Team:Peking/Modeling/Luminesensor/Simulation
From 2012.igem.org
TheTinaChen (Talk | contribs) |
|||
| (23 intermediate revisions not shown) | |||
| Line 8: | Line 8: | ||
</script> | </script> | ||
<div class="PKU_context floatR first"> | <div class="PKU_context floatR first"> | ||
| - | <h3 id="title1">ODE | + | <h3 id="title1">ODE Model</h3> |
<p> | <p> | ||
| - | According to the previous network and ODE model, we listed all the differential equations <!--(<a href="/Team:Peking/Modeling/Appendix/ODE">detail here</a>)--> and simulated this system with MATLAB with equations listed as | + | According to the previous network and ODE model, we listed all the differential equations <!--(<a href="/Team:Peking/Modeling/Appendix/ODE">detail here</a>)--> and simulated this system with MATLAB with equations listed as below: |
</p> | </p> | ||
<div class="floatC"> | <div class="floatC"> | ||
| - | <img src="" alt="Formulae"/> | + | <img src="/wiki/images/5/5f/Peking2012_Formula000.png" alt="Formulae" style="width:500px;"/> |
</div> | </div> | ||
<p> | <p> | ||
| Line 25: | Line 25: | ||
<td>k<sub>1</sub></td><td>3.x10<sup>-4</sup></td><td>s<sup>-1</sup></td><td>vivid decay rate constant</td><td></td> | <td>k<sub>1</sub></td><td>3.x10<sup>-4</sup></td><td>s<sup>-1</sup></td><td>vivid decay rate constant</td><td></td> | ||
</tr><tr> | </tr><tr> | ||
| - | <td>k<sub>2</sub></td><td>5.6x10<sup>-5</sup></td><td>s<sup>-1</sup></td><td>vivid dissociation rate constant</td><td><a href="#ref3" title="Mechanism-based tuning of a LOV domain photoreceptor | + | <td>k<sub>2</sub></td><td>5.6x10<sup>-5</sup></td><td>s<sup>-1</sup></td><td>vivid dissociation rate constant</td><td><a href="#ref3" title="Zoltowski, B.D., Vaccaro, B., and Crane, B.R. (2009). Mechanism-based tuning of a LOV domain photoreceptor. Nat. Chem. Biol. 5: 827: 834">[3]</a></td> |
</tr><tr> | </tr><tr> | ||
<td>k<sub>3</sub></td><td>8.x10<sup>-4</sup></td><td>s<sup>-1</sup></td><td>monomer LexA releasing rate constant from specific binding site</td><td></td> | <td>k<sub>3</sub></td><td>8.x10<sup>-4</sup></td><td>s<sup>-1</sup></td><td>monomer LexA releasing rate constant from specific binding site</td><td></td> | ||
| Line 37: | Line 37: | ||
<td>K<sub>1</sub>(Light)</td><td>1.x10<sup>+3</sup></td><td>1</td><td>equilibrium excitation constant on light</td><td></td> | <td>K<sub>1</sub>(Light)</td><td>1.x10<sup>+3</sup></td><td>1</td><td>equilibrium excitation constant on light</td><td></td> | ||
</tr><tr> | </tr><tr> | ||
| - | <td>K<sub>2</sub></td><td>7.7x10<sup>-5</sup></td><td>(n mol/L)<sup>-1</sup></td><td>vivid association equilibrium constant</td><td><a href="# | + | <td>K<sub>2</sub></td><td>7.7x10<sup>-5</sup></td><td>(n mol/L)<sup>-1</sup></td><td>vivid association equilibrium constant</td><td><a href="#ref1" title="Zoltowski, B.D., Crane, B.R.(2008). Light Activation of the LOV Protein Vivid Generates a Rapidly Exchanging Dimer.Biochemistry, 47: 7012: 7019 ">[1]</a></td> |
</tr><tr> | </tr><tr> | ||
| - | <td>K<sub>3</sub></td><td>1.x10<sup>-3</sup></td><td>(n mol/L)<sup>-1</sup></td><td>monomer LexA binding equilibrium constant with specific binding site</td><td><a href="#ref2" title=" | + | <td>K<sub>3</sub></td><td>1.x10<sup>-3</sup></td><td>(n mol/L)<sup>-1</sup></td><td>monomer LexA binding equilibrium constant with specific binding site</td><td><a href="#ref2" title="2. Mohana-Borges, R., Pacheco, A.B., Sousa, F.J., Foguel, D., Almeida, D.F., and Silva, J.L. (2000). LexA repressor forms stable dimers in solution. The role of specific DNA in tightening protein-protein interactions. J. Biol. Chem., 275: 4708: 4712">[2]</a></td> |
