Team:TU Munich/Modeling/Priors
From 2012.igem.org
(→Reference) |
|||
| Line 109: | Line 109: | ||
=Reference= | =Reference= | ||
---- | ---- | ||
| - | *[ | + | *[Pelechano et al., 2010] Pelechano, V., Ch ́avez, S., and P ́erez-Ort ́ın, J. E. (2010). A complete set of nascent transcription rates for yeast genes. ''PLoS One'', 5(11):e15442. |
| - | *[[http://arohatgi.info/webplotdigitizer/app/ | + | *[[http://arohatgi.info/webplotdigitizer/app/ Rohatgi, 2012]] Rohatgi, A. (2012). http://arohatgi.info/webplotdigitizer/app/. |
| - | *[[http://www2.maths.ox.ac.uk/chebfun/ | + | *[[http://www2.maths.ox.ac.uk/chebfun/ Trefethen, 2012]] Trefethen, N. (2012). http://www2.maths.ox.ac.uk/chebfun/. |
| - | *[ | + | *[Varshavsky, 1997] Varshavsky, A. (1997). The n-end rule pathway of protein degradation. Genes Cells, 2(1):13–28. |
| - | *[ | + | *[Wang et al., 2002] Wang, Y., Liu, C. L., Storey, J. D., Tibshirani, R. J., Herschlag, D., and Brown, P. O. (2002). Precision and functional specificity in mrna decay. ''Proc Natl Acad Sci U S A'', 99(9):5860–5. |
Revision as of 21:06, 19 September 2012



Contents |
Prior Data
Yeast mRNA Degradation Rate
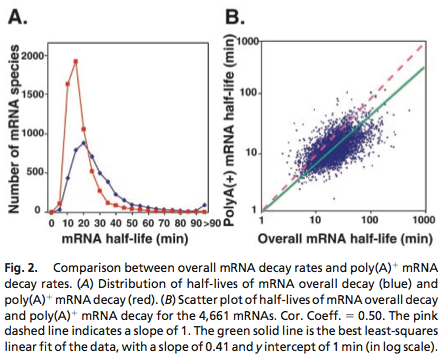
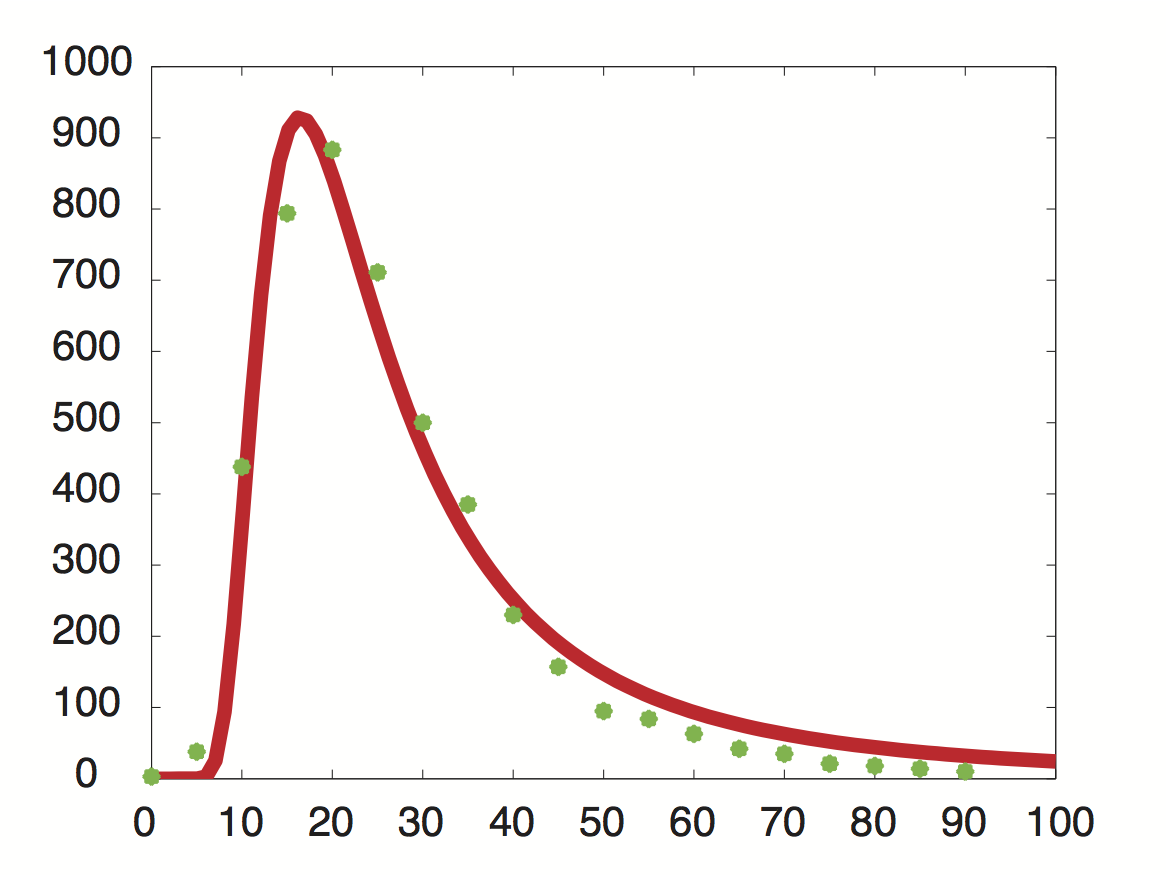
Data was obtained from the Paper (Wang et al., 2002 [http://www.pnas.org/content/99/9/5860.long]) and processed by [http://arohatgi.info/WebPlotDigitizer/app/] to obtain raw data. Using a least-squared error approximation the distribution of the half life time in was approximated as noncentral t-distribution with parameters μ = 1.769 and ν = 20.59.
