Team:UC Chile2/Protocols
From 2012.igem.org
(Difference between revisions)
| (42 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{UC_Chile4}} | {{UC_Chile4}} | ||
| - | <p>Here you will find various protocols we have used during the development of our project. We have separated them into 4 main sections: <br /> [[#General Protocols]] | + | |
| + | [[File: Chile_pipeting.jpg| 300px| right]] | ||
| + | <p>Here you will find various protocols we have used during the development of our project. We have separated them into 4 main sections: <br /><br /> [[#General Protocols]] <br /> [[#Cyanobacteria Protocols]] <br /> [[#Bactomithril Protocols]] <br /> [[#Characterization Protocols]] </p> | ||
<br /> | <br /> | ||
<div id="General Protocols"> | <div id="General Protocols"> | ||
<h1>General Protocols</h1> | <h1>General Protocols</h1> | ||
</div> | </div> | ||
| - | |||
<div id="Growth media"> | <div id="Growth media"> | ||
<h2>Growth media</h2> | <h2>Growth media</h2> | ||
</div> | </div> | ||
| - | |||
<h3>LB media</h3> | <h3>LB media</h3> | ||
| - | |||
| - | |||
<ul> | <ul> | ||
<p>Final concentrations:</p> | <p>Final concentrations:</p> | ||
| Line 44: | Line 42: | ||
<li>Adjust pH to 7.5 with NaOH 1M</li> | <li>Adjust pH to 7.5 with NaOH 1M</li> | ||
</ul> | </ul> | ||
| + | |||
| + | <h3>BG-11</h3> | ||
| + | <br> | ||
| + | |||
| + | For 1L: | ||
| + | <ul> | ||
| + | <li>1.5gr NaNO3</li> | ||
| + | <li>1L distilled H2O </li> | ||
| + | <li>10mL Concentrated salts solution 100X (4°C)</li> | ||
| + | <li>200µL H2PO4 1M (Room T°)</li> | ||
| + | |||
| + | <li>10gr BactoAgar -optional-</li> | ||
| + | |||
| + | </ul> | ||
| + | |||
| + | <h3>BG-110</h3> | ||
| + | <br> | ||
| + | |||
| + | For 1L: | ||
| + | <ul> | ||
| + | <li>1L distilled H2O </li> | ||
| + | <li>10mL Concentrated salts solution 100X (4°C)</li> | ||
| + | <li>200µL H2PO4 1M (Room T°)</li> | ||
| + | <li></li> | ||
| + | <li>10gr BactoAgar -optional-</li> | ||
| + | </ul> | ||
| + | |||
| + | <h3>BG-11C</h3> | ||
| + | <br> | ||
| + | For 1L: | ||
| + | <ul> | ||
| + | <li>1.5gr NaNO3</li> | ||
| + | <li>1L distilled H2O </li> | ||
| + | <li>10mL Concentrated salts solution 100X (4°C)</li> | ||
| + | <li>200µL H2PO4 1M (Room T°)</li> | ||
| + | <li>0.84gr NaHCO3</li> | ||
| + | <li>10gr BactoAgar -opcional-</li> | ||
| + | </ul> | ||
| + | |||
| + | <h3>Concentrated salts solution 100X</h3> | ||
| + | For 1L: | ||
| + | <ul> | ||
| + | <li>7.5gr SO4Mg * 7H2O</li> | ||
| + | <li>3.6gr CaCl2 * 2H2O</li> | ||
| + | <li>0.6gr Citric Acid</li> | ||
| + | <li>0.6gr Ammonium Ferric Citrate</li> | ||
| + | <li>0.093gr EDTA-Na2 </li> | ||
| + | <li>2.0gr CO3 Na2 </li> | ||
| + | <li>0.286gr BO3H3</li> | ||
| + | <li>0.181gr MnCl2 * 4H2O</li> | ||
| + | <li>0.022gr ZnSO4 * 7H2O</li> | ||
| + | <li>0.039gr NaMoO4 * 2H2O</li> | ||
| + | <li>0.0079gr CuSO4 * 5H2O</li> | ||
| + | <li>0.0049gr CoCl2 * 6H2O</li> | ||
| + | <li>1L distilled H2O </li> | ||
| + | </ul> | ||
| + | <br> | ||
| + | |||
| + | Autoclave and store at 4°C. | ||
<div id="Buffers"> | <div id="Buffers"> | ||
