Team:UC Chile2/Characterization
From 2012.igem.org
| Line 10: | Line 10: | ||
<br> | <br> | ||
| - | Methods for the growth curve characterization of Synechocystis PCC 6803 over [[Team:UC_Chile2/Protocols| here]]. | + | Methods for the growth curve characterization of Synechocystis PCC 6803 over [[Team:UC_Chile2/Protocols#SyneGrowth| here]]. |
Revision as of 08:10, 25 September 2012

Contents |
Synechocystis PCC 6803 growth curve
When we started working on the project we realized that although we could manage to acquire an inoculum of Synechocystis PCC 6803 from a laboratory in our faculty, no protocols were standardized in that lab for using Synechocystis for research. That meant obtaining protocols and trying them out. Also, it meant we had to set the conditions for growing them to do all the experiments in due time. Actually, we started out using the conditions set for the growth of many other cyanobacteria, but after a couple of failed transformations we decided to build our own Synechocysistis incubator (you can find a DIY guide over here).
Here we show our growth curve for Synechocystis PCC 6803 using the conditions set in our shaking incubator (40uE/s²/m²), 90 RPM, 30°C.
Methods for the growth curve characterization of Synechocystis PCC 6803 over here.
LuxBrick Characterization - (BBa_K325909)
As a part of our project, we set out to investigate the conditions which affect the output of the LuxBrick. As we are using various elements from this Biobrick, we believe that is cornersome to know how we may increase its output by modifying the conditions that affect its behaviour.
