Team:TU Darmstadt/Modeling Docking
From 2012.igem.org
| (37 intermediate revisions not shown) | |||
| Line 27: | Line 27: | ||
<li><a href="/Team:TU_Darmstadt/Safety" title="Safety">Safety</a></li> | <li><a href="/Team:TU_Darmstadt/Safety" title="Safety">Safety</a></li> | ||
<li><a href="/Team:TU_Darmstadt/Downloads" title="Downloads">Downloads</a></li></ul></li> | <li><a href="/Team:TU_Darmstadt/Downloads" title="Downloads">Downloads</a></li></ul></li> | ||
| - | <li><a href="/Team:TU_Darmstadt/Human_Practice" title="Human Practice">Human Practice</a | + | <li><a href="/Team:TU_Darmstadt/Human_Practice" title="Human Practice">Human Practice</a></li> |
| - | + | ||
| - | + | ||
| - | + | ||
<li><a href="/Team:TU_Darmstadt/Sponsors" title="Sponsors">Sponsors</a><ul> | <li><a href="/Team:TU_Darmstadt/Sponsors" title="Sponsors">Sponsors</a><ul> | ||
<li><a href="/Team:TU_Darmstadt/Sponsors" title="Sponsors">Overview</a></li> | <li><a href="/Team:TU_Darmstadt/Sponsors" title="Sponsors">Overview</a></li> | ||
| Line 51: | Line 48: | ||
==Docking== | ==Docking== | ||
| - | |||
===Goal=== | ===Goal=== | ||
While PET.er is degrading PET into its monomers various chemical compounds like polymer plasticizer are released. Since PET is itself a waste material, it will never be in a pure state. Due to this, it is important to study possible interaction of plasticizer as well as PET with the degradation machinery. Hence an experimental design will give a deeper understanding of the interaction of our enzymes with additives. | While PET.er is degrading PET into its monomers various chemical compounds like polymer plasticizer are released. Since PET is itself a waste material, it will never be in a pure state. Due to this, it is important to study possible interaction of plasticizer as well as PET with the degradation machinery. Hence an experimental design will give a deeper understanding of the interaction of our enzymes with additives. | ||
===Methods=== | ===Methods=== | ||
| - | Due to the absence of forcefield terms of plasticizer we choose a global docking approach to study the interaction. For the docking simulations we uses the Autodock plugin of Yasara structure. Autodock is | + | Due to the absence of forcefield terms of plasticizer we choose a global docking approach to study the interaction. For the docking simulations we uses the Autodock plugin of Yasara structure. Autodock is a highly cited docking program developed at the Scripps Research Institute by Dr. Garrett M. Morris et al. [1] |