| Line 127: | Line 184: | ||
<h3>Gibson Assembly</h3> | <h3>Gibson Assembly</h3> | ||
| - | <p>Gibson Assembly is a DNA assembly method created by Daniel Gibson during the development of the first Synthetic Genome (Synthia) | + | <p>Gibson Assembly is a DNA assembly method created by Daniel Gibson during the development of the first Synthetic Genome (Synthia) [[#2|2]]. Its adaptation to a cloning method allows fast and accurate production of increasingly complex constructions. The strategy behind the method relies on PCR to obtain different parts which share a 40bp homology region, and a 3 enzyme reaction which produces cohesive ends, fills the gaps between the parts and ligates the resulting construct into a scarless assembly of various (>2) parts.</p> |
| - | <h4> | + | <h4>Primer design</h4> |
<p>The easiest way to design primers to obtain amplicons with the required overlaps (40bp final overlaps) is to make an <i>in sillico</i> design of the final construct</p> | <p>The easiest way to design primers to obtain amplicons with the required overlaps (40bp final overlaps) is to make an <i>in sillico</i> design of the final construct</p> | ||
<ul> | <ul> | ||
| Line 169: | Line 226: | ||
</div> | </div> | ||
<ul> | <ul> | ||
| - | <li>Keep 50uL of [ | + | <li>Keep 50uL of [[#Competent Cells| chemically competent cells]] in ice (for no more than 10 minutes until transformation)</li> |
<li>Mix 0.01 to 1 total ng of plasmid with competent cells (depending on size of plasmid and competence of your cells)</li> | <li>Mix 0.01 to 1 total ng of plasmid with competent cells (depending on size of plasmid and competence of your cells)</li> | ||
<p>Flick very softly to mix (competent cells are very fragile)</p> | <p>Flick very softly to mix (competent cells are very fragile)</p> | ||
| Line 178: | Line 235: | ||
<li>Plate 50uL of transformed cells to a LB agar petri dish with corresponding antibiotic (30ug/mL Kanamycin or 30ug/mL Chloramphenicol or 100ug/mL Ampicillin)</li> | <li>Plate 50uL of transformed cells to a LB agar petri dish with corresponding antibiotic (30ug/mL Kanamycin or 30ug/mL Chloramphenicol or 100ug/mL Ampicillin)</li> | ||
</ul> | </ul> | ||
| + | |||
| + | </div> | ||
| + | <div id="Competent Cells"> | ||
| + | <h2>Chemically Competent cells</h2> | ||
| + | <p>To prepare our competent cells we used a protocol we found in openwetware.org. Starting from commercial TOP10 cells as the starting inoculus for the whole protocol, we get competence levels close to 5*10⁸ cfu/ug pUC19 DNA</p> | ||
| + | <br /> | ||
| + | [http://openwetware.org/wiki/TOP10_chemically_competent_cells Openwetware protocol here] | ||
<div id="Cyanobacteria Protocols"> | <div id="Cyanobacteria Protocols"> | ||
<h1>Cyanobacteria Protocols</h1> | <h1>Cyanobacteria Protocols</h1> | ||
</div> | </div> | ||
| - | |||
<div id="Transformation of Synechocystis"> | <div id="Transformation of Synechocystis"> | ||
| + | |||
<h2>Transformation of Synechocystis PCC. 6803</h2> | <h2>Transformation of Synechocystis PCC. 6803</h2> | ||
