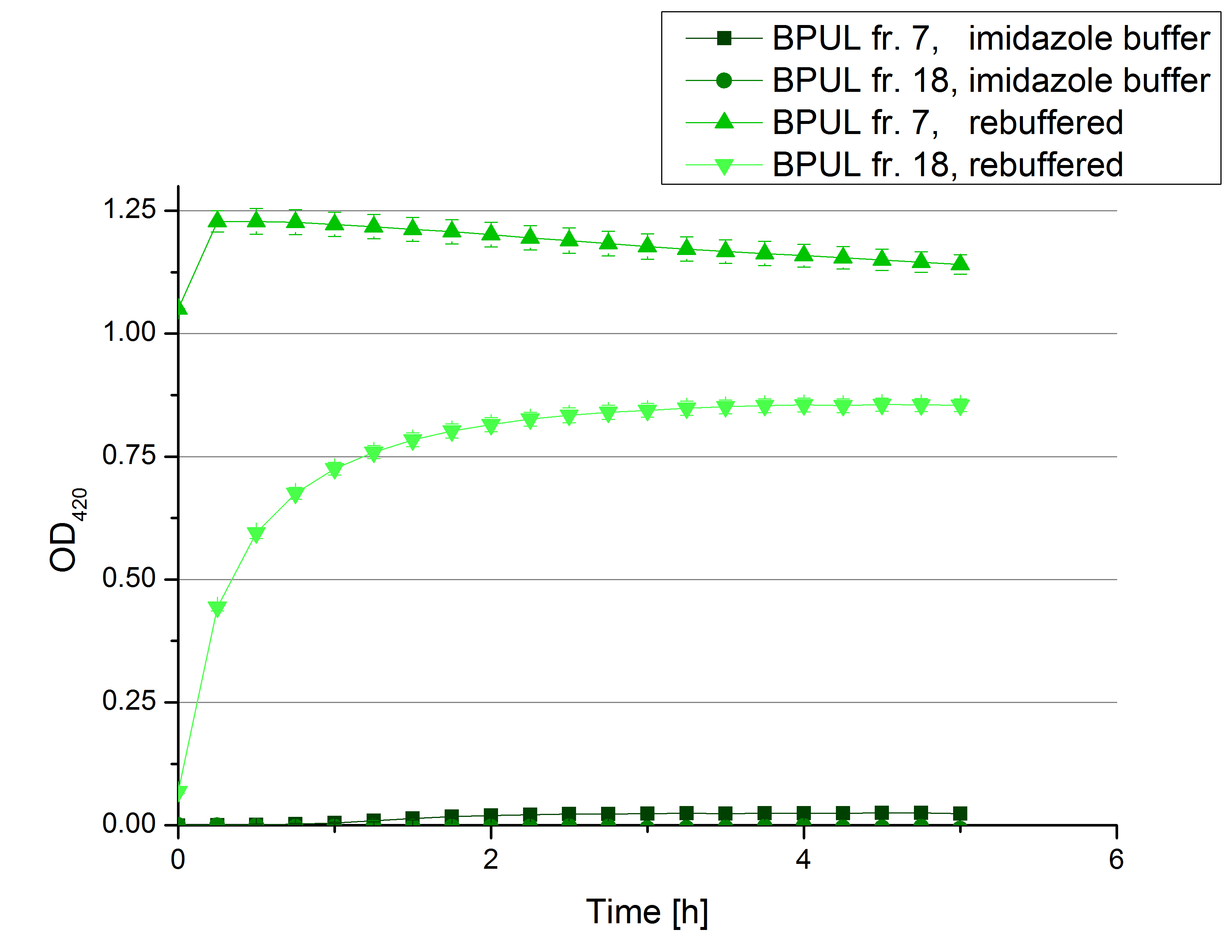
Fractions 6 and 7 show the highest activity leading to the assumption, that faction 7 contains as much protein as fraction 6 and that this amount is almost represented through BPUL laccase. Additionally we were interested in the impact on copper incubation. We have chosen fr. 18 and compared the activity of a copper incubated and a not copper incubated sample. With a incubation time of 2 hours with 0.5 mM CuCl2 we can double the activity of the BPUL laccase (see Fig. 4). Another interesting topic was the impact of imidazole buffer on the BPUL activity. For that we tested fr. 7 and fr. 18 in imidazole buffer and rebuffered in H2O. The results we got were unambiguously: BPUL in imidazole buffer didn't show much activity. Unfortunately we can't omit the rebuffering.
Sunday September 9th
- Team Cultivation & Purification:
- Fermentation of E. coli KRX containing BBa_K863005 (fermenter: Infors) or containing BBa_K863000 (fermenter: Braun) to get a growth kinetics.
- Settings: autoinduction medium, 60 µg/mL chloramphenicol, final volume: 3 L, 37 °C, stirrer on cascade to hold a pO2 of 50 %, airflow: 2 NL/m, 12 hours.
- Harvesting and centrifugation of today's 3 L fermentations.
- Fermentation of E. coli KRX containing BBa_K863000
- Settings: fermenter: Bioengineering NFL22 (7 L), autoinduction medium, final volume: 6 L, 37 °C, pH 7 stirrer increased 2 % if the pO2 got below 30 %, airflow: 5 NL/m, 12 hours.
- Flask cultivation of E. coli Rosetta-Gami 2 cells containing BBa_K863012 and <partinfo>BBa_K863022</partinfo>.
- Settings: flask without baffles, LB medium, 200 mL, 60 µg/mL chloramphenicol and 300 µg/mL ampicillin, 37 °C, 120 rpm. We cultivated for 48 hours and stored the culture then overnight at 4 °C.
- Fermentation of E. coli KRX containing BBa_K863005 (fermenter: Infors) or containing BBa_K863000 (fermenter: Braun) to get a growth kinetics.
| 55px | | | | | | | | | | |
 "
"