Team:TU-Eindhoven/LEC/Lab
From 2012.igem.org
| Line 12: | Line 12: | ||
[[File:Yeast_transformation.jpg|300px|link=]] | [[File:Yeast_transformation.jpg|300px|link=]] | ||
<p>A weekly item on our schedule was yeast transformation. It takes about a week to complete a transformation, that is, to add one plasmid to an existing strain variant. Because success is not guaranteed we choose to introduce plasmids in various orders in parallel. This resulted in many variants that we assigned a unique number for convenience. The same number was used on plates, cultures and cryostocks. In total 49 variants were made.</p> | <p>A weekly item on our schedule was yeast transformation. It takes about a week to complete a transformation, that is, to add one plasmid to an existing strain variant. Because success is not guaranteed we choose to introduce plasmids in various orders in parallel. This resulted in many variants that we assigned a unique number for convenience. The same number was used on plates, cultures and cryostocks. In total 49 variants were made.</p> | ||
| - | [[File:Transformation chart.png| | + | |
| + | <p>To reach the aim of this project and prepare a multi-colored screen, three kind of cells, each with a different color, are needed. Therefore, one single plasmid strain would have to consist of both calcium channel parts MID1 and CCH1 sequences and one of the fluorescent GECO protein sequence. Each of these DNA sequences, coding for the three different parts, is transformed into a yeast plasmid (InvSC1). Moreover, each of these sequences contain a small sequence coding for an amino acid, causing the yeast strain to be able to synthesize this specific amino acid. The synthesis of this specific amino acid will allow strain selection by retaining this amino acid from the culture media. Cells containing the coding CCH1, MID1 and GECO sequence are able to synthesize leucine, uracil and tryptophan respectively. </p> | ||
| + | <p>Looking at Scheme 1, which represents the transformations needed to come to the desired yeast strain, it seems and is logical to start with the MID1 or CCH1 vectors. By doing so, less transformations can be performed due to the transformation intensive introduction of the three different GECO proteins at last. To make sure that the transformations would lead to the desired strain, it was decided to start the first step with transforming all possibilities. When step one was transformed successfully, visualized by the arrows in Scheme 1, the transformation of step two was performed and so on.</p> | ||
| + | [[File:Transformation chart.png|500px|link=]] | ||
| + | <p>The several transformation steps are clarified in Scheme 1. The arrows represent a successful transformation as at the same time the lack of an arrow represents a transformation failure. Of course, not every transformation was needed to obtain the desired strain. E.g. in step one the transformation of the R-GECO failed, however, the strain containing the R-GECO was obtained in step two due to other successful transformations. | ||
| + | <p>As can be seen, in step 2 the first co-transformation are performed. This brought some complications because the regular transformation protocol did not work for these co-transformations. For the co- and co-co-transformations (performed in step 3) we used another high-efficiency transformation [[Team:TU-Eindhoven/Protocols|protocol]] which worked out successfully. After 3 steps of transformations, a couple of different transformation paths reached the three desired strains a complete calcium channel and one of the three GECO proteins.</p> | ||
| + | |||
<p>Initially, yeast transformations failed or yielded only several colonies, a lot less than expected. A review of transformation protocols showed that the shock protocol we used should be more efficient than electroporation or other common methods. We tried to transform another yeast strain with our plasmids which yielded the expected high transformation efficiency. Unfortunately that strain was not compatible with the auxotrophic markers on the plasmids we wanted to introduce. Our problem was remedied by using more plasmid DNA for the transformation, as described in our [[Team:TU-Eindhoven/Protocols|modified yeast transformation protocol]].</p> | <p>Initially, yeast transformations failed or yielded only several colonies, a lot less than expected. A review of transformation protocols showed that the shock protocol we used should be more efficient than electroporation or other common methods. We tried to transform another yeast strain with our plasmids which yielded the expected high transformation efficiency. Unfortunately that strain was not compatible with the auxotrophic markers on the plasmids we wanted to introduce. Our problem was remedied by using more plasmid DNA for the transformation, as described in our [[Team:TU-Eindhoven/Protocols|modified yeast transformation protocol]].</p> | ||
Revision as of 01:02, 27 September 2012

GECO protein expression and isolation

The three GECOs have been expressed in E. coli BL21 to yield a large amount of the proteins for characterization. Cells were cultured in large Erlenmeyer flasks. The results were visually impressive: After lysis the flasks turned vibrantly red and green respectively. The blue GECO however looked much the same as the green GECO.
More information about the GECOs can be found at Our BioBricks.