| - | + | ||
===Docking Protocol=== | ===Docking Protocol=== | ||
| - | For the docking approach we utilized the standard protocol dock_run.mcr and parallelized it up to six processors. Moreover, we | + | For the docking approach we utilized the standard protocol dock_run.mcr and parallelized it up to six processors. Moreover, we choose three additives out of the literature acting as the ligand within the simulations. |
*OET-dimeric | *OET-dimeric | ||
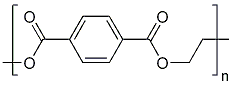
[[File: PET.png |250px|center]] | [[File: PET.png |250px|center]] | ||
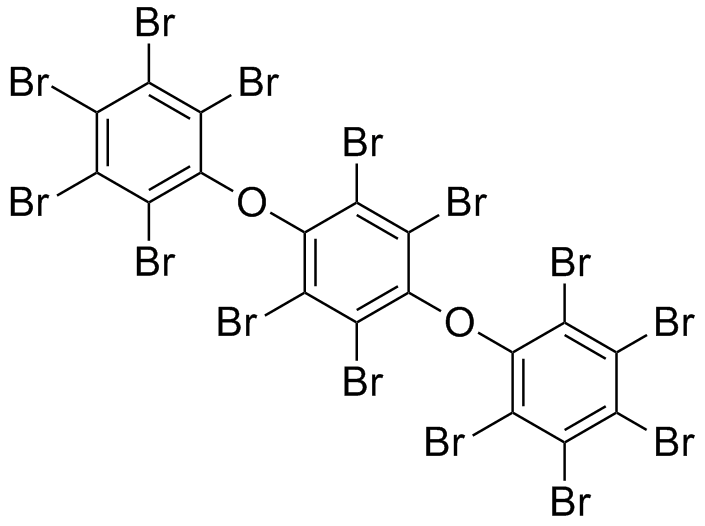
*S-120 | *S-120 | ||
| - | [[File: | + | [[File: S120neu.png|250px|center]] |
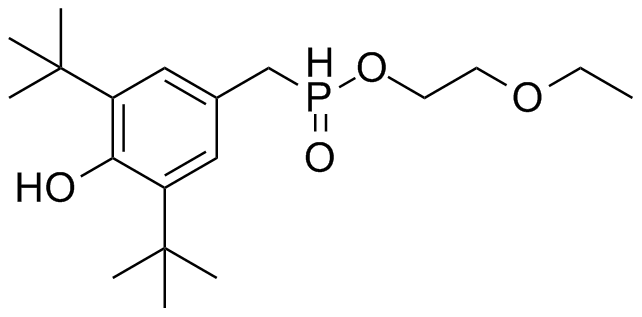
*AO-8 | *AO-8 | ||
| - | [[File: | + | [[File:Ao-8.png|250px|center]] |
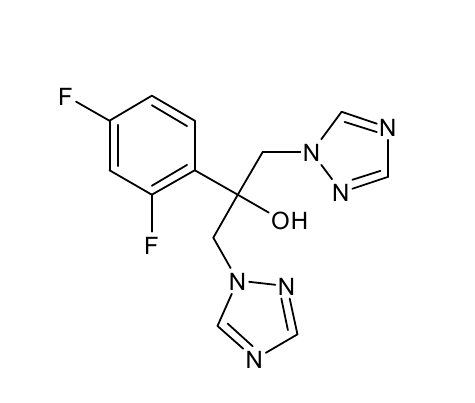
*TPF | *TPF | ||
[[File: TPF.png |250px|center]] | [[File: TPF.png |250px|center]] | ||
====Parameter for the Docking Experiment==== | ====Parameter for the Docking Experiment==== | ||
*Ligands = S120, AO-8, TPF, dimer OET | *Ligands = S120, AO-8, TPF, dimer OET | ||
| + | *Receptors = PnB Esterase 13 (homology model) and Fs. Cutinase (PDB: [http://www.rcsb.org/pdb/explore/explore.do?structureId=1CEX 1CEX]) | ||
*Number of docking runs = 200 | *Number of docking runs = 200 | ||
*ForceField AMBER03 | *ForceField AMBER03 | ||
| - | *Flexible Ligand | + | *Flexible Ligand |
| + | |||
===Analytics=== | ===Analytics=== | ||
| - | We extract binding energy from the atomic B-factors and the binding constant from the | + | [[File: S120.min.gif |right | 200 docking results from S120 with Fs. cutinase ]] |
| + | We extract binding energy from the atomic B-factors and the binding constant from the atomic property derived from all our calculations. We used scatter plots with the binding constant and the inhibition constant in order to compare the plasticizer as well as the PET substrate. We compared the binding energies as well as the inhibition constant from the best ranked docking runs. | ||
| + | |||