</tr><tr> | </tr><tr> | ||
<td>K<sub>4</sub></td><td>K<sub>2</sub>xK<sub>5</sub>/K<sub>3</sub></td><td>(n mol/L)<sup>-1</sup></td><td>binded monomer LexA association equilibrium constant</td><td>Thermal Principle</td> | <td>K<sub>4</sub></td><td>K<sub>2</sub>xK<sub>5</sub>/K<sub>3</sub></td><td>(n mol/L)<sup>-1</sup></td><td>binded monomer LexA association equilibrium constant</td><td>Thermal Principle</td> | ||
</tr><tr> | </tr><tr> | ||
| - | <td>K<sub>5</sub></td><td>1.</td><td>(n mol/L)<sup>-1</sup></td><td>dimered LexA binding equilibrium constant</td><td><a href="#ref2" title=" | + | <td>K<sub>5</sub></td><td>1.</td><td>(n mol/L)<sup>-1</sup></td><td>dimered LexA binding equilibrium constant</td><td><a href="#ref2" title="Mohana-Borges, R., Pacheco, A.B., Sousa, F.J., Foguel, D., Almeida, D.F., and Silva, J.L. (2000). LexA repressor forms stable dimers in solution. The role of specific DNA in tightening protein-protein interactions. J. Biol. Chem., 275: 4708: 4712">[2]</a></td> |
</tr><tr> | </tr><tr> | ||
| - | <td>[L<sub>G</sub>]<sub>0</sub></td><td>1000</td><td>n mol/L</td><td>initial concentration of Luminesensor in ground state</td><td></td> | + | <td>[L<sub>G</sub>]<sub>0</sub></td><td>1000</td><td>n mol/L</td><td>initial concentration of <i>Luminesensor</i> in ground state</td><td></td> |
</tr><tr> | </tr><tr> | ||
| - | <td>[L<sub>A</sub>]<sub>0</sub></td><td>0</td><td>n mol/L</td><td>initial concentration of Luminesensor in active state</td><td></td> | + | <td>[L<sub>A</sub>]<sub>0</sub></td><td>0</td><td>n mol/L</td><td>initial concentration of <i>Luminesensor</i> in active state</td><td></td> |
</tr><tr> | </tr><tr> | ||
| - | <td>[L<sub>A</sub><sup>2</sup>]<sub>0</sub></td><td>0</td><td>n mol/L</td><td>initial concentration of dimered Luminesensor</td><td></td> | + | <td>[L<sub>A</sub><sup>2</sup>]<sub>0</sub></td><td>0</td><td>n mol/L</td><td>initial concentration of dimered <i>Luminesensor</i></td><td></td> |
</tr><tr> | </tr><tr> | ||
<td>[D<sub>L</sub>]<sub>0</sub></td><td>100</td><td>n mol/L</td><td>initial concentration of free specific binding site on DNA</td><td>high-copy plasmid</td> | <td>[D<sub>L</sub>]<sub>0</sub></td><td>100</td><td>n mol/L</td><td>initial concentration of free specific binding site on DNA</td><td>high-copy plasmid</td> | ||
</tr><tr> | </tr><tr> | ||
| - | <td>[L<sub>G</sub>D<sub>L</sub>]<sub>0</sub></td><td>0</td><td>n mol/L</td><td>initial concentration of dimered Luminesensor binded Luminesensor in ground state</td><td></td> | + | <td>[L<sub>G</sub>D<sub>L</sub>]<sub>0</sub></td><td>0</td><td>n mol/L</td><td>initial concentration of dimered <i>Luminesensor</i> binded <i>Luminesensor</i> in ground state</td><td></td> |
</tr><tr> | </tr><tr> | ||
| - | <td>[L<sub>A</sub>D<sub>L</sub>]<sub>0</sub></td><td>0</td><td>n mol/L</td><td>initial concentration of dimered Luminesensor binded Luminesensor in active state</td><td></td> | + | <td>[L<sub>A</sub>D<sub>L</sub>]<sub>0</sub></td><td>0</td><td>n mol/L</td><td>initial concentration of dimered <i>Luminesensor</i> binded <i>Luminesensor</i> in active state</td><td></td> |
</tr><tr> | </tr><tr> | ||