Yeast transformation
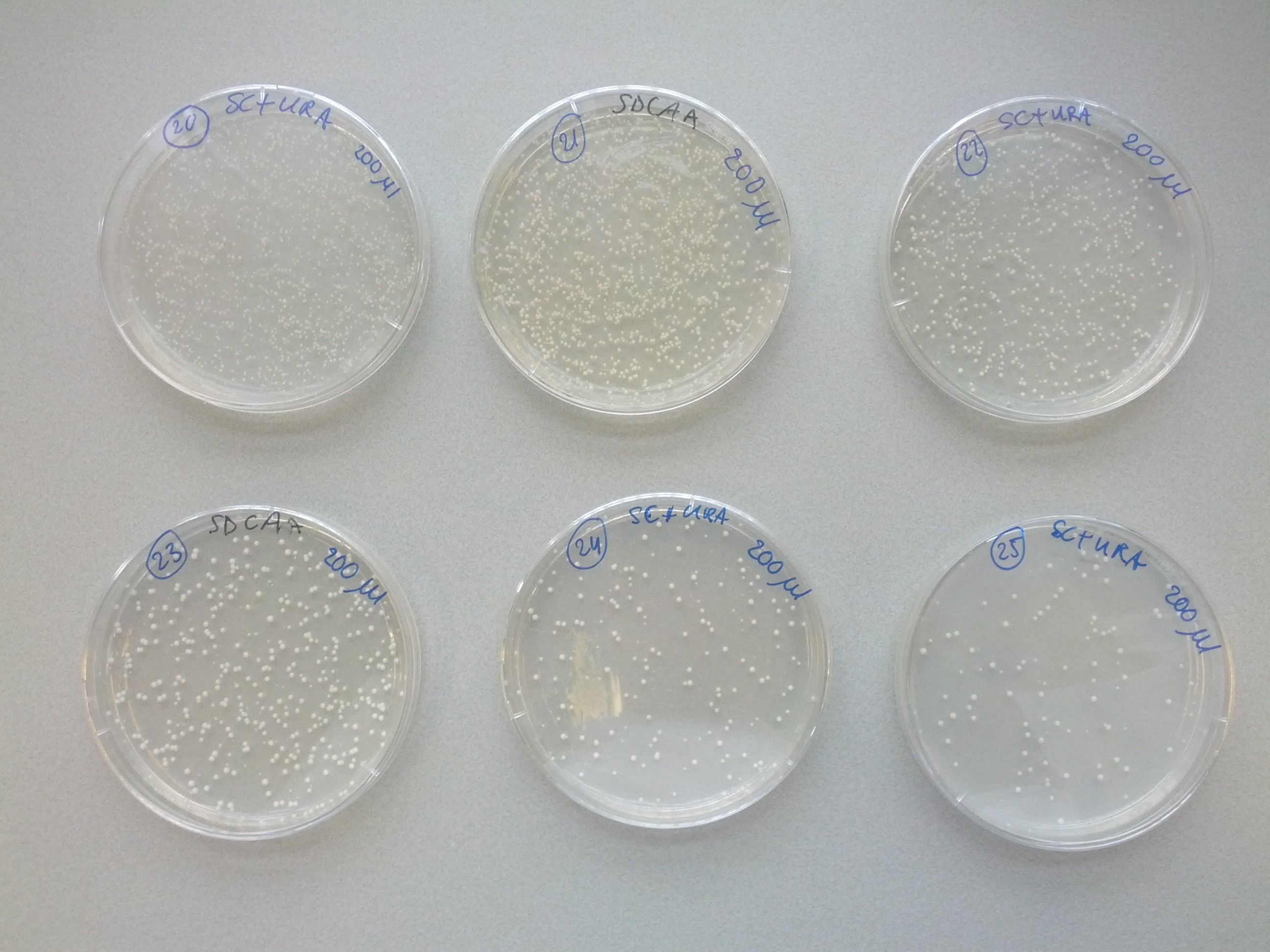
A weekly item on our schedule was yeast transformation. It takes about a week to complete a transformation, that is, to add one plasmid to an existing strain variant. Because success is not guaranteed we choose to introduce plasmids in various orders in parallel. This resulted in many variants that we assigned a unique number for convenience. The same number was used on plates, cultures and cryostocks. In total 49 variants were made.
To reach the aim of this project and prepare a multi-colored screen, three kind of cells, each with a different color, are needed. Therefore, one single plasmid strain would have to consist of both calcium channel parts MID1 and CCH1 sequences and one of the fluorescent GECO protein sequence. Each of these DNA sequences, coding for the three different parts, is transformed into a yeast plasmid (InvSC1). Moreover, each of these sequences contain a small sequence coding for an amino acid, causing the yeast strain to be able to synthesize this specific amino acid. The synthesis of this specific amino acid will allow strain selection by retaining this amino acid from the culture media. Cells containing the coding CCH1, MID1 and GECO sequence are able to synthesize leucine, uracil and tryptophan respectively.