| + | Here, OET (dimeric state) became our reference substrate, meaning if an additive exhibits a higher inhibition constant it may effect our protein. Otherwise, if a binding energy exhibits a higher level than OET we can conclude that an additive may accumulate on our protein surface. Moreover, we compare the binding as well as the inhibition energies of the additives with itself to identify the strongest binder. | ||
| + | |||
| + | Since we know the active site of the Fs. Cutinase and the PnB Est. 13, it will be interesting if a plastizicer may dock there. In order to analyze that, we counted the residues which are involved within every docking simulation. Therefore, we measured the distance of every ligand to its corresponding receptor with an appropriate threshold and thus identified the residues. | ||
===Results=== | ===Results=== | ||
| + | |||
| + | ====Interaction with Fs. Cutinase==== | ||
| + | |||
| + | First we study the interaction of OET with the Fs. Cutinase. The following bar plot illustrates the occurrence of residues in direct contact with different ligand positions and conformations. | ||
| + | |||
| + | [[File: PET.fsc.hist.png |900px|center]] | ||
| + | |||
| + | Within the best runs we got a wide range of involved residues. Since we we set no surface constrains, this is typically for a global docking approach. Here, the residues of the active site, ASP 165 (4 times), SER 120 (2 times) and HIS 188 (2 times), occurs under the best ranked runs. Interestingly, other hydrophobic residues like ALA 164 and VAL 169 are among the high-scoring residues. Since these residues are located near the active site they may help to stabilize the PET Chains on top of the surface during the derogation process. It indicates that OET regularly docked in the active site. | ||
| + | |||
| + | Here we illustrate the docking results of S-120 with Fs. Cutinase: | ||
| + | |||
| + | [[File: S120.fsc.hist.png |900px|center]] | ||
| + | |||
| + | Only a few residues are highly involved during the docking simulation. We can conclude that S-120 may not effect the active site. For this example we identified possible accumulation sites: CYS 171, LYS 206 and LYS 168 . Although, we didn't identify active site residues, we measured ALA 164 and VAL 169 as one of the high-scoring residues. | ||
| + | |||
| + | Here we illustrate the the docking results of AO-8 with Fs. Cutinase: | ||