| - | <td>[L<sub>A</sub><sup>2</sup>D<sub>L</sub>]<sub>0</sub></td><td>0</td><td>n mol/L</td><td>initial concentration of binded and dimered Luminesensor</td><td></td> | + | <td>[L<sub>A</sub><sup>2</sup>D<sub>L</sub>]<sub>0</sub></td><td>0</td><td>n mol/L</td><td>initial concentration of binded and dimered <i>Luminesensor</i></td><td></td> |
</tr> | </tr> | ||
</table> | </table> | ||
| Line 87: | Line 87: | ||
<img src="/wiki/images/7/71/Peking2012_ode_YL.png" alt="Simulation Result" style="width:600px;"/> | <img src="/wiki/images/7/71/Peking2012_ode_YL.png" alt="Simulation Result" style="width:600px;"/> | ||
<div> | <div> | ||
| - | <p class="description"> | + | <p class="description" style="text-align:center;"> |
| - | Figure | + | Figure 1. ODE Simulation Result of the prototype Luminesensor. |
</p> | </p> | ||
</div> | </div> | ||
</div> | </div> | ||
<p> | <p> | ||
| - | From the | + | From the Figure 1 above, we discovered that the activation and decay of <i>Luminesensor</i> are the key points of progress, and the activating rate is the most sensitive to light intensity. The promoter will be repressed even though the <i>Luminesensor</i> does not totally dimerized. |
</p> | </p> | ||
</div> | </div> | ||
| - | |||
<div class="PKU_context floatR"> | <div class="PKU_context floatR"> | ||
| - | <h3 id=" | + | <h3 id="title2">Stochastic Simulation</h3> |
<p> | <p> | ||
| - | In order to | + | In order to verify the robustness of <i>Luminesensor</i> function, we simulated this reaction network with a <!--<a href="/Team:Peking/Modeling/Appendix/Stochastic">-->stochastic model<!--</a>-->. By estimating the volume of a cell, we converted the concentration of a component into the number of molecules by 1 n mol/L : 1. The results are shown below: |
<p> | <p> | ||
<div class="floatC"> | <div class="floatC"> | ||
<img src="/wiki/images/c/c3/Peking2012_sto_YL.png" alt="Simulation Result" style="width:600px;"/> | <img src="/wiki/images/c/c3/Peking2012_sto_YL.png" alt="Simulation Result" style="width:600px;"/> | ||
<div> | <div> | ||
| - | <p class="description"> | + | <p class="description" style="text-align:center;"> |
| - | Figure | + | Figure 2. Stochastic Simulation Result of Prototype <i>Luminesensor</i>. |
</p> | </p> | ||
</div> | </div> | ||
</div> | </div> | ||
<p> | <p> | ||
| - | According to | + | According to Figure 2 above, noise does not influence this system. Thus the <i>Luminesensor</i> is expected to work theoretically. Besides, the average value of stochastic simulation is consistent with the result of ODE model, which in turn proves the self-consistency of our ODE model. |
</p> | </p> | ||
</div> | </div> | ||
| - | |||
<div class="PKU_context floatR"> | <div class="PKU_context floatR"> | ||
| - | <h3 id="title3">Simulation for GFP | + | <h3 id="title3">Simulation for GFP Expression <br />Regulated by the <i>Luminesensor</i></h3> |
<p> | <p> | ||
| - | + | In order to see whether our model is predictive for the downstream gene expression under control of the <i>Luminesensor</i>, transcription and translation process were incorporated into the modeling of DNA binding process. In addition, we considered the delay of translation initiation time and the growth of cell. The simulation below(Figure 3) represents the GFP expression regulated by the <i>Luminesensor</i>. After a long time in light condition, where GFP expression is inhibited, from <i>t=0h</i>, the cells are moved into dark and begin to express GFP. The GFP expression level varying with time was recorded in this simulation. | |
