Team:NTU-Taida/Modeling/Stochastic-Analysis
From 2012.igem.org
Contents |
Stochastic Analysis
Overview
Besides the parameter space search, we performed another type of analysis to verify the robustness and the mono-stability of our system. Because gene expression is an intrinsically stochastic process, we performed stochastic simulations to see how our system reacts to noise and how it responds to perturbations. We were especially interested in the filter function, whether it is always present and whether the amount of GLP-1 produced in the cells for different concentrations of fatty acid has large fluctuations.
Method
We performed the stochastic analysis using the Gillespie algorithm by the numerical Matlab solver. In the Gillespie algorithm, propensity theory is used to describe the behavior of the system. Each reaction occurring in the cell has a certain propensity. We separated the degradation terms from the activation/repression terms in our single cell ODEs, and converted the species values, production rates and repression coefficients from concentrations (in μM) to number of molecules to derive the propensity for each reaction involved in our system. A reaction with a relatively high propensity will have more chance to occur than another one. In order to determine the next event in a stochastic simulation, the propensity of all possible changes to the state of the model are computed, and then ordered in an array. Next, the cumulative sum of the array is taken, and the final cell contains the number R, where R is the total event propensity. This cumulative array is now a discrete cumulative distribution, and can be used to choose the next event by picking a random number z~U(0,R) and choosing the first event, such that z is less than the propensity associated with that event. In this way, the reaction to happen at the time point is determined, the concentrations of each species are changed according to the reaction, and the same process is repeated until the ending time of the simulation is reached. Following these steps we can get a curve of the evolution of a molecule in the cell that takes into account the randomness of reactions occurring in the cell.
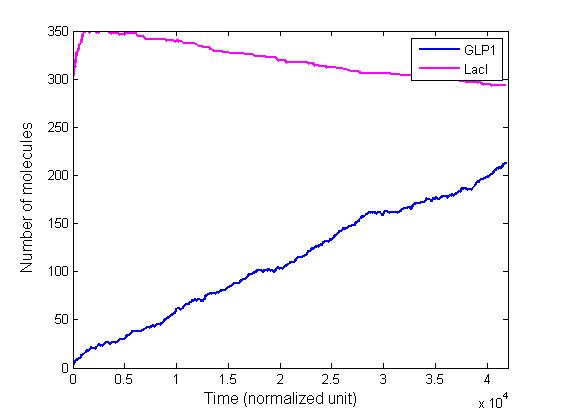
Since the calculation is partially based on a random selection from the reactions, each run of the algorithm will produce a slightly different curve. However, the curves will converge to approximately the same level, which is predicted by the deterministic equations.
Analysis of our system
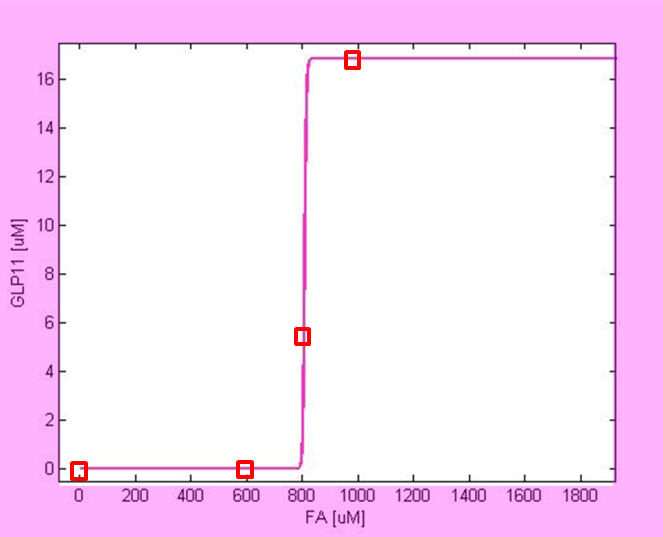
For analyzing the robustness of the system, we chose 4 different fatty acid concentrations (see Figure 2), and for each of them we performed stochastic simulations. We selected the following Acetaldehyde concentrations:
- 0 μM : At this concentration the GLP-1 output should be very low
- 600 μM: This concentration is below the threshold, and therefore the GLP-1 output should also be very low.
