Team:Buenos Aires/Results/DevicesTesting
From 2012.igem.org
Manugimenez (Talk | contribs) (→Protocol) |
Manugimenez (Talk | contribs) (→Plasmids) |
||
| Line 6: | Line 6: | ||
== Experimental Setup == | == Experimental Setup == | ||
| - | === Plasmids === | + | === Plasmids and BBs === |
In order to construct the yeast expression plasmids we choosed 3 vectors, 2 with a tryptophan marker and 1 with an histidine marker: | In order to construct the yeast expression plasmids we choosed 3 vectors, 2 with a tryptophan marker and 1 with an histidine marker: | ||
Revision as of 02:51, 27 October 2012

Contents |
Devices testing
Experimental Setup
Plasmids and BBs
In order to construct the yeast expression plasmids we choosed 3 vectors, 2 with a tryptophan marker and 1 with an histidine marker:
- pCM182/5, which are centromeric plasmids with TRP1 marker, and with a doxycycline repressible promoter [Gari et al 1996].
- pEG202, with a 2 ori, HIS3 marker and a constitutive promoter (PADH1).
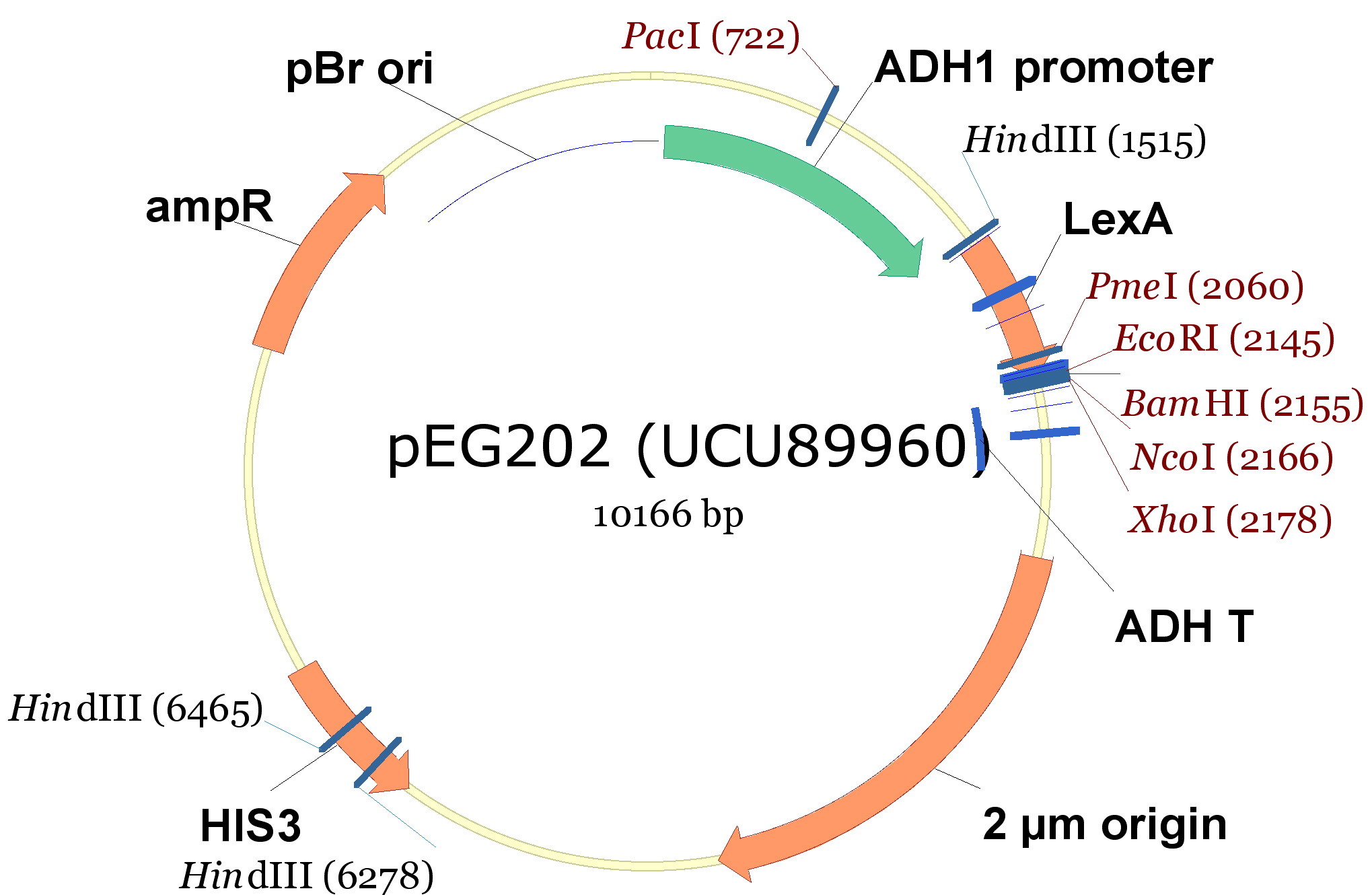
| 340px |
The cloning we did was:
| <partinfo>BBa_K792009</partinfo> (PoliHa) | <partinfo>BBa_K792010</partinfo> (TRPZipper2) | <partinfo>BBa_K792011</partinfo> (PoliHb) | <partinfo>BBa_K792012</partinfo> (PoliWb) | |
|---|---|---|---|---|
| pCM182 (TRPa) | X | X | ||
| pCM185 (TRPb) | X | X | ||
| pEG202 (HIS) | X | X |
Protocol
- Digestion of plasmids and TRP/HIS export device:
- pCM182; pCM 185; BBa_K792010 y BBa_K792012 were digested with BamHI and Pst1 restriction enzymes.