</p> | </p> | ||
<div class="floatC"> | <div class="floatC"> | ||
| - | <img src="/wiki/images/ | + | <img src="/wiki/images/0/07/Wild_type.png" alt="Simulation Result" style="width:500px;"/> |
<div> | <div> | ||
| - | <p class="description"> | + | <p class="description" style="text-align:center;"> |
| - | Figure | + | Figure 3. ODE Simulation Result is correspond to the experiment data of GFP expression level according to time from, which suggests that our model is effective to present the experiment situation. |
</p> | </p> | ||
</div> | </div> | ||
</div> | </div> | ||
| + | </div> | ||
| + | <div class="PKU_context floatR"> | ||
| + | <h3 id="title4">Reference</h3> | ||
| + | <p></p> | ||
| + | <ul class="refer"><li id="ref1"> | ||
| + | 1. Zoltowski, B.D., Crane, B.R.(2008). Light Activation of the LOV Protein Vivid Generates a Rapidly Exchanging Dimer. <i>Biochemistry</i>, 47: 7012: 7019 | ||
| + | </li><li id = "ref2"> | ||
| + | 2. Mohana-Borges, R., Pacheco, A.B., Sousa, F.J., Foguel, D., Almeida, D.F., and Silva, J.L. (2000). LexA repressor forms stable dimers in solution. The role of specific DNA in tightening protein-protein interactions. <i>J. Biol. Chem.</i>, 275: 4708: 4712 | ||
| + | </li><li id = "ref3"> | ||
| + | 3. Zoltowski, B.D., Vaccaro, B., and Crane, B.R. (2009). Mechanism-based tuning of a LOV domain photoreceptor. <i>Nat. Chem. Biol.</i> 5: 827: 834 | ||
| + | </li></ul> | ||
</div> | </div> | ||
</html>{{Template:Peking2012_Color_Epilogue}} | </html>{{Template:Peking2012_Color_Epilogue}} | ||
Latest revision as of 02:49, 27 September 2012
ODE Model
According to the previous network and ODE model, we listed all the differential equations and simulated this system with MATLAB with equations listed as below:

And parameters as
| Parameter | Value | Unit | Description | Source |
| k1 | 3.x10-4 | s-1 | vivid decay rate constant | |
| k2 | 5.6x10-5 | s-1 | vivid dissociation rate constant | [3] |
| k3 | 8.x10-4 | s-1 | monomer LexA releasing rate constant from specific binding site | |
| k4 | 1.x10-3 | s-1 | binded monomer LexA dissociation rate constant | |
| k5 | 1.x10-4 | s-1 | dimered LexA releasing rate constant from specific binding site | |
| K1(Dark) | 0 | 1 | equilibrium excitation constant on dark | |
| K1(Light) | 1.x10+3 | 1 | equilibrium excitation constant on light | |
| K2 | 7.7x10-5 | (n mol/L)-1 | vivid association equilibrium constant | [1] |
| K3 | 1.x10-3 | (n mol/L)-1 | monomer LexA binding equilibrium constant with specific binding site | [2] |
| K4 | K2xK5/K3 | (n mol/L)-1 | binded monomer LexA association equilibrium constant | Thermal Principle |
| K5 | 1. | (n mol/L)-1 | dimered LexA binding equilibrium constant | [2] |
| [LG]0 | 1000 | n mol/L | initial concentration of Luminesensor in ground state | |
| [LA]0 | 0 | n mol/L | initial concentration of Luminesensor in active state | |
| [LA2]0 | 0 | n mol/L | initial concentration of dimered Luminesensor | |
| [DL]0 | 100 | n mol/L | initial concentration of free specific binding site on DNA | high-copy plasmid |
| [LGDL]0 | 0 | n mol/L | initial concentration of dimered Luminesensor binded Luminesensor in ground state | |
| [LADL]0 | 0 | n mol/L | initial concentration of dimered Luminesensor binded Luminesensor in active state | |
| [LA2DL]0 | 0 | n mol/L | initial concentration of binded and dimered Luminesensor |
We specifically watched three expressions to understand the mechanism for Luminesensor.