| + | [[File: AO8.fsc.hist.png |900px|center]] | ||
| + | Interestingly, we can observe nearly a uniform distribution of residue occurrence. Due to the hydrophobic nature of AO-8 it is obvious that it distributes ubiquity over the surface. | ||
| + | |||
| + | Here we illustrate the the docking results of TPF Fs. Cutinase: | ||
| + | [[File: TPF.fsc.hist.png |900px|center]] | ||
| + | Similar to the results of S-120, the active site residues are not identified under the best ranked docking results (accept for SER 120). | ||
| + | The scatter plots of the binding constant are stored in the gallery below: | ||
| + | <gallery widths="400px" heights="250px" perrow="2"> | ||
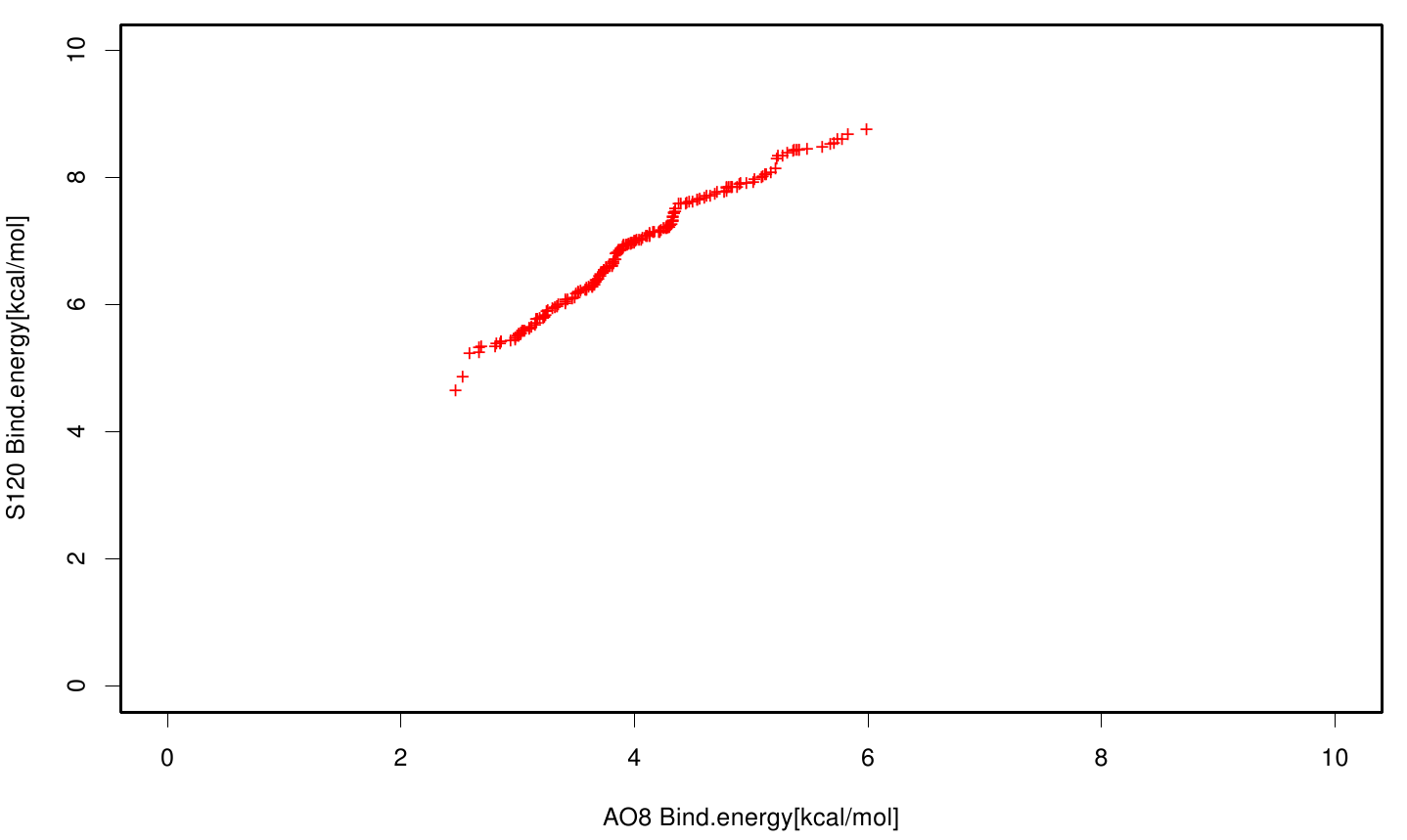
| + | File: AO8S120.fsc.scatter.bind.png | Scatter plot of AO8 vs. S120 | ||
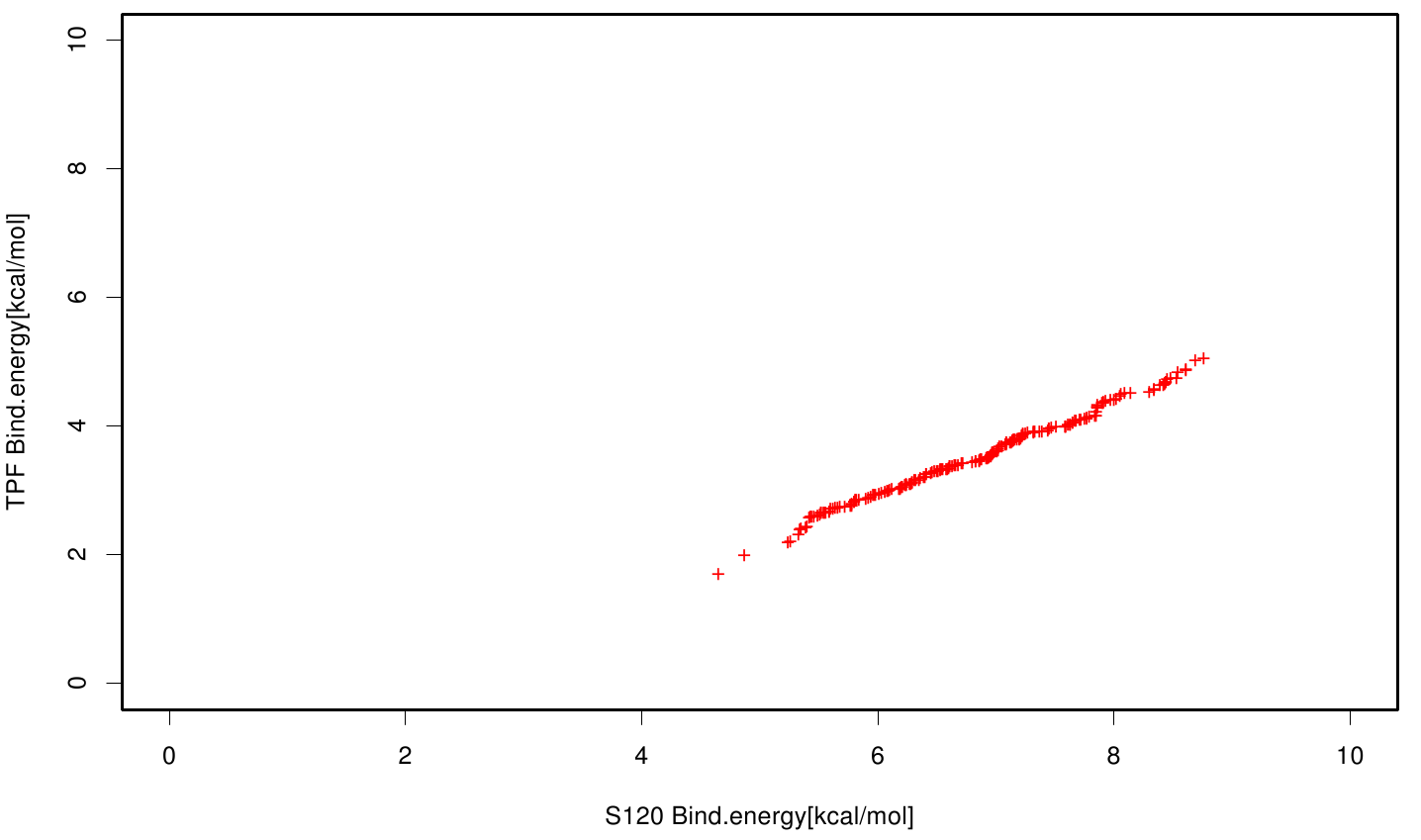
| + | File: S120TPF.fsc.scatter.Bind.enery.png | Scatter plot of TPF vs. S120 | ||
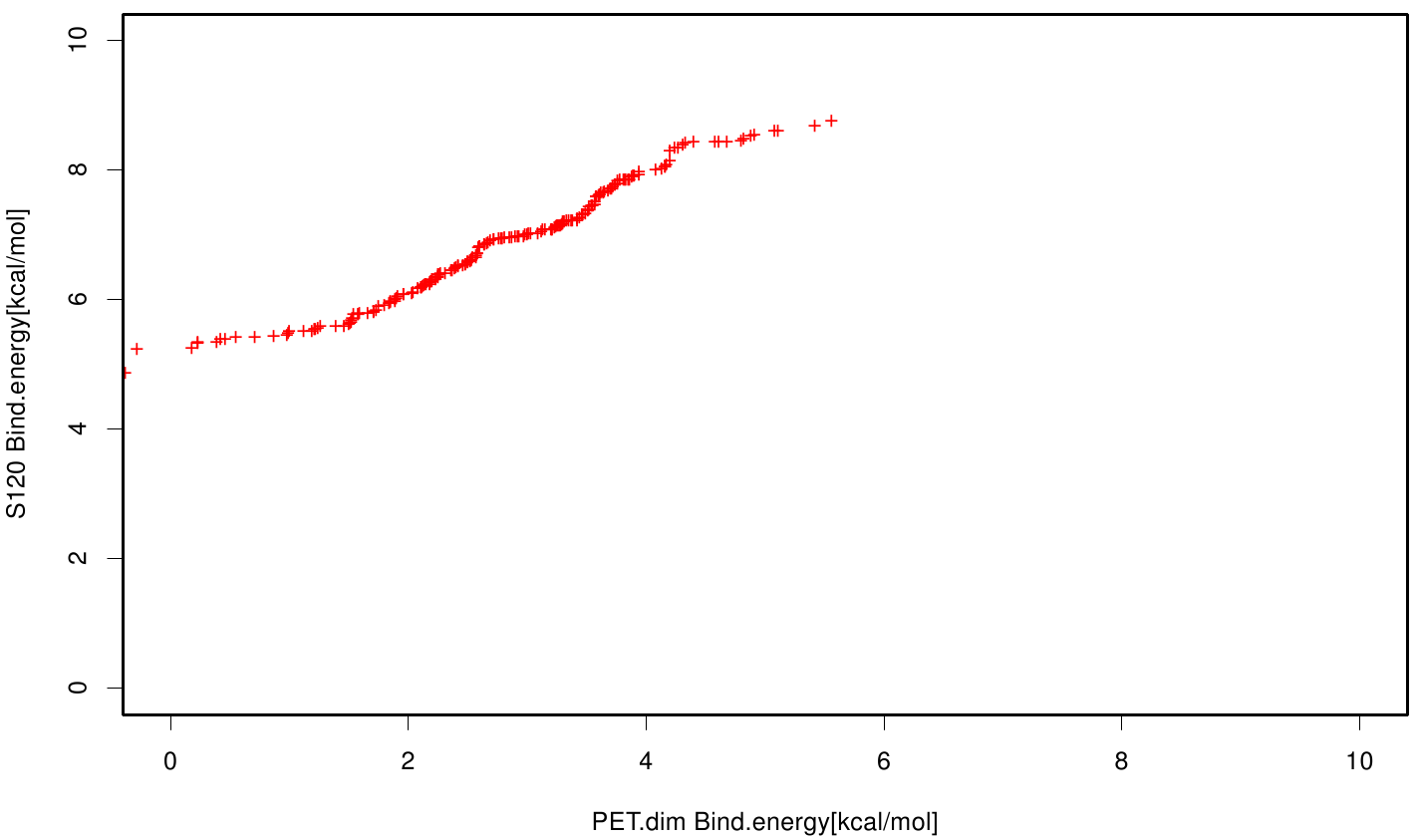
| + | File: S120PET.fsc.scatter.bind.png | Scatter plot of PET vs. S120 | ||
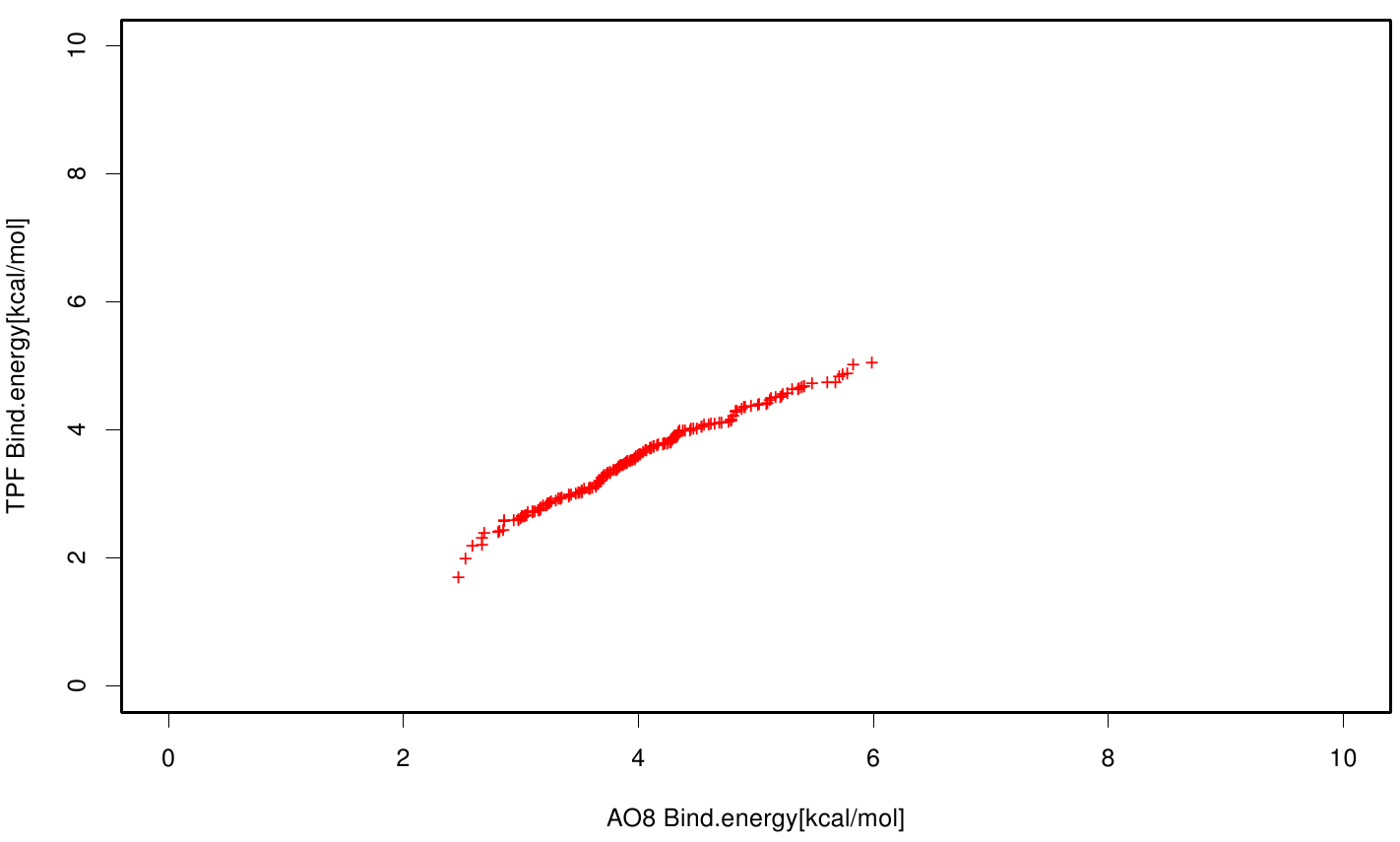
| + | File: AO8TPF.fsc.scatter.bind.png | Scatter plot of AO8 vs. TPF | ||
| + | File: TPFPET.fsc.scatter.bind.png | Scatter plot of TPF vs. PET | ||
| + | File: AO8PET.fsc.scatter.bind.png | Scatter plot of AO8 vs. PET | ||
| + | </gallery> | ||
| + | S120 exhibits the highest binding constant compared to all ligands. Although it results in a strong affinity to the molecule, the target residues are far away from the active site and thus not actually relevant for the degradation process. Interestingly, only TPF possesses overall smaller values than OET. Notably that the binding energies from AO8 are in the same range than TPF. | ||
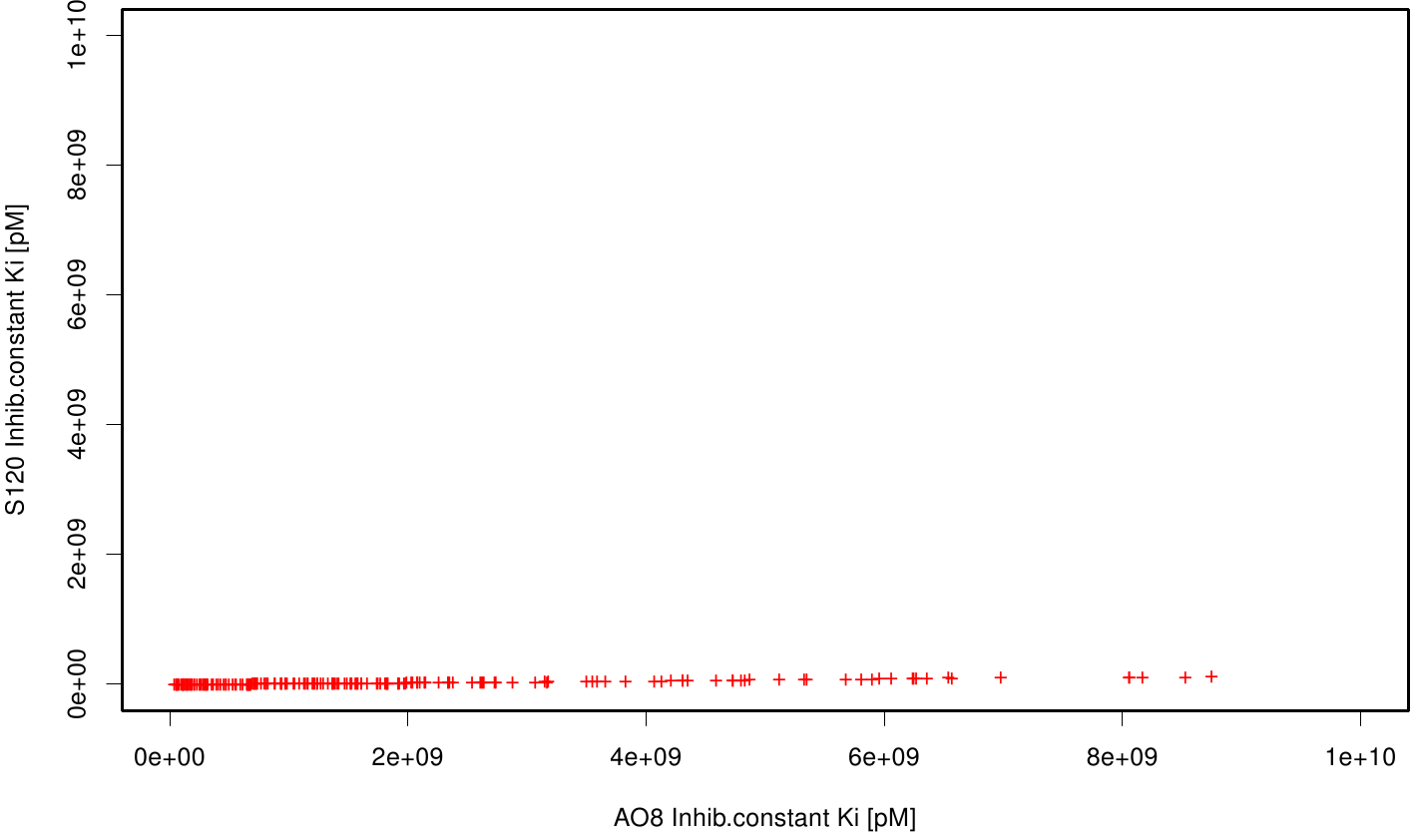
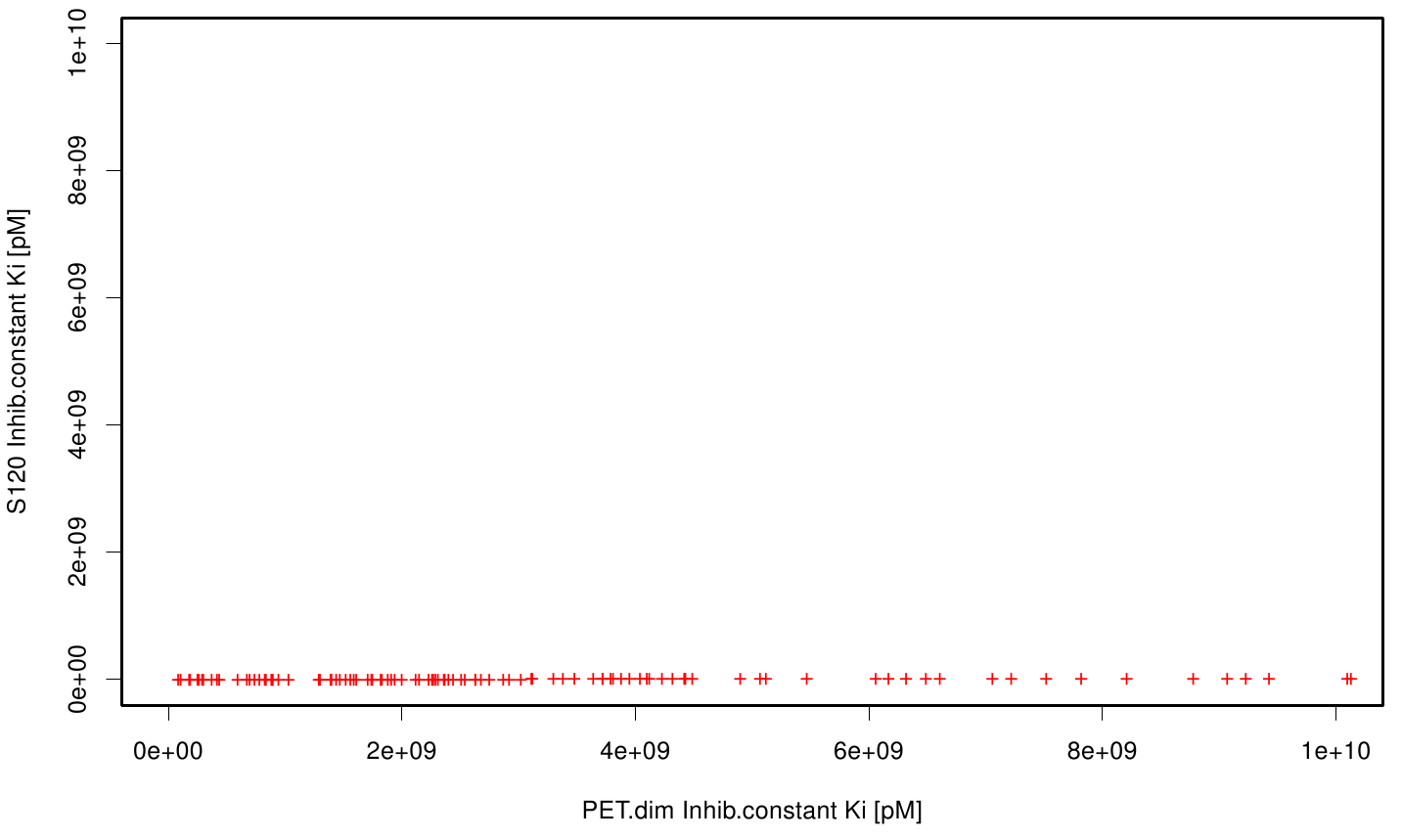
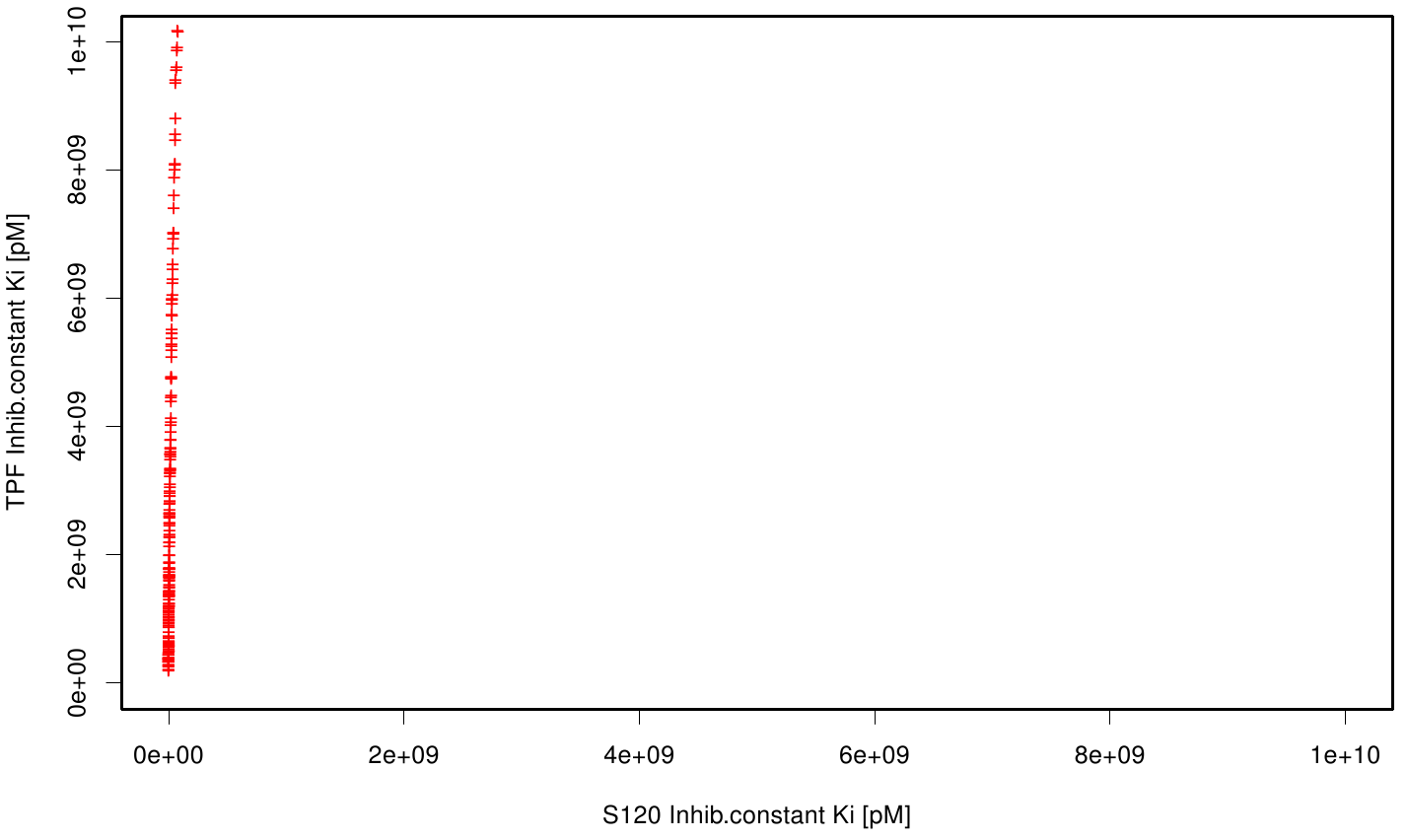
| + | The scatter plots of the inhibition constant are stored in the gallery below: | ||
| + | <gallery widths="400px" heights="250px" perrow="2"> | ||
| + | File: S120AO8.fsc.scatter.inhib.png | Scatter plot of AO8 vs. S120 | ||
| + | File: S120PET.fsc.scatter.inhib.png | Scatter plot of PET vs. S120 | ||
| + | File: S120TPF.fsc.scatter.inhib.png | Scatter plot of TPF vs. S120 | ||
| + | File: TPFPET.fsc.scatter.inhib.png | Scatter plot of TPF vs. PET | ||
