Team:TU Munich/Modeling/Priors
From 2012.igem.org
Revision as of 15:38, 6 September 2012 by FabianFroehlich (Talk | contribs)



Contents |
Prior Data
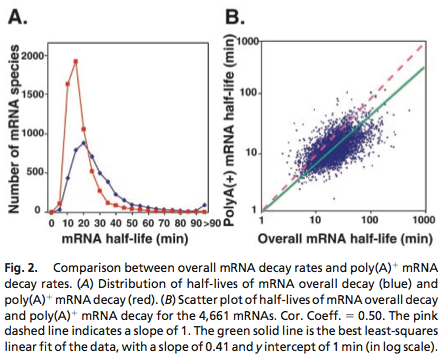
Yeast mRNA Degradation Rate
Data was obtained from the Paper (Wang et. Al 2001 [http://www.pnas.org/content/99/9/5860.long]) and processed by [http://arohatgi.info/WebPlotDigitizer/app/] to obtain raw data. Using a least-squared error approximation the distribution of the half life time in was approximated as noncentral t-distribution with parameters μ = 1.769 and ν = 20.59;.
dataGraph = [ 0.0018691649126431735,0.0016851538590669062 0.05978099456360327,0.01885629059542104 0.11548146330755026,0.21910551258377348 0.17122389948476902,0.396902157771723 0.2253457470848775,0.4417136917136917 0.2815821076690642,0.3552607791738227 0.3359848142456839,0.249812760682326 0.39216629434020744,0.19272091011221448 0.4465173486912618,0.11490683229813668 0.5026600896166115,0.07854043723608946 0.5569239808370243,0.04735863431515607 0.6111394480959699,0.04208365077930302 0.667233765059852,0.031624075102336016 0.7233280820237343,0.021164499425369035 0.777540321018582,0.017616637181854626 0.8373665112795547,0.010604847561369285 0.8897063570976615,0.008787334874291503 0.9420462029157682,0.006969822187213598 0.9999935434718044,0.005142624707842157 ]; X = round(dataGraph(:,1)*90); y = round(dataGraph(:,2)*2000); k(1) = 1.769292045467269; k(2) = 20.589996419308118; k(3) = 24852.48237036381; k=fminunc(@(z) sum((y-z(3)*nctpdf(X,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*nctpdf(X,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*nctpdf(X,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*nctpdf(X,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*nctpdf(X,z(1),z(2))).^2),k);
Yeast Protein Degradation Rate
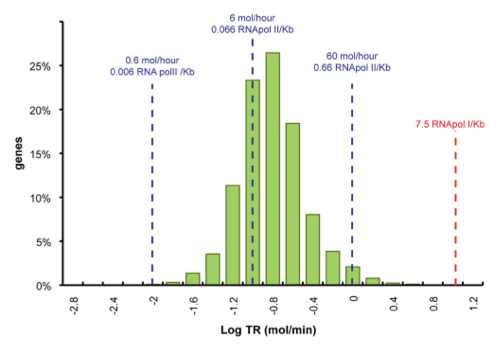
Yeast Transcription Rate Rate
Data was obtained from the Paper (Wang et. Al 2001 [http://www.pnas.org/content/99/9/5860.long]) and processed by [http://arohatgi.info/WebPlotDigitizer/app/] to obtain raw data. Using a least-squared error approximation the distribution of the transcription rate was approximated as log-normal distribution with parameters μ = -1.492 and σ = 0.661;.
dataGraph = [ -1.8,0.3442950751957339 -1.6,1.3525375039897853 -1.4,3.5492668181220783 -1.2,11.28874786429094 -1.0,23.213749272450762 -0.8,26.31522126884587 -0.6,18.273455248681024 -0.4,7.913623476840467 -0.2,3.7755111620134825 0,1.9559339854677913 0.2,0.6458759692833385 0.4,0.12767315671880167 ]; x = 10.^dataGraph(:,1); y = dataGraph(:,2); k(1) = -0.8; k(2) = 0.2; k(3) = 25; k=fminunc(@(z) sum((y-z(3)*lognpdf(x,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*lognpdf(x,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*lognpdf(x,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*lognpdf(x,z(1),z(2))).^2),k); figure(1) clf plot(linspace(-2.8,0.8,100),k(3)*lognpdf(linspace(-2.8,0.8,100),k(1),k(2)),'r-') hold on plot(x,y,'g*')
 "
"