Looking at Scheme 1, which represents the transformations needed to come to the desired yeast strain, it seems and is logical to start with the MID1 or CCH1 vectors. By doing so, less transformations can be performed due to the transformation intensive introduction of the three different GECO proteins at last. To make sure that the transformations would lead to the desired strain, it was decided to start the first step with transforming all possibilities. When step one was transformed successfully, visualized by the arrows in Scheme 1, the transformation of step two was performed and so on.

The several transformation steps are clarified in Scheme 1. The arrows represent a successful transformation as at the same time the lack of an arrow represents a transformation failure. Of course, not every transformation was needed to obtain the desired strain. E.g. in step one the transformation of the R-GECO failed, however, the strain containing the R-GECO was obtained in step two due to other successful transformations. <p>As can be seen, in step 2 the first co-transformation are performed. This brought some complications because the regular transformation protocol did not work for these co-transformations. For the co- and co-co-transformations (performed in step 3) we used another high-efficiency transformation protocol which worked out successfully. After 3 steps of transformations, a couple of different transformation paths reached the three desired strains a complete calcium channel and one of the three GECO proteins.
Initially, yeast transformations failed or yielded only several colonies, a lot less than expected. A review of transformation protocols showed that the shock protocol we used should be more efficient than electroporation or other common methods. We tried to transform another yeast strain with our plasmids which yielded the expected high transformation efficiency. Unfortunately that strain was not compatible with the auxotrophic markers on the plasmids we wanted to introduce. Our problem was remedied by using more plasmid DNA for the transformation, as described in our modified yeast transformation protocol.