- pEG202; BBa_K792009 y BBa_K792011 were digested with BamHI and Not1.
- Purification of digested vectors (pCM185; pCM182; pEG202)
- Ligation of vectors and devices according the anterior table (T4 ligase protocol, overnight)
- Transformation of E. Coli DH5a with the ligation products. Bacterias were plated on LB-agar with Ampicillin, and incubated over night at 37 °C.
- Colonies were used for liquid cultures (LB + Ampicillin) and minipreps were made.
- Constructions (vector + insert) were checked by digestion with restriction enzymes, and 1%-agarose gel (1kb and 100bp as markers).
Yeast strains
Once obtained the desired constructions, we transformed yeast strains:
- TCY3081: W303, bar1-, ura3::PAct1-YFP
- TCY3128: W303, bar1-, leu2:: Pprm1-CFP 405
Final transformed strains
We got the following transformed strains:
|
|
|
|
Measurement of Trp in medium and Basal Production
To check the efectiveness of our biobricks, we must first determine the ammount of tryptophan secreted by natural strains to the medium, so we can compare. With that end in mind, we designed a protocol for measurement of tryptophan in medium, based in its fluorescense at 350nm, when excited with 295nm light. As a previous step, we checked that none of the other aminoacids used in the medium interferes, by graphically comparing the spectres for uncomplemented medium and medium complemented with leucine, uracile and histidine, at an appropiate range.
To determine Trp concentration, we must first have a way to transform our readings (intensity) to a more useful output, so we made a calibration curve, through serialized 1:2 dilutions of our medium, which Trp's concentration is 50mg/mL, until approximately constant intensity.
The procedure to measure secretion rates will be growing the strain from a known OD in exponential growth phase in -T medium and plotting it's OD over time, spin-drying at time=t, retrieving the supernatant's Trp concentration and dividing it by the integral of OD vs. time between time=0 and time=t, so we get to a rate which will be proportional to the number of cells in the culture, which means we can actually compare between different strains. Since our medium is free from Trp, all of it should come from within the cells, and if the culture is growing at exponential rates, lysis should be negligible, so the only explanation would be cells exporting their own Trp.
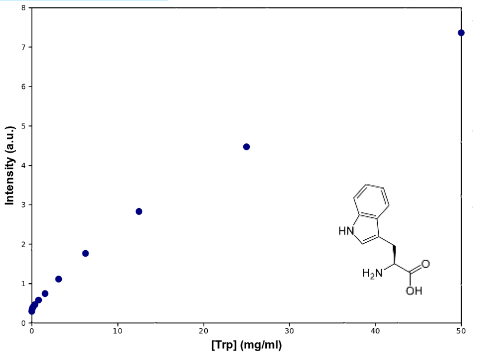
|
| Graph:Tryptophan calibration curve |
Results
As can be seen from the graph the screening of the concentration of the Trp in medium describes an almost lineal function. Through this experiment we can be sure that we would be able to measure increase of Trp in medium as it is exported from the cells, within the biological range of export. The sensitivity of this method seems to be enough to detect concentrations as low as ~0.02mg/mL, and as high as 50mg/mL, maybe more. Since our medium is 50mg/mL, we assume that's the saturation point of the curve. If we get bigger intensities than the one corresponding to it, we will dilute the sample.
Because of time constraints, we haven't been able to check the method with either our designed strains nor the non-exporting ones.
 "
"