Team:Peking/Modeling/Ring/Simulation
From 2012.igem.org
| Line 65: | Line 65: | ||
</table> | </table> | ||
</div> | </div> | ||
| - | |||
| - | |||
<p> | <p> | ||
The simulation result is shown below: | The simulation result is shown below: | ||
| Line 84: | Line 82: | ||
</p> | </p> | ||
</div> | </div> | ||
| + | |||
<div class="PKU_context floatR"> | <div class="PKU_context floatR"> | ||
<h3 id="title2">Parameter Analysis</h3> | <h3 id="title2">Parameter Analysis</h3> | ||
<p> | <p> | ||
| - | + | After modeling the prototype <i>Luminesensor</i>, we attempted to optimize it in a rational way. We have tuned the parameters both up and down, one by one, and finally discovered four parameters which predominantly influence the performance of the <i>Luminesensor</i>. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
</p> | </p> | ||
| + | <table style="width:600px;"><tr> | ||
| + | <td>Function</td> | ||
| + | <td>Parameter</td> | ||
| + | <td>Description</td> | ||
| + | <td>Remark</td> | ||
| + | </tr><tr> | ||
| + | <td rowspan="2">Reduce responsing time</td> | ||
| + | <td>k<sub>1</sub></td><td>Vivid lighting decay rate constant</td><td>Mainly on process from Light to Dark</td> | ||
| + | </tr><tr> | ||
| + | <td>k<sub>3</sub></td><td>rate constant of monomer LexA releasing from specific binding site</td><td></td> | ||
| + | </tr><tr> | ||
| + | <td rowspan="2">Enhance contrast</td> | ||
| + | <td>K<sub>2</sub></td><td>Vivid association equilibrium constant</td><td>More dimerization provides more binding opportunity</td> | ||
| + | </tr><tr> | ||
| + | <td>K<sub>5</sub></td><td>dimered LexA binding equilibrium constant</td><td>More binding affinity</td> | ||
| + | </tr></table> | ||
| + | |||
</div> | </div> | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
<div class="PKU_context floatR"> | <div class="PKU_context floatR"> | ||