We set out to understand diverse factors that could be influencing the total bioluminescence yield of the brick. Some of them have direct relation with some of the aspects of our project. These factors are:
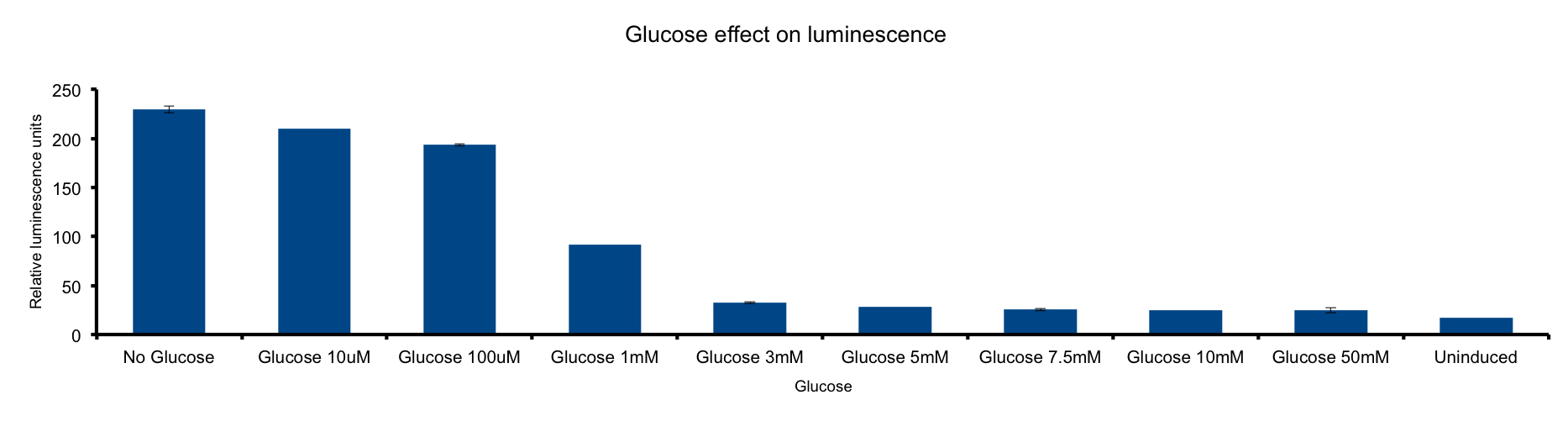
Glucose concentration
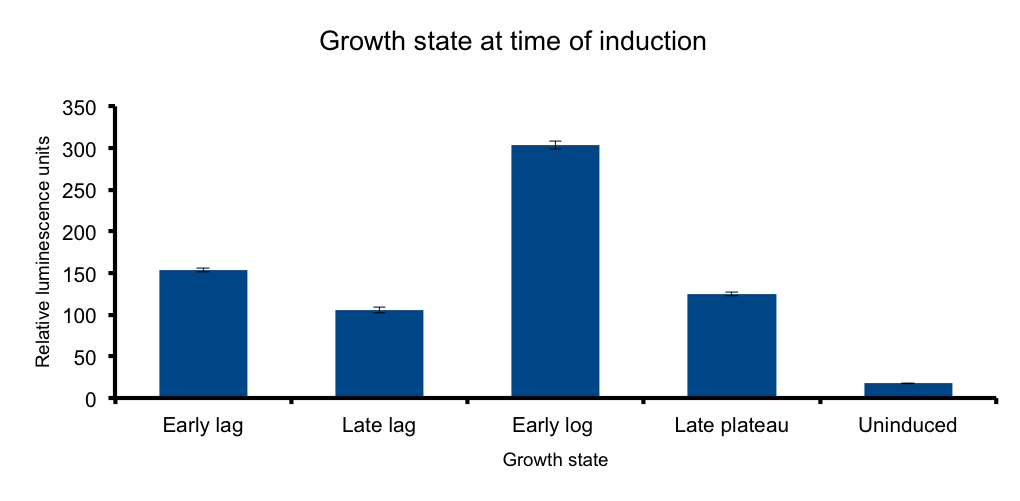
Growth state at time of induction
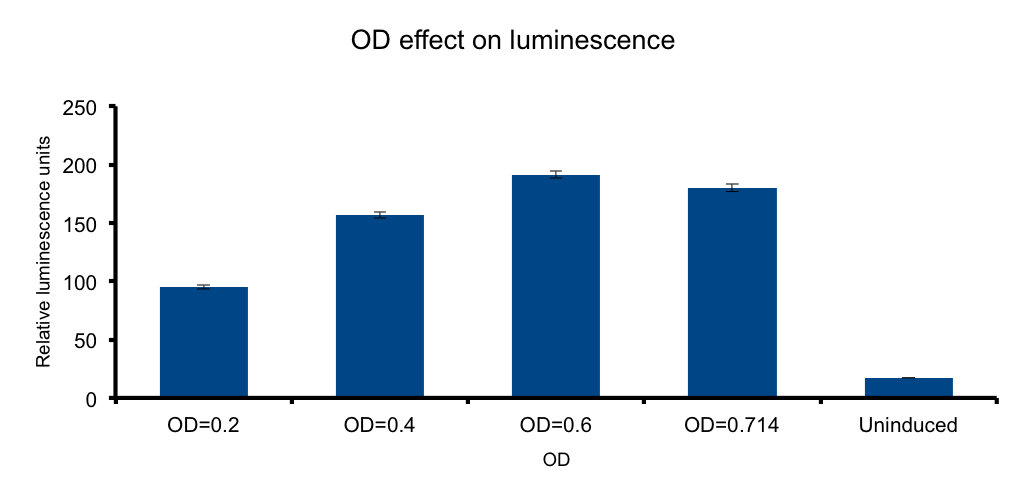
Cell mass
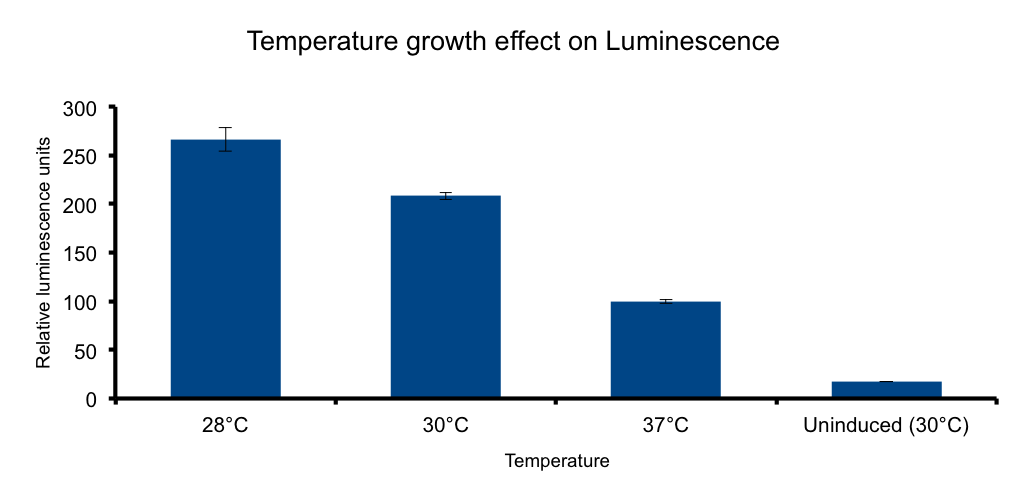
Temperature
Here you'll find detailed the characterization methods we used in this experiment.
Glucose concentration in media
HERE THERE SHOULD BE SOME TEXT SUPPORTING THE EXPERIMENT HOPEFULLY WITH CITATION.
FOR EXAMPLE:
Glucose is most important energy source of bacteria. As glucose is catabolized into pyrovate ...
Growth state at-time-of-induction
HERE THERE SHOULD BE SOME TEXT SUPPORTING THE EXPERIMENT HOPEFULLY WITH CITATION. It has been know for some time that general bacterial metabolism is principally determined its growth state. Growth states involve the expression of different sigma factors according to the metabolic status of the bacteria. The transcription of certain sequences directly depend on this, as some promoters are only read by the RNA polymerase when sigma70 is present in the cell. (TO BE CONTINUED)
Growth temperature
HERE THERE SHOULD BE SOME TEXT SUPPORTING THE EXPERIMENT HOPEFULLY WITH CITATION. Protein folding, vibrio fisherii free living organism... blablabla
sfGFP vs mRFP1
Improvement of an existing BioBrick