| Expression | Description | Remark |
| rb = 1 - [DL]/[DT] | This indicates the repressing degree. | DT indicates the total specific binding sites, while the DL indicates the free ones among DT. |
| rd = 2[LA2X]/[LT] | This indicates the dimerizing degree. | LT indicates the total Luminesensor molecules, and LA2X indicates all dimered Luminesensor molecules, i.e. LA2 + LA2DL. |
| ra = ( [LAX] + 2[LA2X] ) /[LT] |
This indicates the activating degree. | LAX indicates all monomer Luminesensor molecules, i.e. LA + LADL. |
The simulation result is shown below:
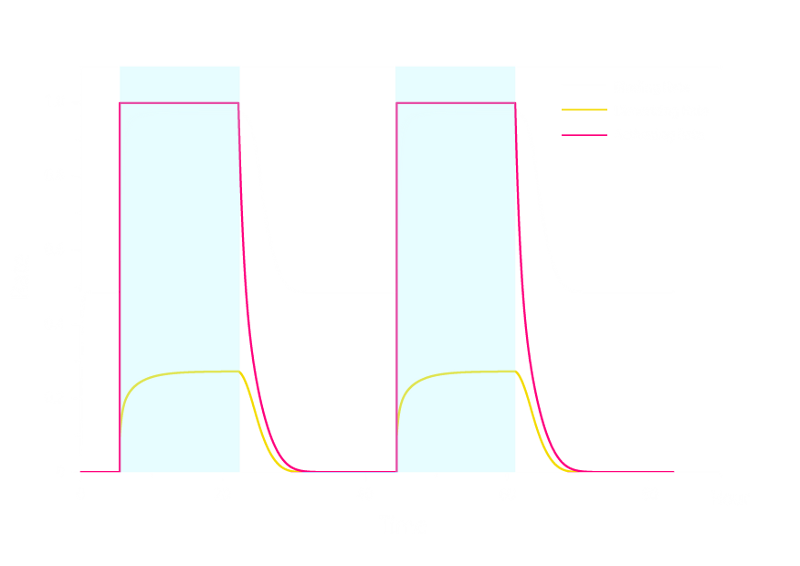
Figure 1. ODE Simulation Result of the prototype Luminesensor.
From the Figure 1 above, we discovered that the activation and decay of Luminesensor are the key points of progress, and the activating rate is the most sensitive to light intensity. The promoter will be repressed even though the Luminesensor does not totally dimerized.
Stochastic Simulation
In order to verify the robustness of Luminesensor function, we simulated this reaction network with a stochastic model. By estimating the volume of a cell, we converted the concentration of a component into the number of molecules by 1 n mol/L : 1. The results are shown below:
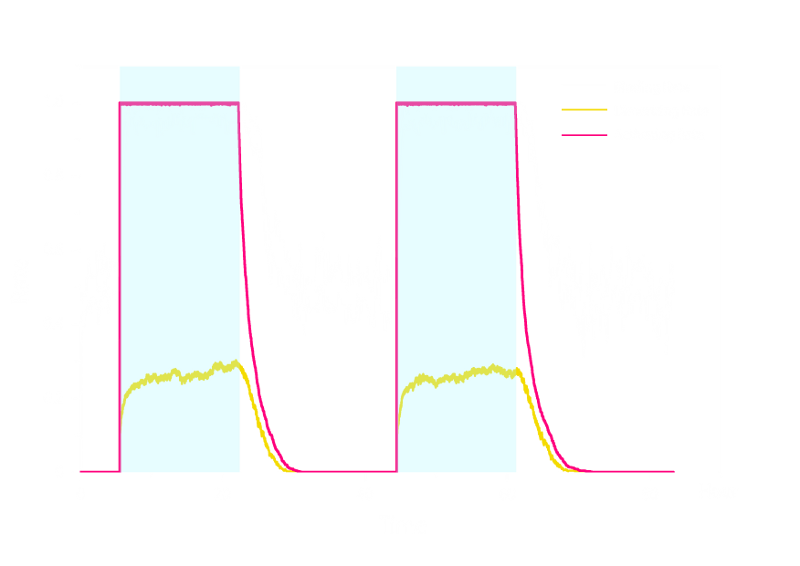
Figure 2. Stochastic Simulation Result of Prototype Luminesensor.
According to Figure 2 above, noise does not influence this system. Thus the Luminesensor is expected to work theoretically. Besides, the average value of stochastic simulation is consistent with the result of ODE model, which in turn proves the self-consistency of our ODE model.
Simulation for GFP Expression
Regulated by the Luminesensor
In order to see whether our model is predictive for the downstream gene expression under control of the Luminesensor, transcription and translation process were incorporated into the modeling of DNA binding process. In addition, we considered the delay of translation initiation time and the growth of cell. The simulation below(Figure 3) represents the GFP expression regulated by the Luminesensor. After a long time in light condition, where GFP expression is inhibited, from t=0h, the cells are moved into dark and begin to express GFP. The GFP expression level varying with time was recorded in this simulation.