Device tests
The device was tested with yeast containing all three plasmids, thus over expressing channels and expressing GECOs. A quick inspection with the naked eye did not show any sign of visual light coming off the device, not even in a dark room. We think that the yeast was not concentrated enough to show a clear response.
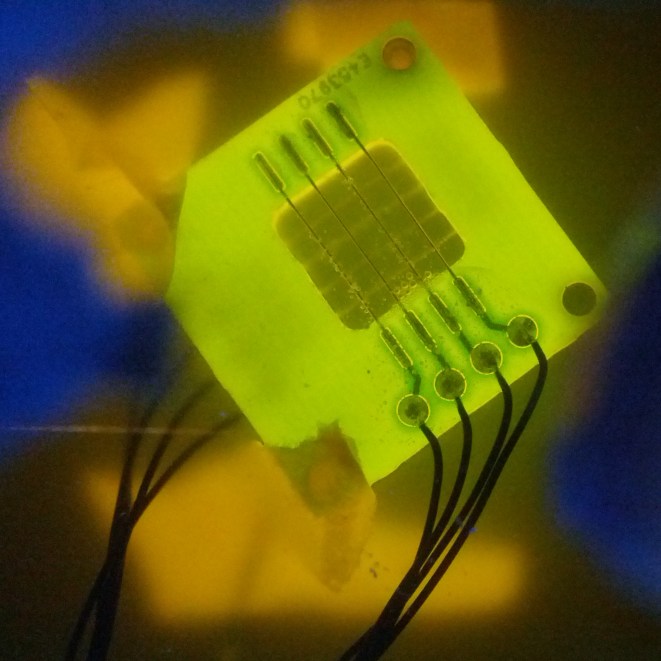

Spectrophotometry
Plasmid construction
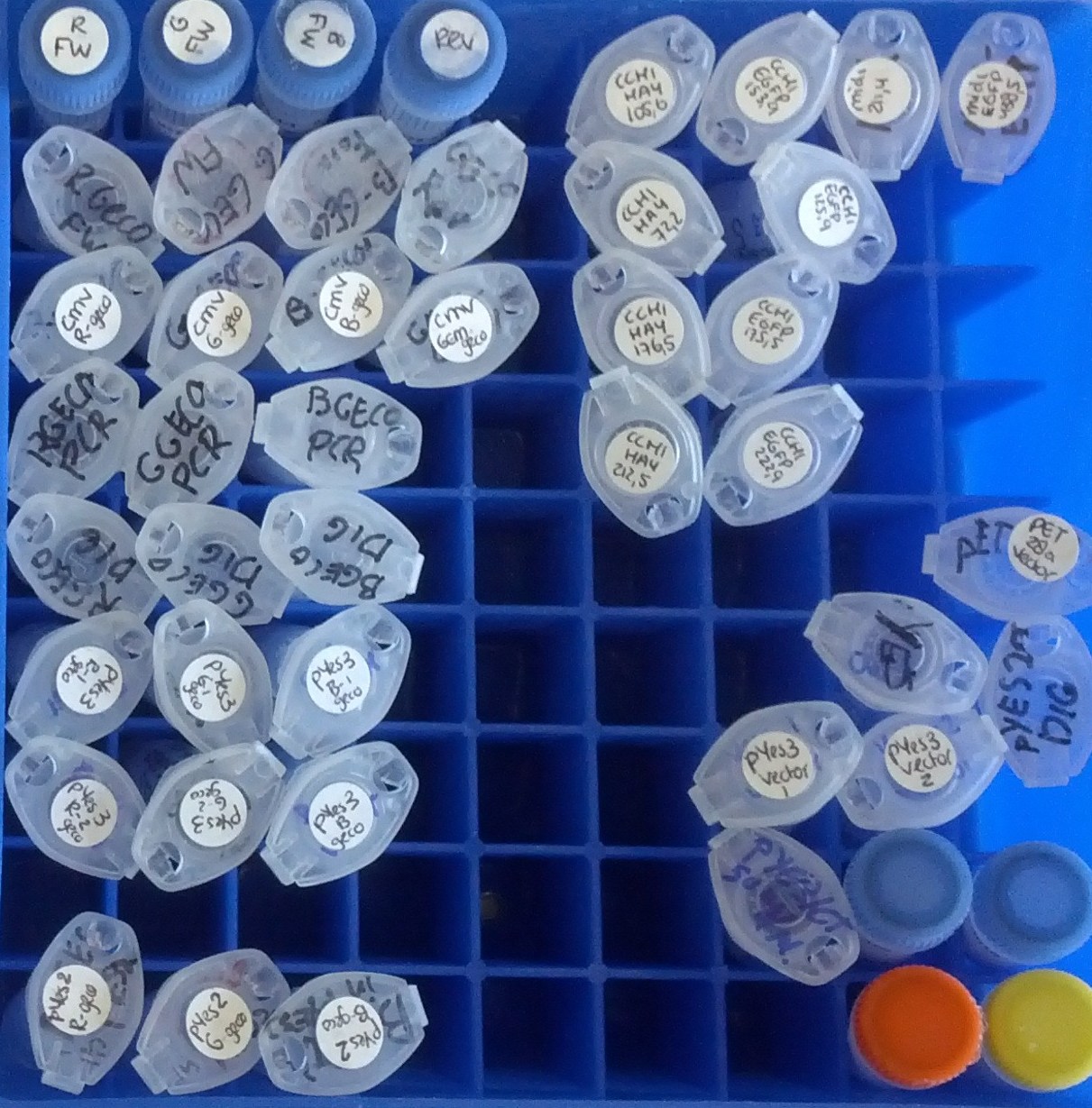

The following non-BioBrick plasmids have been constructed:
| name | vector | inserts | description |
|---|---|---|---|
| pYES2/CT-X-GECO | pYES2/CT | R-GECO1, G-GECO1.1, B-GECO1 | High copy-number shuttle vector |
| pYES3/CT-X-GECO | pYES3/CT | R-GECO1, G-GECO1.1, B-GECO1 | High copy-number shuttle vector |
| pET28a | X-GECO/pET28a | R-GECO1, G-GECO1.1, B-GECO1 | Expression vector for E. coli |
The following plasmids were received from addgene.org:
| name | vector | inserts | description |
|---|---|---|---|
| CMV-X-GECO | CMV | R-GECO1, G-GECO1.1, B-GECO1 | Shipping vector use by addgene.org, CMV vector was not used in this project |
These plasmids were kindly donated by H. Iida & K. Iida:
| name | vector | inserts | description |
|---|---|---|---|
| pBCT -CCH1H | pBCT | CCH1H | Low copy-number shuttle vector, encodes the CCH1 protein |
| pBCT-CCH1H-HA4 | pBCT | CCH1H-HA4 | Low copy-number shuttle vector, encodes HA4-tagged CCH1 |
| pBCT-CCH1H-EGFP | pBCT | CCH1H-EGFP | Low copy-number shuttle vector, encodes EGFP-tagged CCH1 |
| YCpT-MID1 | YCpT | MID1 | Low copy-number shuttle vector, encodes the MID1 protein |
| YCpT-MID1-EGFP | YCpT | MID1-EGFP | Low copy-number shuttle vector, encodes EGFP-tagged MID1 |
References
 "
"