</div> | </div> | ||
| - | < | + | By iGEM - UC_Chile Team 2012 (Based on [[#1 | 1]]) |
| + | <h3>Materials</h3> | ||
| + | <ul> | ||
| + | <li>Liquid BG-11 Media</li> | ||
| + | <li>Solid BG-11 Media</li> | ||
| + | <li>Sterile 250 mL Erlenmeyer flask</li> | ||
| + | <li>Sterile 50 mL Falcon tube (2)</li> | ||
| + | <li>Sterile 1.5ml Eppendorf tubes</li> | ||
| + | <li>Petri Dishes</li> | ||
| + | <li>Millipore Membrane Filters, 0.45um HA (HAWP09000) </li> | ||
| + | <li>Pasteur pipette</li> | ||
| + | <li>Antibiotics (according to your selection resistance)</li> | ||
| + | <li>DNA for transformation</li> | ||
| + | </ul> | ||
| + | <h3>To be prepared previously</h3> | ||
| + | <br> | ||
| + | Concentrate the DNA to be transformed into Synechocystis to 1µg/µL in 10µL for each transformation (A total of 10 ug of the transformation plasmid). | ||
| + | <br> | ||
| + | Inoculate Synechocystis PCC 6803 into a sterile 250mL Erlenmeyer flask with 150mL* of liquid BG-11 and grow culture until you reach an OD730nm of 0.8 – 1.0 (That should take between 6 to 10 days dependending on the amount of initial inoculum). | ||
| + | * For each 50 mL of Synechocystis PCC 6803 you will have 5 individual transformations. | ||
| + | <br> | ||
| + | Once you have reached the OD730nm and have enough DNA, proceed to transform. | ||
| + | <br> | ||
| + | <h3>Transformation</h3> | ||
| + | |||
| + | All subsequent actions are to be realized under sterile conditions. | ||
| + | <ul> | ||
| + | <li>1.-Transfer Synechocystis cells into 50 mL Falcon tubes and centrifuge 10 minutes at 2760g (Room temperature)</li> | ||
| + | |||
| + | <li>2.- Wash cells: </li> | ||
| + | Discard supernatant and resuspend pellet in the following manner: | ||
| + | <ul>- Add 1mL of liquid BG-11 and softly pipet cells until resuspended.</ul> | ||
| + | <ul>- Add 4mL of liquid BG-11 and pipet cells until completely mixed.</ul> | ||
| + | <ul>- Add 5ml of BG-11 and pipet cells.</ul> | ||
| + | * It is important that cells are throroughly resuspended so that recieve fresh media and do not form aggregates. | ||
| + | <li>3.- Centrifuge 10 minutos at 2760g (Room temperature).</li> | ||
| + | <li>4.- Discard supernatant and extract any remaining media by pipeting.</li> | ||
| + | <li>5.- Resuspend pellet with 1mL of BG-11 media. Once resuspended, add 0.5 mL more of BG-11. </li> | ||
| + | <li>6.- Distribuite 300 uL of resuspended cells into sterile 1,5 mL Eppendorf tube.</li> | ||
| + | <li>* Make sure to pipet into the bottom of the tube so that none of the cells remain in the walls of the tube.</li> | ||
| + | <li>7.- Add 10 μL of plasmid DNA (1μg/μL) to the tube containing the 300 uL of the Synechocystis cells.</li> | ||
| + | <li>Softly mix contents with the point of the pipet. Repeat the procedure to all but the transformation control tube (without DNA). </li> | ||
| + | <li>8.- Leave the tubes in a light chamber (50 uE/s²/m²) at 30°C for 5 hours. Flick the tubes at 2.5 hours. </li> | ||
| + | </ul> | ||
| + | <h3>Plate preparation</h3> | ||
| + | <br> | ||
