Team:TU Munich/Modeling/Priors
From 2012.igem.org
(→Yeast Transcription Rate Rate) |
(→Yeast mRNA Degradation Rate) |
||
| Line 7: | Line 7: | ||
[[File:TUM12_mRNA_degradation.png|300px||right|Taken from Wang et. Al 2001]] | [[File:TUM12_mRNA_degradation.png|300px||right|Taken from Wang et. Al 2001]] | ||
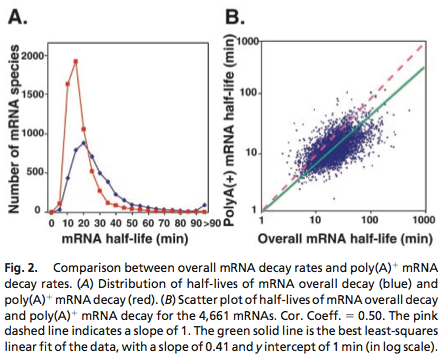
Data was obtained from the Paper (Wang et. Al 2001 [http://www.pnas.org/content/99/9/5860.long]) and processed by [http://arohatgi.info/WebPlotDigitizer/app/] to obtain raw data. | Data was obtained from the Paper (Wang et. Al 2001 [http://www.pnas.org/content/99/9/5860.long]) and processed by [http://arohatgi.info/WebPlotDigitizer/app/] to obtain raw data. | ||
| - | Using a least-squared error approximation the distribution of the half life time in was approximated as noncentral ''t''-distribution with parameters μ = 1.769 and ν = 20.59;. | + | Using a least-squared error approximation the distribution of the half life time in was approximated as '''noncentral ''t''-distribution''' with parameters μ = 1.769 and ν = 20.59;. |
<pre> | <pre> | ||
dataGraph = [ | dataGraph = [ | ||
Revision as of 15:52, 6 September 2012



Contents |
Prior Data
Yeast mRNA Degradation Rate
Data was obtained from the Paper (Wang et. Al 2001 [http://www.pnas.org/content/99/9/5860.long]) and processed by [http://arohatgi.info/WebPlotDigitizer/app/] to obtain raw data. Using a least-squared error approximation the distribution of the half life time in was approximated as noncentral t-distribution with parameters μ = 1.769 and ν = 20.59;.
dataGraph = [ 0.0018691649126431735,0.0016851538590669062 0.05978099456360327,0.01885629059542104 0.11548146330755026,0.21910551258377348 0.17122389948476902,0.396902157771723 0.2253457470848775,0.4417136917136917 0.2815821076690642,0.3552607791738227 0.3359848142456839,0.249812760682326 0.39216629434020744,0.19272091011221448 0.4465173486912618,0.11490683229813668 0.5026600896166115,0.07854043723608946 0.5569239808370243,0.04735863431515607 0.6111394480959699,0.04208365077930302 0.667233765059852,0.031624075102336016 0.7233280820237343,0.021164499425369035 0.777540321018582,0.017616637181854626 0.8373665112795547,0.010604847561369285 0.8897063570976615,0.008787334874291503 0.9420462029157682,0.006969822187213598 0.9999935434718044,0.005142624707842157 ]; X = round(dataGraph(:,1)*90); y = round(dataGraph(:,2)*2000); k(1) = 1.769292045467269; k(2) = 20.589996419308118; k(3) = 24852.48237036381; k=fminunc(@(z) sum((y-z(3)*nctpdf(X,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*nctpdf(X,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*nctpdf(X,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*nctpdf(X,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*nctpdf(X,z(1),z(2))).^2),k);
Yeast Protein Degradation Rate
For the Degradation Rate the N-end rule (Varshavsky 1997 [http://openwetware.org/images/c/c2/The_N-end_Rule_Pathway_of_Protein_Degradation.pdf]) served as approximation for the half life time. It states that the half life time in S. cerevisiae can be approximated based on the amino acid after the initial start codon.
| Residue ! Half-life | |
|---|---|
| Arg | 2 min |
| Lys, Phe, Leu, Trp, His, Asp, Asn | 3 min |
| Tyr, Gln | 10 min |
| Ile, Glu | 30 min |
| Pro | > 5 h |
| Cys, Ala, Ser, Thr, Gly, Val, Met | > 3 h |
Yeast Transcription Rate Rate
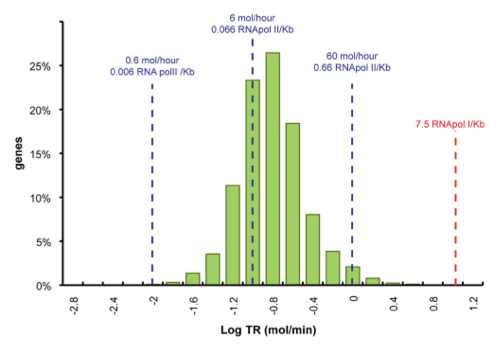
Data was obtained from the Paper (Pelechano et. Al 2010 [http://www.pnas.org/content/99/9/5860.long]) and processed by [http://arohatgi.info/WebPlotDigitizer/app/] to obtain raw data. Using a least-squared error approximation the distribution of the transcription rate was approximated as log-normal distribution with parameters μ = -1.492 and σ = 0.661;.
dataGraph = [ -1.8,0.3442950751957339 -1.6,1.3525375039897853 -1.4,3.5492668181220783 -1.2,11.28874786429094 -1.0,23.213749272450762 -0.8,26.31522126884587 -0.6,18.273455248681024 -0.4,7.913623476840467 -0.2,3.7755111620134825 0,1.9559339854677913 0.2,0.6458759692833385 0.4,0.12767315671880167 ]; x = 10.^dataGraph(:,1); y = dataGraph(:,2); k(1) = -0.8; k(2) = 0.2; k(3) = 25; k=fminunc(@(z) sum((y-z(3)*lognpdf(x,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*lognpdf(x,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*lognpdf(x,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*lognpdf(x,z(1),z(2))).^2),k); figure(1) clf plot(linspace(-2.8,0.8,100),k(3)*lognpdf(linspace(-2.8,0.8,100),k(1),k(2)),'r-') hold on plot(x,y,'g*')
 "
"