| + | File: AO8PET.fsc.scatter.inhib.png | Scatter plot of AO8 vs. PET | ||
| + | File: AO8TPF.fsc.scatter.inhib.png | Scatter plot of AO8 vs. TPF | ||
| + | </gallery> | ||
| + | All plots showed smaller inhibition energies than OET except S120 with the overall smallest values. Overall we can conclude that tested additives not behave like the substrate. We observed that they can't fit in the active site pocket. Thats why we can exclude a direct inhibition of the Fs. cutinase from additives. Notably they could also influent the mechanical behavior of our enzyme while accumulating on the surface. | ||
| + | |||
| + | ====Results for the pNB==== | ||
| + | For the pNB Esterase 13 we study the interaction of OET. The following bar plot illustrate the occurrence of residues in direct contact with different ligand positions and conformations. | ||
| + | |||
| + | [[File: PET.pnb.hist.png |900px|center]] | ||
| + | |||
| + | One major problem of the pNB Esterase seems to be the recognition of the active site. In contrast to the Fs. Cutinase, with its active site at the surface, the pNB exhibits its active site cavity in the center of the protein. That might be the reason for the absence of the following compounds near the active site during every docking approach. For the same reason, we got a nearly uniform distributed occurrence of surface residues complexed with our ligand. | ||
| + | ==References== | ||
| + | [1] Automated Docking Using a Lamarckian Genetic Algorithmand and Empirical Binding Free Energy Function Morris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK and Olson AJ (1998), J.Comput.Chem. 19, 1639-1662 | ||
| + | |||
| + | * [https://2012.igem.org/Team:TU_Darmstadt/Materials/Software Software] | ||
Latest revision as of 20:29, 26 September 2012
| Homology Modeling | | Gaussian Networks | | Molecular Dynamics | | Information Theory | | Docking Simulation |
|---|
Contents |
Docking
Goal
While PET.er is degrading PET into its monomers various chemical compounds like polymer plasticizer are released. Since PET is itself a waste material, it will never be in a pure state. Due to this, it is important to study possible interaction of plasticizer as well as PET with the degradation machinery. Hence an experimental design will give a deeper understanding of the interaction of our enzymes with additives.
