Team:Bielefeld-Germany/Labjournal/week16
From 2012.igem.org
Contents |
Week 16 (08/13 - 08/19/12)
weekly meeting
- As part of our collaboration the iGEM team from SDU-Denmark will arrive on saturday (25th of august).
- Our StreetScience booth will have 16 m2.
- We already ordered all info-material we want to provide at StreetScience.
- Just in case we are going to qualify for the world finals in Boston, everyone needs a visa. Especially our team members without a european nationality have to inform about visa terms.
- Derya is responsible for planning a new, project-driven video.
- As deadlines come closer we have to think about a team name.
- He have decided to rent a bus to travel to Amsterdam.
Monday August 13th
- Team Cloning of Bacterial Laccases:
- After cleaning up the gradient PCR from the 09th August we did an digestion with the S. goettingen DNA for cloning it into the pSB1C3. The pSB1C3 was additionaly treated with Shrimp-alkaline-phosphotase. After this we set the ligation of insert DNA and vector DNA.
- Team Cultivation & Purification:
- Today we discussed that we may get activity if we start to purify our laccases. So from now we will purify our products before measuring them. We hope this will bring promising results. Therefore we searched for a useful method.
- Team Site Directed Mutagenesis:
- Gradient-PCR of tvel10 55 to 66°C with DMSO (12 Steps) resulted in a lot of product at 55°C and 63°C to 66°C; a little product at 56°-59° and 61°-62°C and no product at 60°C (59°C was the temperature Clonemanager predicted and I used before). Merged the products and cleaned them up.
- Digested the product with EcoRI, PstI and DpnI.
- Team Cellulose Binding Domain:
- We did another PCR on the <partinfo>I13522</partinfo> with the GFP_His-primers with positive result.
- Plasmid-isolation of two positive clones of CBDclos(T7) followed by digestion with NotI that confirmed that the plasmid is CBDclos(T7). Prepared both for sequencing.
- Plated three positve CBDcex(T7)-colonies.
- Restriction of the GFP_His-PCR-product with EcoRI and PstI to bring it into the pSB1C3-backbone.
- Also: Restriction of the CBDcex(T7)-PCR-product with NgoMIV and PstI for an assembly.
- Restriktion of the CBDcex(T7)-PCR-product with AgeI and EcoRI for the assembly.
- All digestions were done with DpnI to get rid of the templates.
- Ligation of GFP_His with the pSB1C3-backbone.
- Ligation of CBDcex(T7) and GFP_His with the pSB1C3-backbone.
- Transformed all ligations into KRX.
Tuesday August 14th
- Team Cloning of Bacterial Laccases: We finally had positive bacteria for the S. goettingen. So we decide to do a colony PCR. But the result has shown that there were no insert DNA but we didn't want to give it up so we transfered some colonies to a new plate an let them grown over night again.
- Team Fungal and Plant Laccase: Today our cDNA took a bath in ethanol and got cleaned. We are pretty sure our next PCR will be a lot more successful. Check protocols for further information about the cDNA washing via ethanol precipitation.
- Team Cultivation & Purification:
- Made preculture of E. coli KRX without plasmid and with BBa_K863020, BBa_K863000, BBa_K863005, BBa_K863010 and BBa_K863015 as well as BBa_K525710.
- Team Site Directed Mutagenesis:
- Ligation of tvel (PCR-product) and pSB1C3 and transformation into KRX
- Gradient-PCRs (55°C bis 72 °C) with xccl-plasmid using xccl-g3633c and xccl-g2247c primer-mixes, respectively, resulted in no product of the right size.
- Team Cellulose Binding Domain:
- Sequencing results showed, that the insertion of GFP_His wasn't successful at all, CBDcex(T7) had a lot of mutations and CBDclos(T7) had one silent mutation.
- On the GFP_His-transformation-dish did not grow any colony. Did a new transformation with a little more ligation-product and plated the cells on selection-agar.
- Plasmid-isolation of the three positive CBDcex(T7)-plasmids and digestion showed not the correct fragments. Plated some more positive colonies on selection-agar for plasmid-isolation.
- The CBDcex(T7)+GFP_His-dish had only nine colonies and a colony-PCR showed that none of them had the right insert.