dataGraph = [ 0.0018691649126431735,0.0016851538590669062 0.05978099456360327,0.01885629059542104 0.11548146330755026,0.21910551258377348 0.17122389948476902,0.396902157771723 0.2253457470848775,0.4417136917136917 0.2815821076690642,0.3552607791738227 0.3359848142456839,0.249812760682326 0.39216629434020744,0.19272091011221448 0.4465173486912618,0.11490683229813668 0.5026600896166115,0.07854043723608946 0.5569239808370243,0.04735863431515607 0.6111394480959699,0.04208365077930302 0.667233765059852,0.031624075102336016 0.7233280820237343,0.021164499425369035 0.777540321018582,0.017616637181854626 0.8373665112795547,0.010604847561369285 0.8897063570976615,0.008787334874291503 0.9420462029157682,0.006969822187213598 0.9999935434718044,0.005142624707842157 ]; %scale the data X = round(dataGraph(:,1)*90); y = round(dataGraph(:,2)*2000); k(1) = 1.769292045467269; k(2) = 20.589996419308118; k(3) = 24852.48237036381; k=fminunc(@(z) sum((y-z(3)*nctpdf(X,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*nctpdf(X,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*nctpdf(X,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*nctpdf(X,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*nctpdf(X,z(1),z(2))).^2),k);
The matlab nctpdf.m script needs to compute the limit of an infinite series to calculate the probabilities for the noncentral t-distribution. As this the function will be called several million times during the generation of samples, the computation was quite time consuming and the funtion was approximated using chebyshev interpolation [http://www.mathworks.de/matlabcentral/fileexchange/23972-chebfun].
Yeast Protein Degradation Rate
For the Degradation Rate the N-end rule (Varshavsky et al., 1997 [http://openwetware.org/images/c/c2/The_N-end_Rule_Pathway_of_Protein_Degradation.pdf]) served as approximation for the half life time. It states that the half life time in S. cerevisiae can be approximated based on the amino acid after the initial start codon.
| Residue ! Half-life | |
|---|---|
| Arg | 2 min |
| Lys, Phe, Leu, Trp, His, Asp, Asn | 3 min |
| Tyr, Gln | 10 min |
| Ile, Glu | 30 min |
| Pro | > 5 h |
| Cys, Ala, Ser, Thr, Gly, Val, Met | > 30 h |
As these values do not give enough information to infer a proper distribution, only the two lower bounds 5 h and 30 h will serve as approximate lower bounds for the optimization routines.