Our chosen target was Biobrick J04450 which is a RFP Coding Device (link: http://partsregistry.org/Part:BBa_J04450). Its usage as a cloning tool has increased in time due to the easy detection of transformed colonies to the naked eye. Nonetheless, the advancement of automatic measurement systems (and yada yada) opens up a whole new range of applications in which markers are detected by highly sensitive machines and not by human observers. In these cases mRFP does not provide a significant advantage over other marker systems. In fact, it could be detrimental for the efficiency of the experiment (?)
Therefore, we propose to exchange part mRFP for sfGFP. sfGFP would prove to be a better part for the following reasons:
sfGFP is of shorter length than mRFP1 (proposed implication: Lower metabolic cost. )
sfGFP has higher quantum yield/faster maturation rate or something of the sort!
To demonstrate the improvement of the biobrick, we designed and run the following experiments:
Experiment : Culture growth
Three flasks were inoculated in equal concentrations with: psB4K5-J04450, psB4K5 – ours and wildtype TOP 10 cells respectively. Inoculum concentrations were checked by culture OD measurement at 600 nm to normalize the amount of bacteria that would be initially present.
Then, OD measurements at 600 nm, 584 nm and 492 nm were reported every 30 minutes for each of the flasks. 584 nm and 492 nm are the peak absortion wavelengths of mRFP1 and sfGFP respectively.
Insert:Explanation of why this should prove improvement
Results 1. Growth rate Experimental data was adjusted in order to…. Waiting for results ☺
Results 2.
Experimental data was adjusted in order to….
Waiting for results ☺
References
 "
"