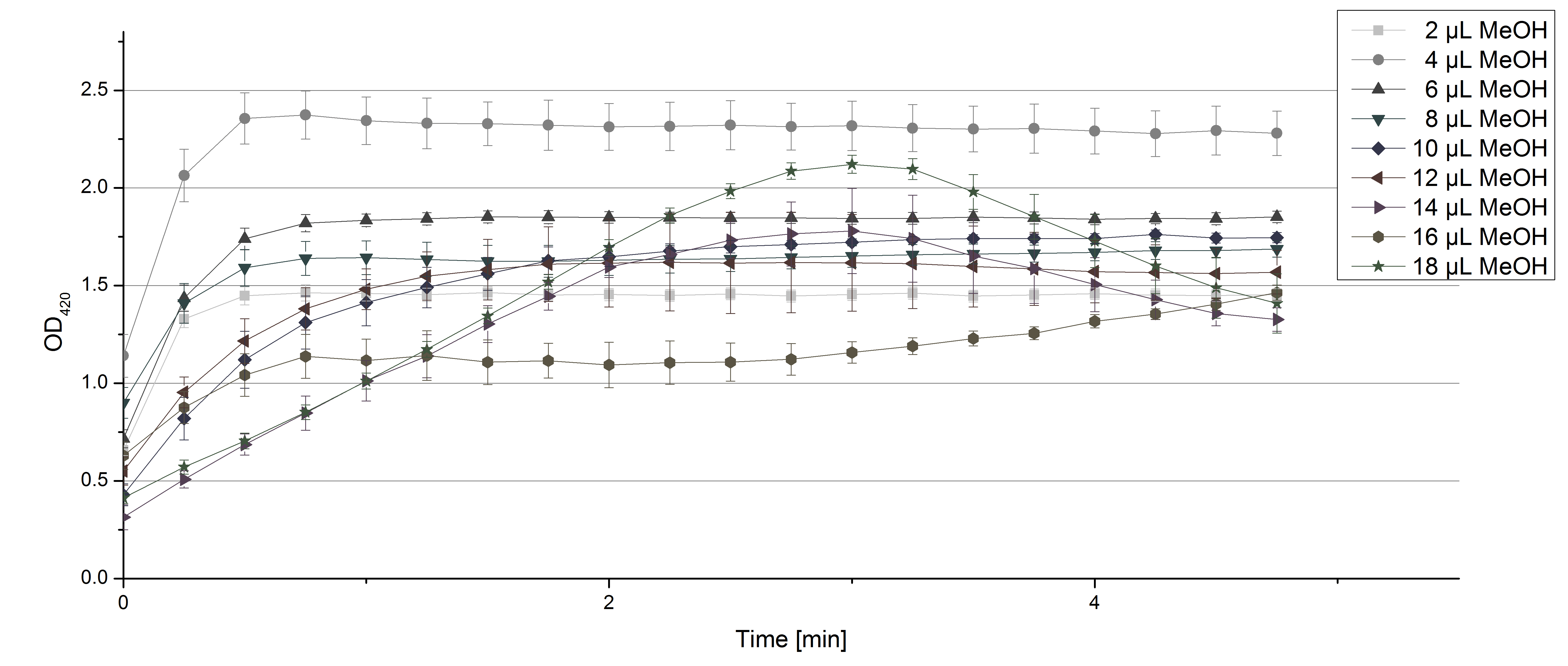
- Team Activity Tests: Again our skills in measuring laccase activity were in demand today. We had a job incoming from Team Substrate Analytic. Since they need to dissolve their substrates in methanol or acetonitrile, they were curious about laccase activity related to these solvents. And they infected us with their curiosity! So we started measurements using 0.1 U TVEL0 Laccase, 40 µL sodium acetate buffer and different amounts of MeOH or acetonitrile respectively, ad 200 µL H2O. Our tested range went from 2 µL to 18 µL. The results show that MeOH and acetonitril have an impact on the ability of TVEL0 to oxidize ABTS. Using 16 µL MeOH (which means 8% of the amount of the sample contained MeOH) led to a different saturation curve as usual. The saturation couldn't be reached any more (see Fig. 1). Acetonitrile had a noticeable impact when using 5% (14 µL) of it in relation to the sample (see Fig. 2). But in the end both solvents couldn't stop TVEL0 laccase from being active. We are happy to tell Team Substrate Analytics that they can proceed as planned and dissolve the substrates in the tested solvents.