<h3 id="title4">Reference</h3> | <h3 id="title4">Reference</h3> | ||
Revision as of 02:53, 23 October 2012
ODE Model
According to the previous circuit and ODE model, we listed all the differential equations and simulated this system with MATLAB with equations listed as below:
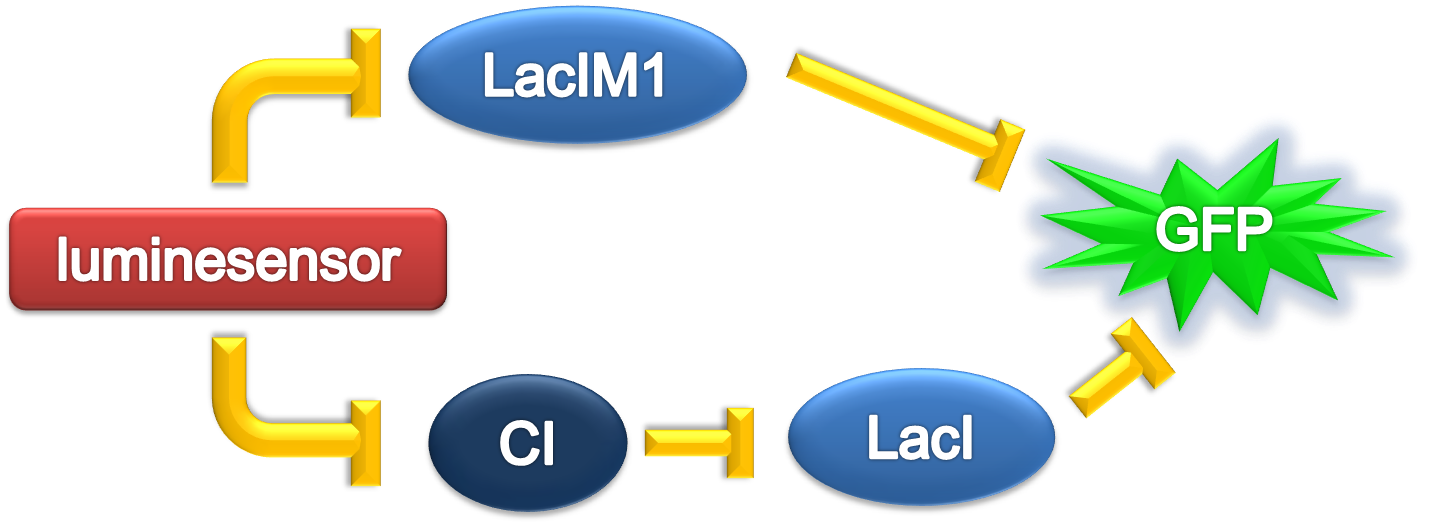
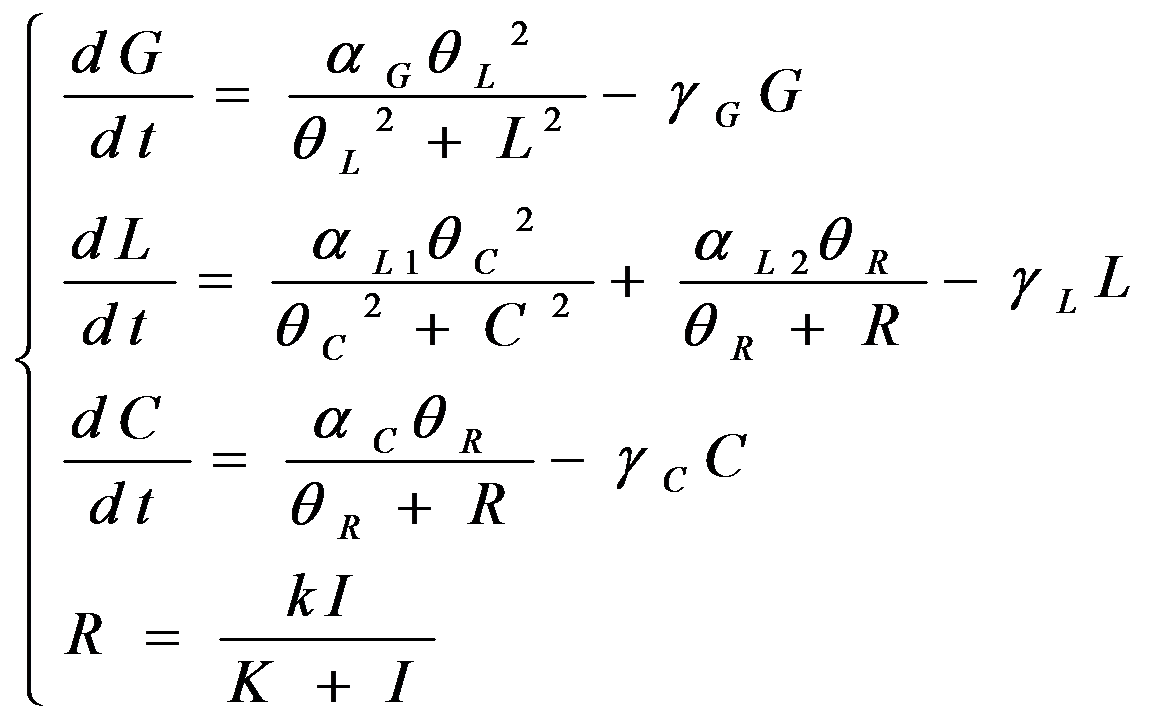
And parameters as
| Parameter | Value | Unit | Description | Source |
| k1 | 3.x10-4 | s-1 | vivid decay rate constant | |
| k2 | 5.6x10-5 | s-1 | vivid dissociation rate constant | [3] |
| k3 | 8.x10-4 | s-1 | monomer LexA releasing rate constant from specific binding site | |
| k4 | 1.x10-3 | s-1 | binded monomer LexA dissociation rate constant | |
| k5 | 1.x10-4 | s-1 | dimered LexA releasing rate constant from specific binding site | |
| K1(Dark) | 0 | 1 | equilibrium excitation constant on dark | |
| K1(Light) | 1.x10+3 | 1 | equilibrium excitation constant on light | |
| K2 | 7.7x10-5 | (n mol/L)-1 | vivid association equilibrium constant | [1] |
| K3 | 1.x10-3 | (n mol/L)-1 | monomer LexA binding equilibrium constant with specific binding site | [2] |
| K4 | K2xK5/K3 | (n mol/L)-1 | binded monomer LexA association equilibrium constant | Thermal Principle |
| K5 | 1. | (n mol/L)-1 | dimered LexA binding equilibrium constant | [2] |
| [LG]0 | 1000 | n mol/L | initial concentration of Luminesensor in ground state | |
| [LA]0 | 0 | n mol/L | initial concentration of Luminesensor in active state | |
| [LA2]0 | 0 | n mol/L | initial concentration of dimered Luminesensor | |
| [DL]0 | 100 | n mol/L | initial concentration of free specific binding site on DNA | high-copy plasmid |
| [LGDL]0 | 0 | n mol/L | initial concentration of dimered Luminesensor binded Luminesensor in ground state | |
| [LADL]0 | 0 | n mol/L | initial concentration of dimered Luminesensor binded Luminesensor in active state | |