Methods
Due to the absence of forcefield terms of plasticizer we choose a global docking approach to study the interaction. For the docking simulations we uses the Autodock plugin of Yasara structure. Autodock is a highly cited docking program developed at the Scripps Research Institute by Dr. Garrett M. Morris et al. [1]
Docking Protocol
For the docking approach we utilized the standard protocol dock_run.mcr and parallelized it up to six processors. Moreover, we choose three additives out of the literature acting as the ligand within the simulations.
- OET-dimeric
- S-120
- AO-8
- TPF
Parameter for the Docking Experiment
- Ligands = S120, AO-8, TPF, dimer OET
- Receptors = PnB Esterase 13 (homology model) and Fs. Cutinase (PDB: [http://www.rcsb.org/pdb/explore/explore.do?structureId=1CEX 1CEX])
- Number of docking runs = 200
- ForceField AMBER03
- Flexible Ligand
Analytics
We extract binding energy from the atomic B-factors and the binding constant from the atomic property derived from all our calculations. We used scatter plots with the binding constant and the inhibition constant in order to compare the plasticizer as well as the PET substrate. We compared the binding energies as well as the inhibition constant from the best ranked docking runs.
Here, OET (dimeric state) became our reference substrate, meaning if an additive exhibits a higher inhibition constant it may effect our protein. Otherwise, if a binding energy exhibits a higher level than OET we can conclude that an additive may accumulate on our protein surface. Moreover, we compare the binding as well as the inhibition energies of the additives with itself to identify the strongest binder.
Since we know the active site of the Fs. Cutinase and the PnB Est. 13, it will be interesting if a plastizicer may dock there. In order to analyze that, we counted the residues which are involved within every docking simulation. Therefore, we measured the distance of every ligand to its corresponding receptor with an appropriate threshold and thus identified the residues.
Results
Interaction with Fs. Cutinase
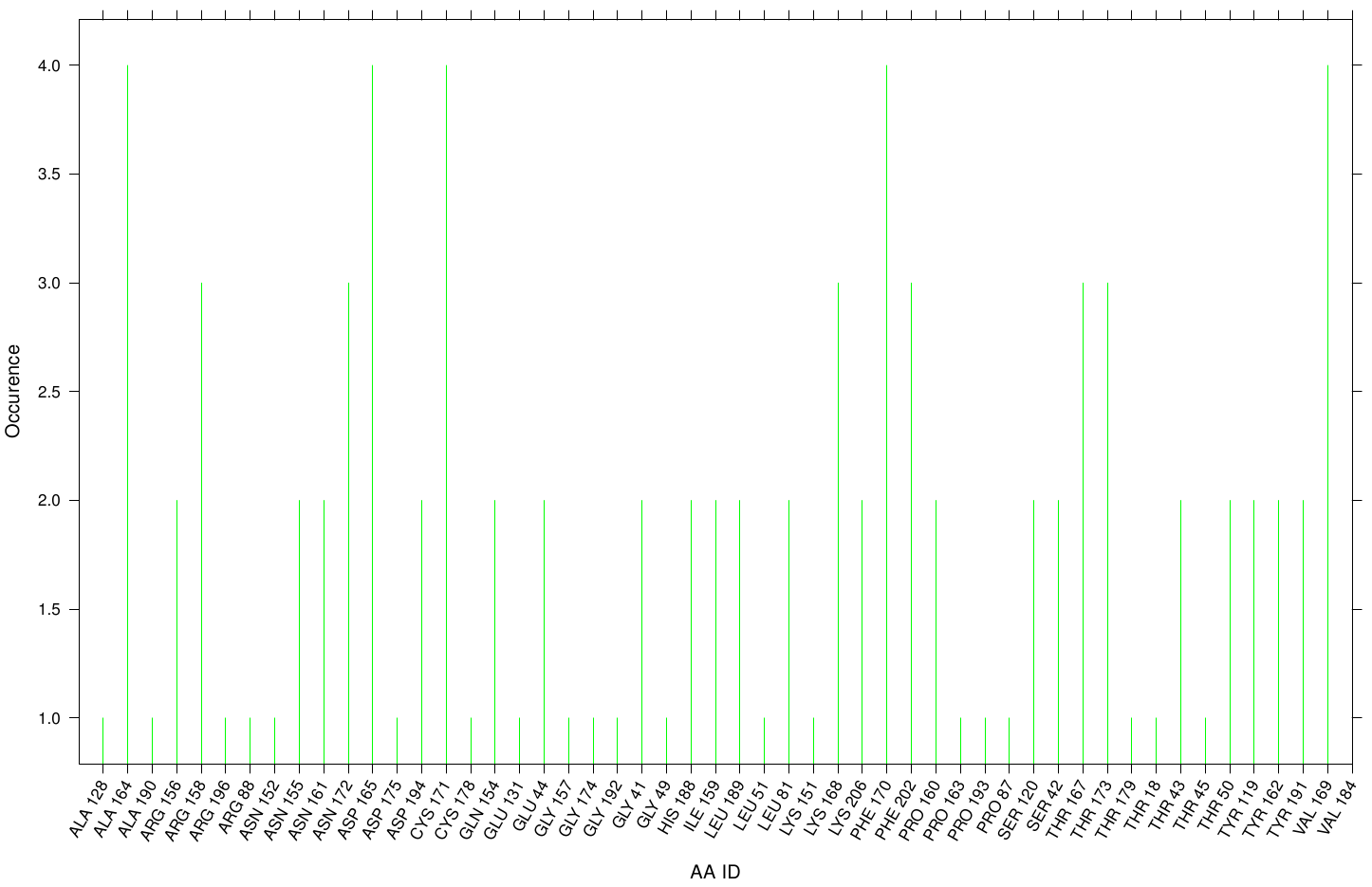
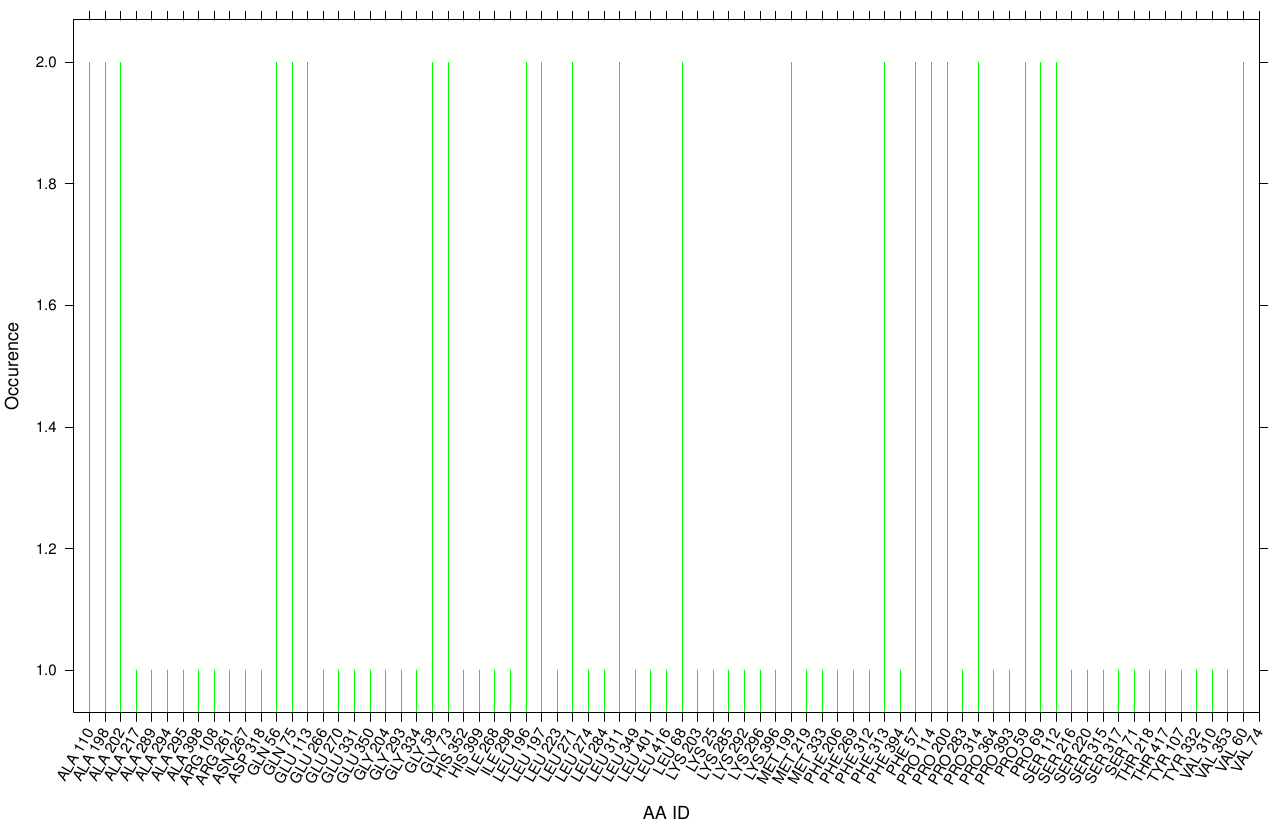
First we study the interaction of OET with the Fs. Cutinase. The following bar plot illustrates the occurrence of residues in direct contact with different ligand positions and conformations.