- Team Substrate Analytic:
- Dominic helped us to set the HPLC method based on the paper „Enzyme-Catalyzed Oxidation of 17β-Estradiol Using Immobilized Laccase from Trametes versicolor”.
- We prepared dilution series from three different estrogens and tryed to mesure them. Sadly all substrates left the column with the same retention time as the solvent.
Wednesday August 15th
- Team Wiki: We set up some more wiki rules. Today´s rules were about citations. We agreed von some standards that will for sure help make the wiki a little prettier. Have a look:
- Paper: Exampleman M et al. (2002). The example paper. The example journal (Volume): pages.
- Website: Name of website, URL, Index, date site visited
- Book: Examplewomen M et al. (1999). The example book. The example publisher (edition).
Next will be a standard for charts.
- Team Cloning of Bacterial Laccases:
- Plasmid isolation from the plate of the 14th August. This was done with the Analytic Jena Kit.
- Team Cultivation & Purification:
- Flask cutivation of E. coli KRX with BBa_K863020, BBa_K863000,BBa_K863005, BBa_K863010 and BBa_K863015. We used E. coli KRX negative control as well as E. coli KRX with BBa_K525710 as positive control.
- Settings: 1 L flasks without baffles, final volume: 250 mL, autoinduction medium supplemented with 60 µg/mL chloramphenicol, 37 °C, 120 rpm, single determination
- Flask cutivation of E. coli KRX with BBa_K863020, BBa_K863000,BBa_K863005, BBa_K863010 and BBa_K863015. We used E. coli KRX negative control as well as E. coli KRX with BBa_K525710 as positive control.
- Team Site Directed Mutagenesis:
- Colony-PCR of tvel10-colonies resulted in small bands. Quickly explained: tvel10 still had illegal PstI-restrictions-sites. Digestion of tvel10 PCR-product with NotI as well as digestion of RFP-pSB1C3 with NotI
- Team Cellulose Binding Domain:
- Plasmid-isolation of the three positive CBDcex(T7)-colonies, but digestion showed no positive result, again.
- Colony-PCR of GFP_His and CBDcex(T7)+GFP_His did not show any positive result, ether.
- Team Substrate Analytic:
- Dominik tryed to regenerate the column. After that we mesured new dilution series from all three estrogens. Once again all substrates left the colmun together with the solvent. That means the column is defect.
Thursday August 16th
- Team Cultivation & Purification:
- Today we harvested the cells from the cultivation on 08/15, disrupted them in equilibration buffer via sonification, centrifugated and purified the supernatant via HisTrap column. The purified samples in 500 mM Imidazol were given to the activity test team.
- Team Site Directed Mutagenesis:
- Clean-up of tvel10 and pSB1C3 (both cut with NotI); Dephosphorilation of pSB1C3 with SAP and ligation. Transformed into KRX and plated on select-agar.
- Team Cellulose Binding Domain:
- Colony-PCR of CBDcex(T7)+GFP_His-clones with no result, even the positive-control was negative, so we plated three colonies for a test-digestion instead.
- Digestion of CBDclos(T7) with AgeI and PstI and digestion of the GFP_His-PCR-product with NgoMIV and PstI over-night.
Friday August 17th
- Team Cellulose Binding Domain:
- Stopped digestion of CBDclos(T7) and GFP_His. Dephosphorylated pSB1C3+CBDclos(T7).
- Plasmid-isolation of the three CBDcex(T7)+GFP_His-dishes.
- The sequencing of the two CBDclos(T7)-plasmids showed the same result as the first: one silent mutation. We decided to carry on with this plasmid.
Saturday August 18th
- Team Cloning of Bacterial laccases:
- Today we performed a ligation from the digestions we have done before. The ligation included of course the T7 promotor, the backbone pSB1C3 and the individual laccase genes. Ligation was performed with T4 ligase for 20 minutes at room temperature and stopped through a five minute incubation at 70 °C.
- Team Cellulose Binding Domain:
- Ligated CBDclos(T7) and GFP_His, transform into KRX and plated on CM-selection-Agar.
Sunday August 19th
- Team Site Directed Mutagenesis:
- Colony-PCR of tvel10-colonies with three positive result.
- Plated those three on select-agar for plasmid-isolation.
- Team Cellulose Binding Domain:
- Colony-PCR of colonies from the CBDclos(T7)+GFP_His-trafo-dish with only negative results.
| 55px | | | | | | | | | | |
 "
"