| + | Prepare 50 mL BG-11 plates with no antibiotic for the transformation day. | ||
| + | Put Millipore membrane filters on BG-11 plates. | ||
| + | <ul> | ||
| + | |||
| + | |||
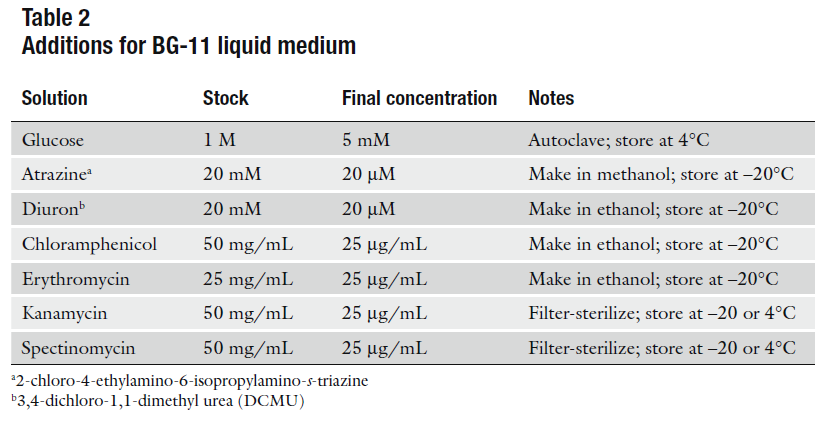
| + | [[File: Ant_table.jpg| 500px| right]] | ||
| + | |||
| + | <li>9.- After the 5 hours of recuperation, take the celular suspension and spread over the Millipore membrane. The suspension is to be spreaded carefully over the whole membrane using a sterile bended Pasteur pipete.</li> | ||
| + | <li>10.- Let the membranes dry before transfering the plates to the growth chamber.</li> | ||
| + | <li>11.- Transfer plates to the growth chamber putting the membrane facing up and leave at 30°C for 12 hours in light.</li> | ||
| + | |||
| + | |||
| + | <li>12.- Transfer membranes to BG-11 plates that contain half of the concentration of the antibiotic (12.5 ug/mL of antibiotic) and leave growing for 3 days at 30°C in light. </li> | ||
| + | <li>13.- After the 3 days have passed, transfer the membrane to BG-11 plates with full concentration of antibiotic (25ug/mL).</li> | ||
| + | </ul> | ||
| + | <br><br> | ||
| + | <i>Colonies should appear between 1 to 2 weeks after the transformation. First you will see most of the cellular suspension dissapear and slowly small colonies should appear in the plate.</i> | ||
| + | |||
| + | |||
<div id="Reinoculation"> | <div id="Reinoculation"> | ||
<h2>Reinoculation</h2> | <h2>Reinoculation</h2> | ||
| + | Protocol right here! | ||
</div> | </div> | ||
| + | |||
<div id="Cyanobacterial DNA extraction"> | <div id="Cyanobacterial DNA extraction"> | ||
<h2>Cyanobacterial DNA extraction</h2> | <h2>Cyanobacterial DNA extraction</h2> | ||
</div> | </div> | ||
<p>Protocol goes here!</p> | <p>Protocol goes here!</p> | ||
| + | |||
<div id="Bactomithril Protocol"> | <div id="Bactomithril Protocol"> | ||
<h1>Bactomithril Protocols</h1> | <h1>Bactomithril Protocols</h1> | ||
</div> | </div> | ||
<br /> | <br /> | ||
| - | |||
<p>You can find a complete protocol to produce spider-silk fibers in the following paper: | <p>You can find a complete protocol to produce spider-silk fibers in the following paper: | ||
Teulé, F., Cooper, A. R., Furin, W. a, Bittencourt, D., Rech, E. L., Brooks, A., & Lewis, R. V. (2009). A protocol for the production of recombinant spider silk-like proteins for artificial fiber spinning. Nature protocols, 4(3), 341-55. doi:10.1038/nprot.2008.250 | Teulé, F., Cooper, A. R., Furin, W. a, Bittencourt, D., Rech, E. L., Brooks, A., & Lewis, R. V. (2009). A protocol for the production of recombinant spider silk-like proteins for artificial fiber spinning. Nature protocols, 4(3), 341-55. doi:10.1038/nprot.2008.250 | ||
| - | Other easier protocols are in the following adresses:<br> | + | <br><br>Other easier protocols are in the following adresses:<br> |
http://www.pnas.org/content/107/32/14059.long<br> | http://www.pnas.org/content/107/32/14059.long<br> | ||
| Line 212: | Line 342: | ||
<h1>Characterization Protocols</h1> | <h1>Characterization Protocols</h1> | ||