Yeast Transcription Rate
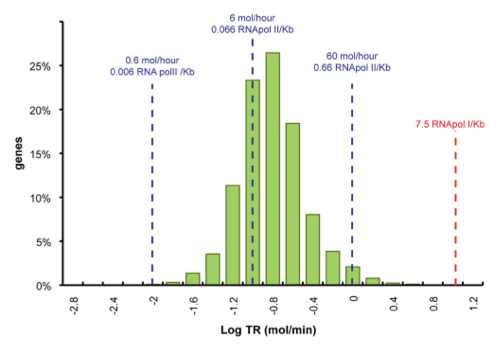
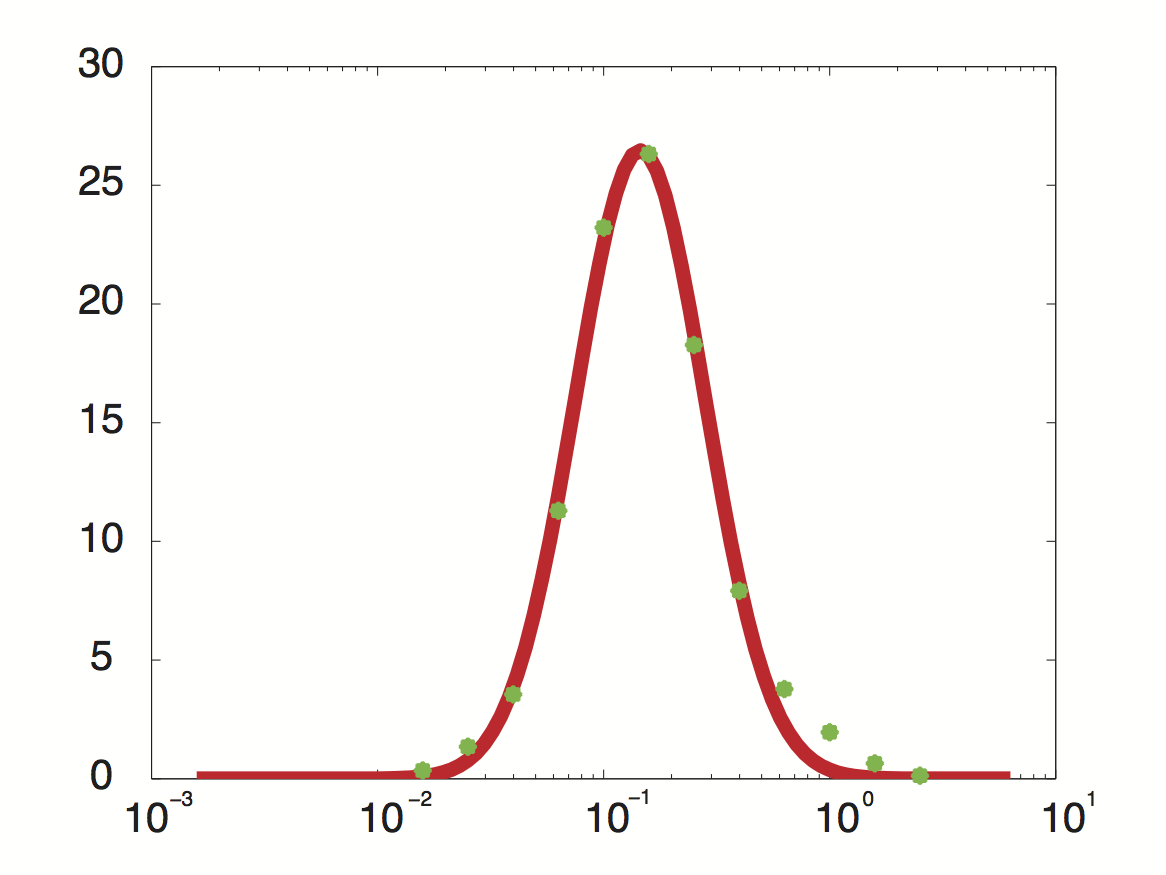
Data was obtained from the Paper (Pelechano et al., 2010 [http://www.pnas.org/content/99/9/5860.long]) and processed by http://arohatgi.info/WebPlotDigitizer/app/ 2 to obtain raw data. Using a least-squared error approximation the distribution of the transcription rate was approximated as log-normal distribution with parameters μ = -1.492 and σ = 0.661;.
dataGraph = [ -1.8,0.3442950751957339 -1.6,1.3525375039897853 -1.4,3.5492668181220783 -1.2,11.28874786429094 -1.0,23.213749272450762 -0.8,26.31522126884587 -0.6,18.273455248681024 -0.4,7.913623476840467 -0.2,3.7755111620134825 0,1.9559339854677913 0.2,0.6458759692833385 0.4,0.12767315671880167 ]; x = 10.^dataGraph(:,1); y = dataGraph(:,2); k(1) = -0.8; k(2) = 0.2; k(3) = 25; k=fminunc(@(z) sum((y-z(3)*lognpdf(x,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*lognpdf(x,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*lognpdf(x,z(1),z(2))).^2),k); k=fminunc(@(z) sum((y-z(3)*lognpdf(x,z(1),z(2))).^2),k);
Reference
- [Pelechano et al., 2010] Pelechano, V., Ch ́avez, S., and P ́erez-Ort ́ın, J. E. (2010). A complete set of nascent transcription rates for yeast genes. PLoS One, 5(11):e15442.
- http://arohatgi.info/webplotdigitizer/app/ Rohatgi, 2012 Rohatgi, A. (2012). http://arohatgi.info/webplotdigitizer/app/.
- http://www2.maths.ox.ac.uk/chebfun/ Trefethen, 2012 Trefethen, N. (2012). http://www2.maths.ox.ac.uk/chebfun/.
- [Varshavsky, 1997] Varshavsky, A. (1997). The n-end rule pathway of protein degradation. Genes Cells, 2(1):13–28.
- [Wang et al., 2002] Wang, Y., Liu, C. L., Storey, J. D., Tibshirani, R. J., Herschlag, D., and Brown, P. O. (2002). Precision and functional specificity in mrna decay. Proc Natl Acad Sci U S A, 99(9):5860–5.
 "
"