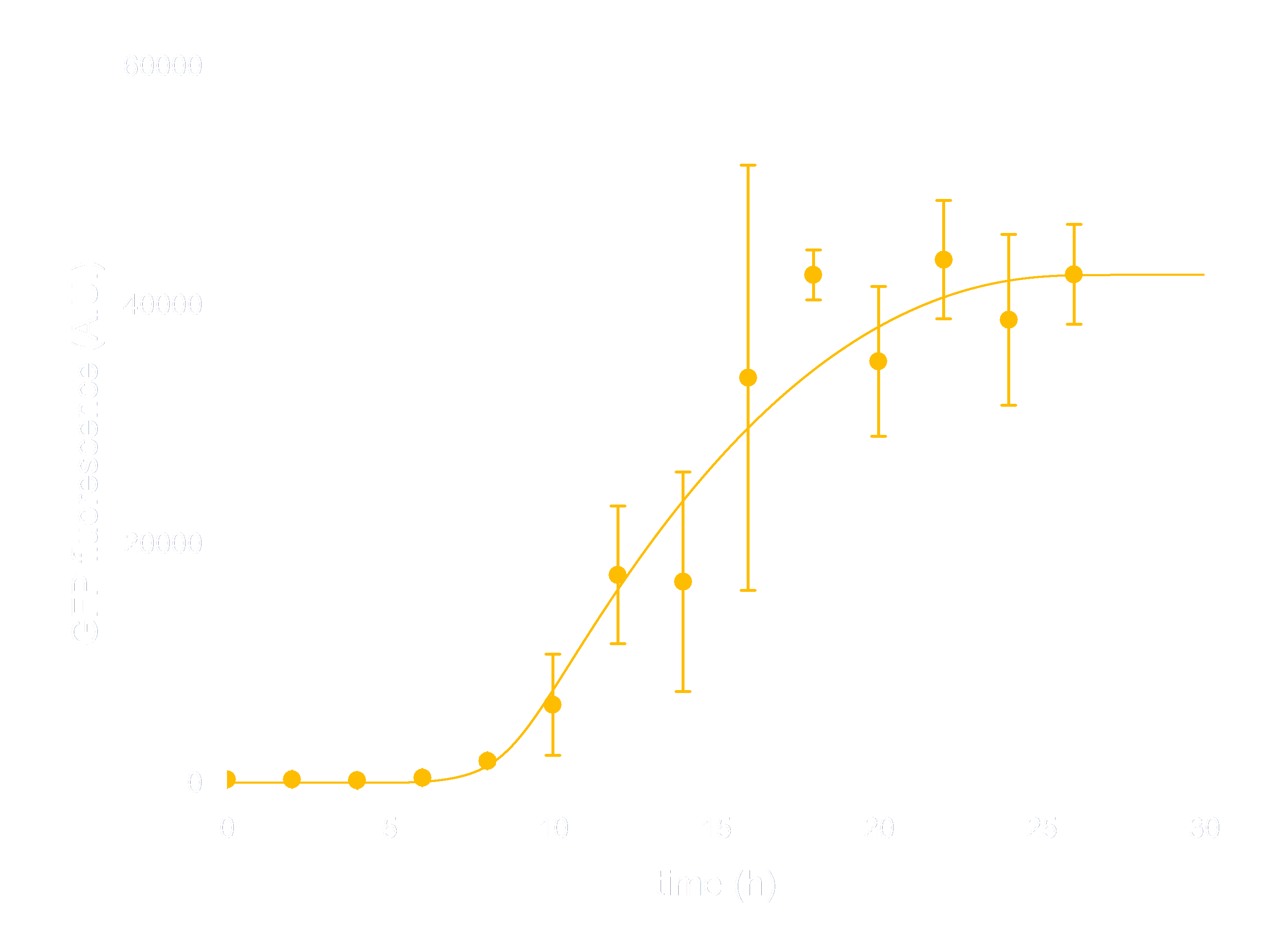
Figure 3. ODE Simulation Result is correspond to the experiment data of GFP expression level according to time from, which suggests that our model is effective to present the experiment situation.
Reference
- 1. Zoltowski, B.D., Crane, B.R.(2008). Light Activation of the LOV Protein Vivid Generates a Rapidly Exchanging Dimer. Biochemistry, 47: 7012: 7019
- 2. Mohana-Borges, R., Pacheco, A.B., Sousa, F.J., Foguel, D., Almeida, D.F., and Silva, J.L. (2000). LexA repressor forms stable dimers in solution. The role of specific DNA in tightening protein-protein interactions. J. Biol. Chem., 275: 4708: 4712
- 3. Zoltowski, B.D., Vaccaro, B., and Crane, B.R. (2009). Mechanism-based tuning of a LOV domain photoreceptor. Nat. Chem. Biol. 5: 827: 834
 "
"