| [LA2DL]0 | 0 | n mol/L | initial concentration of binded and dimered Luminesensor |
The simulation result is shown below:
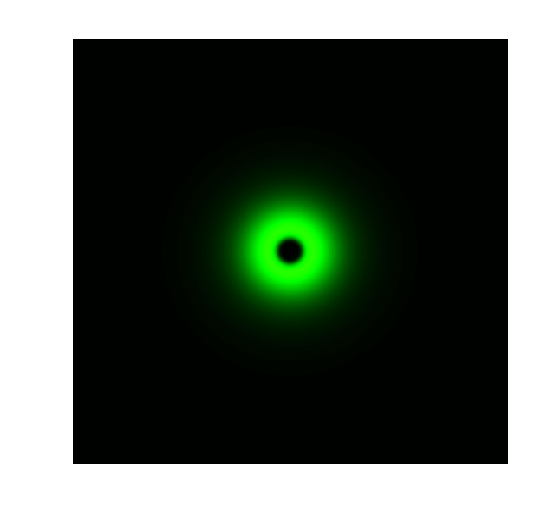
Figure 1. ODE Simulation in a plate of the ring-like pattern formation.
Figure 2. ODE Simulation for the radial expression amplitude of the ring-like pattern formation.
From the Figure 1 above, we discovered that the activation and decay of Luminesensor are the key points of progress, and the activating rate is the most sensitive to light intensity. The promoter will be repressed even though the Luminesensor does not totally dimerized.
Parameter Analysis
After modeling the prototype Luminesensor, we attempted to optimize it in a rational way. We have tuned the parameters both up and down, one by one, and finally discovered four parameters which predominantly influence the performance of the Luminesensor.
| Function | Parameter | Description | Remark |
| Reduce responsing time | k1 | Vivid lighting decay rate constant | Mainly on process from Light to Dark |
| k3 | rate constant of monomer LexA releasing from specific binding site | ||
| Enhance contrast | K2 | Vivid association equilibrium constant | More dimerization provides more binding opportunity |
| K5 | dimered LexA binding equilibrium constant | More binding affinity |
Reference
- 1. Zoltowski, B.D., Crane, B.R.(2008). Light Activation of the LOV Protein Vivid Generates a Rapidly Exchanging Dimer. Biochemistry, 47: 7012: 7019
- 2. Mohana-Borges, R., Pacheco, A.B., Sousa, F.J., Foguel, D., Almeida, D.F., and Silva, J.L. (2000). LexA repressor forms stable dimers in solution. The role of specific DNA in tightening protein-protein interactions. J. Biol. Chem., 275: 4708: 4712
- 3. Zoltowski, B.D., Vaccaro, B., and Crane, B.R. (2009). Mechanism-based tuning of a LOV domain photoreceptor. Nat. Chem. Biol. 5: 827: 834
 "
"