Within the best runs we got a wide range of involved residues. Since we we set no surface constrains, this is typically for a global docking approach. Here, the residues of the active site, ASP 165 (4 times), SER 120 (2 times) and HIS 188 (2 times), occurs under the best ranked runs. Interestingly, other hydrophobic residues like ALA 164 and VAL 169 are among the high-scoring residues. Since these residues are located near the active site they may help to stabilize the PET Chains on top of the surface during the derogation process. It indicates that OET regularly docked in the active site.
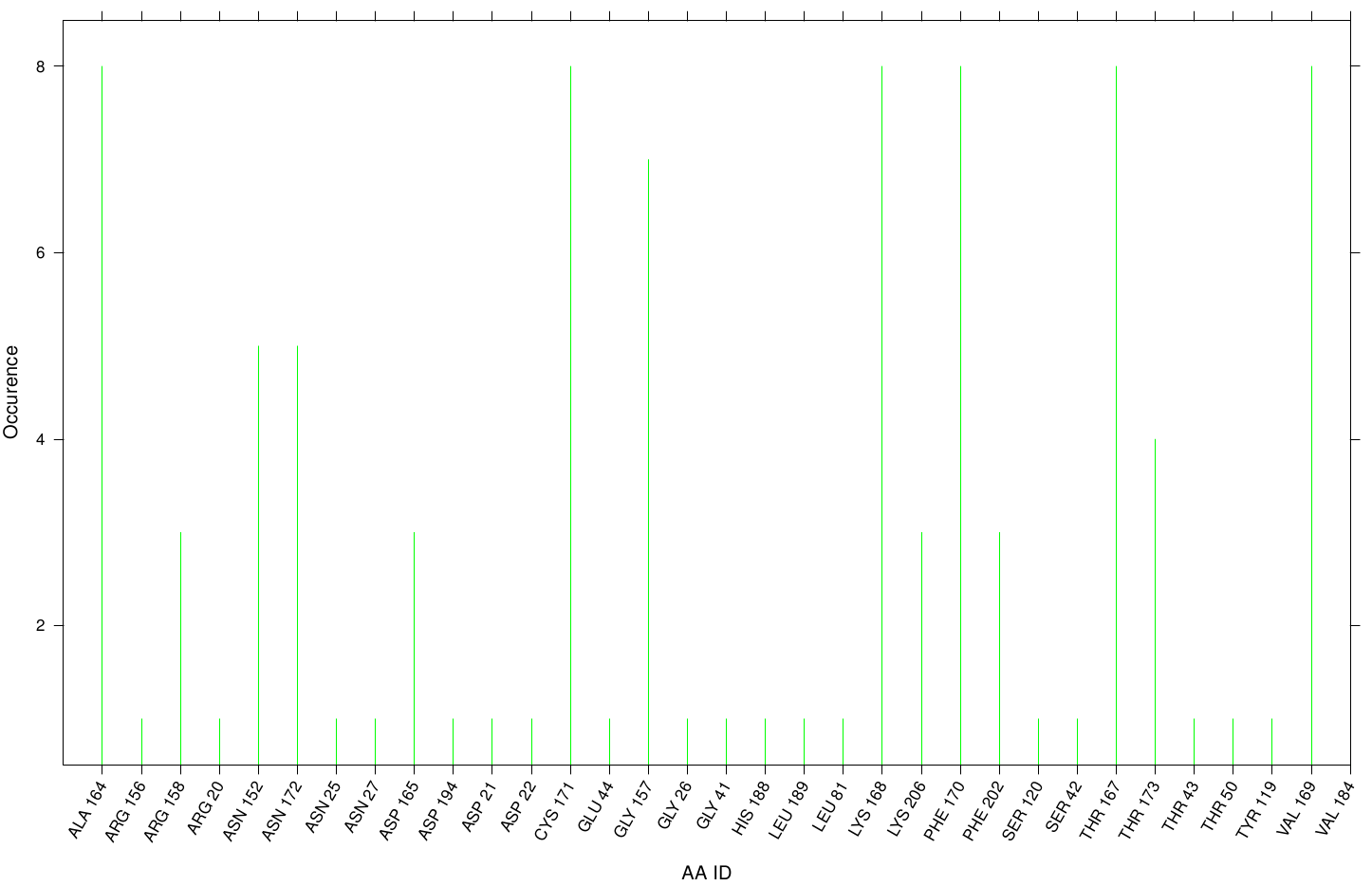
Here we illustrate the docking results of S-120 with Fs. Cutinase:
Only a few residues are highly involved during the docking simulation. We can conclude that S-120 may not effect the active site. For this example we identified possible accumulation sites: CYS 171, LYS 206 and LYS 168 . Although, we didn't identify active site residues, we measured ALA 164 and VAL 169 as one of the high-scoring residues.