</div> | </div> | ||
| + | |||
| + | <div id="SyneGrowth"> | ||
| + | <h2>Methods for the characterization of Synechocystis PCC 6803 growth curve</h2> | ||
| + | This are the methods we used for setting up the growth curve experiment of our Synechocystis PCC 6803 which is further described [[Team:UC_Chile2/Characterization | here ]]. | ||
| + | |||
| + | <h3>Materials</h3> | ||
| + | <ul> | ||
| + | <li>4 sterile (autoclaved) 250 mL Erlenmeyer flasks</li> | ||
| + | <li>1 liter of BG-11</li> | ||
| + | <li>250 mL of BG-110</li> | ||
| + | <li>Axenic culture of Synechocystis PCC 6803</li> | ||
| + | <li>Sterile filtered 1 mL tips</li> | ||
| + | <li>Spectrophotometer</li> | ||
| + | <li>5 1 mL Cuvettes</li> | ||
| + | </ul> | ||
| + | <br> | ||
| + | |||
| + | All procedures should be realized under sterile conditions, preferably in a laminar flow hood. | ||
| + | |||
| + | <ul> | ||
| + | <li>Measure 150 mL of BG-11 with a sterile 50 mL Falcon tube and put into each of the 3 Erlenmeyer flasks.</li> | ||
| + | <li>Measure 150 mL of BG-110 with a sterile 50 mL Falcon tube and put into the remaining Erlenmeyer flask.</li> | ||
| + | <li>Inoculate 1 mL of axenic Synechocystis PCC 6803 with and OD730 of X to each flask (including the one with BG-110). </li> | ||
| + | <li>Place flasks in rotating incubator and try to avoid leaving them out of the incubator for long when taking measurements</li> | ||
| + | <li>Measure OD600 and OD730 twice a day (preferably with at least 8 hours of difference between measurements)</li> | ||
| + | <li>Repeat measurements for 14 days</li> | ||
| + | </ul> | ||
| + | </div> | ||
| + | |||
<div id="LuxBrick Characterization"> | <div id="LuxBrick Characterization"> | ||
| Line 218: | Line 377: | ||
<ul> | <ul> | ||
<li> Pick a single colony from a transformed plate of K325909. Grow in LB media with chloramphenicol 30ug/mL O/N at 30°C.</li> | <li> Pick a single colony from a transformed plate of K325909. Grow in LB media with chloramphenicol 30ug/mL O/N at 30°C.</li> | ||
| - | <li>Measure OD at 600nm, then dilute to OD600=0.6 following direct C*V ratio to obtain bacterial stock for experiments.</li> | + | <li>Measure OD at 600nm, then dilute to OD600 = 0.6 with growth media following direct C*V ratio to obtain bacterial stock for experiments.</li> |
<li>Induce with 3mM (L)-Arabinose (final concentration)</li> | <li>Induce with 3mM (L)-Arabinose (final concentration)</li> | ||
<li> Alicuot 600uL of bacteria and add LB media with chloramphenicol 30ug/mL + additives and inducers up to 1.5mL. This will yield a 0.24 OD600nm.</li> | <li> Alicuot 600uL of bacteria and add LB media with chloramphenicol 30ug/mL + additives and inducers up to 1.5mL. This will yield a 0.24 OD600nm.</li> | ||
<li> Grow at 30°C for 1.5 hours (or at chosen T° condition for characterization).</li> | <li> Grow at 30°C for 1.5 hours (or at chosen T° condition for characterization).</li> | ||
| - | <li> Alicuot 100uL into 4 wells | + | <li> Alicuot 100uL into 4 wells of a 96 well falcon plaque.</li> |