- 800 μM: This is the concentration around the threshold, so the output response level should be between the lowest and the highest level.
- 1000 μM: This concentration exceeds the threshold, and therefore should result in maximum level of GLP-1 value.
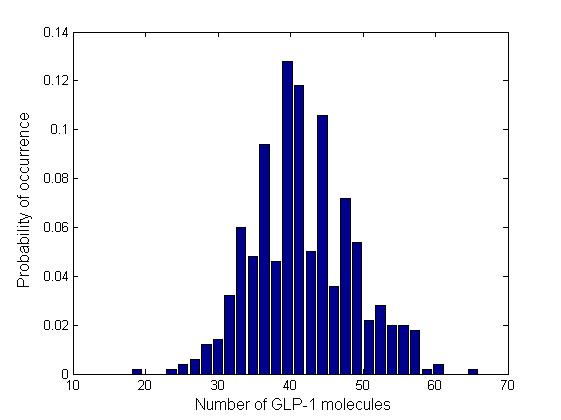
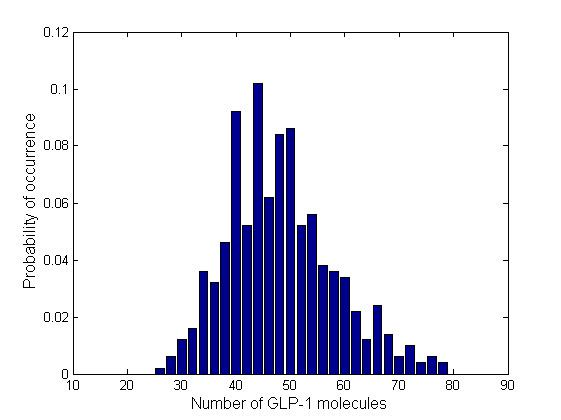
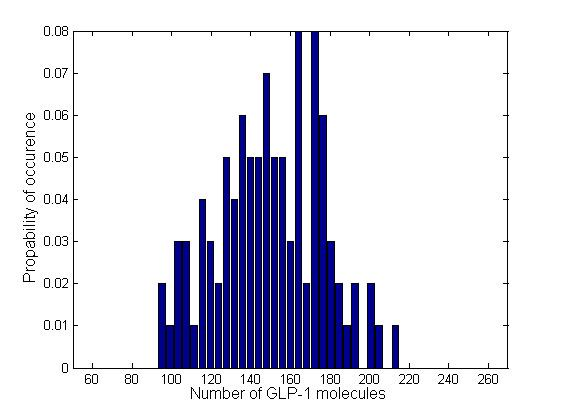
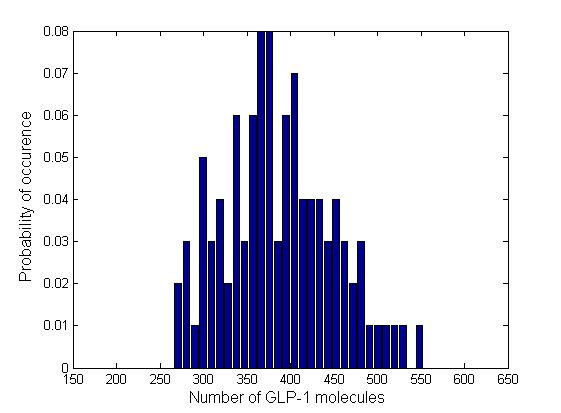
In order to see how the numbers of molecules are distributed and whether there are multiple steady states, we plotted the normalized histograms for the 4 data sets. We used 30 bins for plotting the histograms.
Due to time limitation, the number of runs we had performed is not large enough, therefore the distribution is not very smooth. However, generally the information brought by a few instances of the Gillespie simulation may also give us a glimpse into the general behavior of the system. All of the 4 distributions have a Gaussian-like shape, which means that the number of GLP-1 molecules centralizes around a mean value, confirming the system is mono-stable. We fit the histogram into Gaussian curves and obtained the mean and standard deviation of each distribution. The relative standard deviations are in the range of 23% ~ 28%, which are relatively narrow, which means there are no large fluctuations.
 "
"