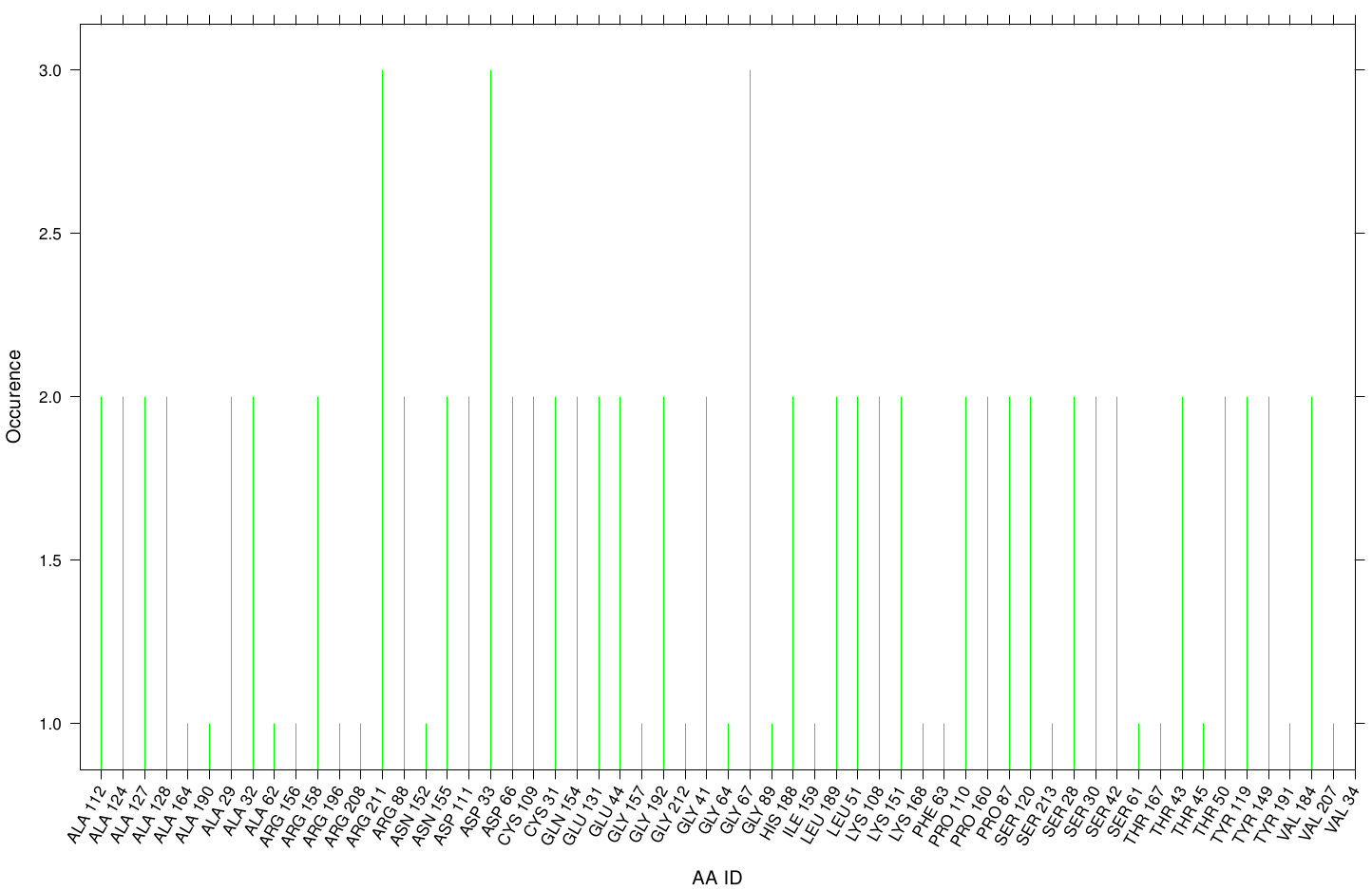
Here we illustrate the the docking results of AO-8 with Fs. Cutinase:
Interestingly, we can observe nearly a uniform distribution of residue occurrence. Due to the hydrophobic nature of AO-8 it is obvious that it distributes ubiquity over the surface.
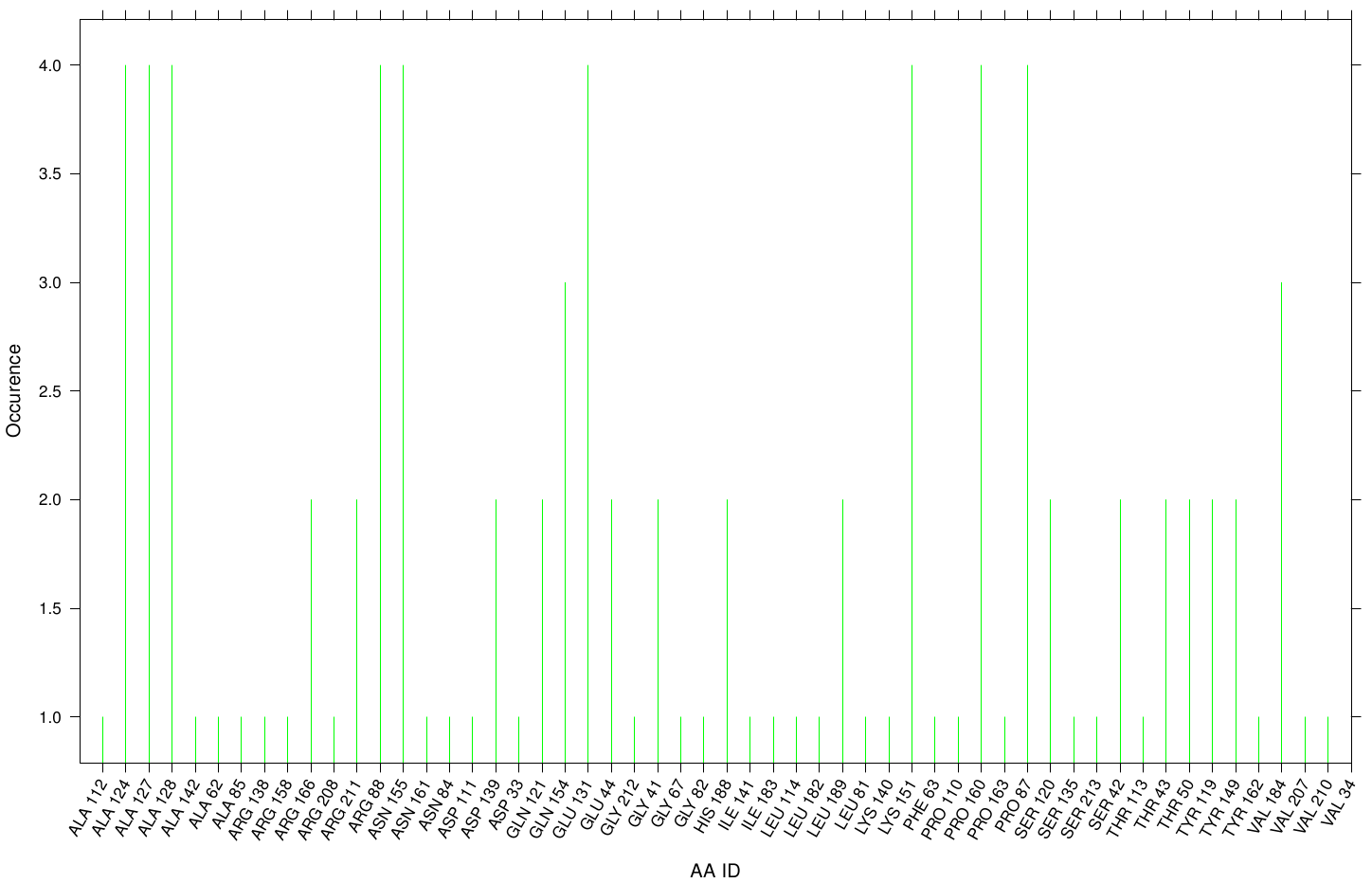
Here we illustrate the the docking results of TPF Fs. Cutinase:
Similar to the results of S-120, the active site residues are not identified under the best ranked docking results (accept for SER 120). The scatter plots of the binding constant are stored in the gallery below:
S120 exhibits the highest binding constant compared to all ligands. Although it results in a strong affinity to the molecule, the target residues are far away from the active site and thus not actually relevant for the degradation process. Interestingly, only TPF possesses overall smaller values than OET. Notably that the binding energies from AO8 are in the same range than TPF. The scatter plots of the inhibition constant are stored in the gallery below:
All plots showed smaller inhibition energies than OET except S120 with the overall smallest values. Overall we can conclude that tested additives not behave like the substrate. We observed that they can't fit in the active site pocket. Thats why we can exclude a direct inhibition of the Fs. cutinase from additives. Notably they could also influent the mechanical behavior of our enzyme while accumulating on the surface.
Results for the pNB
For the pNB Esterase 13 we study the interaction of OET. The following bar plot illustrate the occurrence of residues in direct contact with different ligand positions and conformations.
One major problem of the pNB Esterase seems to be the recognition of the active site. In contrast to the Fs. Cutinase, with its active site at the surface, the pNB exhibits its active site cavity in the center of the protein. That might be the reason for the absence of the following compounds near the active site during every docking approach. For the same reason, we got a nearly uniform distributed occurrence of surface residues complexed with our ligand.
References
[1] Automated Docking Using a Lamarckian Genetic Algorithmand and Empirical Binding Free Energy Function Morris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK and Olson AJ (1998), J.Comput.Chem. 19, 1639-1662
 "
"