<li> Measure luminescence in a ThermoScientific Luminoskan Ascent Microplate Luminometer.</li> | <li> Measure luminescence in a ThermoScientific Luminoskan Ascent Microplate Luminometer.</li> | ||
</ul> | </ul> | ||
| + | |||
| + | <div id="Gibson Assembly for small parts Characterization"> | ||
| + | <h2>Methods for Gibson Assembly for small parts characterization</h2> | ||
| + | </div> | ||
| + | The experiment for the characterization of T5 exonuclease concentration in Gibson Assembly for small parts was set as follows: | ||
| + | <ul> | ||
| + | <li>After the design of the segment to be assembled as described in the [[Team:UC_Chile2/Results/Gibson#Gibson_Assembly_for_small_parts | Characterization section]], amplification of the parts was done using standard Phusion Polymerase PCR conditions but with a total amount of template of 0.05ng</li> | ||
| + | <li>Bands where cut and purification of DNA from agarose gel was realized using a Promega's Wizard SV Gel and PCR Clean-Up System, final elution volume of 20 ul</li> | ||
| + | <li>Purified DNA was quantified using a Nanodrop 2000</li> | ||
| + | <li>According to concentration of DNA, the amount of pmoles/uL of each part was calculated using the following formulae:</li> | ||
| + | pmoles of DNA = weigth in ng * 1000 (conversion factor from nano to pico) / (650 Daltons * base pair length of part) | ||
| + | <li>Volumes of each part were calculated to obtain a final amount of 0.0125 pmoles of the small part and 0.0025 pmoles of the backbone in a final volume of 3 uL (add nuclease-free water to reach 3 uL) for each reaction</li> | ||
| + | <li>For the preparation of Gibson Assembly Master Mixes, different volumes of a 10X T5 exonuclease 1.33X Gibson Assembly Master Mix were mixed with T5 exonuclease-free 1.33X Gibson Assembly Master Mix until appropriate concentrations were reached. If mixing volumes of any of the Master Mixes was lower than 1 uL, then one of the diluted T5 exonuclease 1.33X Master Mixes was used for the mix.</li> | ||
| + | <li>Gibson Assembly was done as described [[#Assembly_reaction| here]], and transformation proceeded as described [[#E.coli_Transformation| here]]</li> | ||
| + | <li>Each of the concentration points of the experiment was done in triplicate</li> | ||
| + | <li>Red colonies and white colonies were counted 20 hours post-transformation</li> | ||
| + | </ul> | ||
| + | |||
| + | |||
| + | <h1>References</h1> | ||
| + | <div id="1"> | ||
| + | [1]Julian J. Eaton-Rye , “Construction of Gene Interruptions and Gene Deletions in the Cyanobacterium Synechocystis sp. Strain PCC 6803 ”, Photosynthesis Research Protocols, Methods in Molecular Biology, vol. 684, DOI 10.1007/978-1-60761-925-3_22 | ||
| + | </div> | ||
| + | |||
| + | <div id="2"> | ||
| + | [2]Gibson DG, Young L, Chuang RY, Venter JC, Hutchison CA 3rd, Smith HO. (2009). "Enzymatic assembly of DNA molecules up to several hundred kilobases". Nature Methods 6 (5): 343–345. doi:10.1038/nmeth.1318 | ||
| + | </div> | ||
| + | {{UC_Chilefooter}} | ||
Latest revision as of 11:06, 25 September 2012
 "
"