Team:Wageningen UR/Journal/week15
From 2012.igem.org
(→Lab work) |
(→Lab work) |
||
| Line 58: | Line 58: | ||
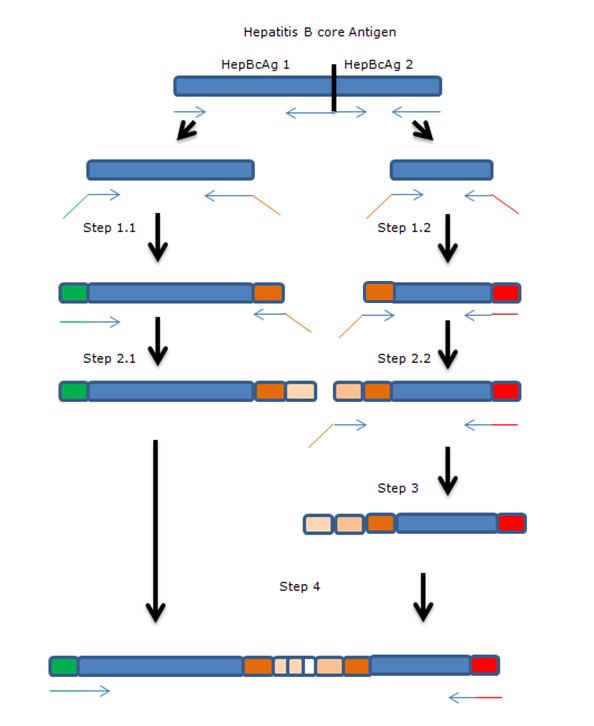
The gene of HepBcAg will be ‘cut’ in two parts using two sets of primers after which the parts will be extended towards each other by using overhanging primers. The overhang codes for linkers and the E-coil. | The gene of HepBcAg will be ‘cut’ in two parts using two sets of primers after which the parts will be extended towards each other by using overhanging primers. The overhang codes for linkers and the E-coil. | ||
| - | Legend: | + | |
| - | Blue: Hepatitis B core antigen gene | + | '''Legend:''' |
| - | Green: iGEM standard 10 prefix | + | * Blue: Hepatitis B core antigen gene |
| - | Red: iGEM standard 10 suffix | + | * Green: iGEM standard 10 prefix |
| - | Orange: insert containing linkers and K-coil | + | * Red: iGEM standard 10 suffix |
| - | Arrows: primers, with overhang. | + | * Orange: insert containing linkers and K-coil |
| + | * Arrows: primers, with overhang. | ||
[[File:PCRHepBout.JPG|500px|center|thumb|<p align="justify">''Overview the PCR steps needed for the modification inside the external loop of HepB'</p>]] | [[File:PCRHepBout.JPG|500px|center|thumb|<p align="justify">''Overview the PCR steps needed for the modification inside the external loop of HepB'</p>]] | ||
Revision as of 12:08, 19 September 2012
week 15: 6 august - 12 august
Office work
Lab work
New electrocompetent DH5 Alfa and Mach1 E. coli cells were prepared. Transformation efficiency was determined: DH5 Alfa: 10^8 Mach1: 10^9
TuYV
- Bricking:
Last week's minipreps were digested (EcoRI, PstI), the inserts ligated into the chloramphenicol backbone from BBa_J04450 and transformed into our electrocompetent DH5 Alfa E. coli. A PCR check gave negative results, but miniprep was performed anyway.
- Expression:
Last week's minipreps were digested (XbaI, Spe1), the inserts ligated into the promotor&RBS-containing backbone from BBa_J04500 and transformed into our electrocompetent BL21 E. coli. ALthough there were colonies colonies on the sample plate, but we got negative results from both colony PCR and growing in LB+ antibiotic medium, so the entire procedure was done again on saturday.
Written by: Wouter and Han
PLRV
Four different PLRV CP's were transformed into E.coli DH5α electrocompetent cells. After culturing the cells, we miniprepped the plasmids. Restriction check with XbaI and SpeI enzymes confirms the presence of the PLRV CP inside the plasmid.
Hepatitis B general
6 August
- Electro-transformation of the HepB+IPTG promoter biobrick into BL21 (expression strain)
- Adding a terminator to the HepB+IPTG promoter biobrick and electro transformation with DH5α
8 August
- colony PCR of those transformants using sequencing primers
-> no fragments where found on the gel
9 August
- the colony PCR was repeated using both sequencing primers as well as Hepatitis core protein primers in parallel
Hepatitis B outside modification
The gene of HepBcAg will be ‘cut’ in two parts using two sets of primers after which the parts will be extended towards each other by using overhanging primers. The overhang codes for linkers and the E-coil.
Legend:
- Blue: Hepatitis B core antigen gene
- Green: iGEM standard 10 prefix
- Red: iGEM standard 10 suffix
- Orange: insert containing linkers and K-coil
- Arrows: primers, with overhang.
8 August
• PCR step 1
Step 1 of the above described pcr scheme was done using primers Rev1 and FWGeneral for part 1.1 of the gene and FW1 with RevGeneral for part 1.2 of the gene. All steps were done using phusion polymerase, because of the low chance of mutations in the gene.
Result of step 1 on 1% agarose gel.
• PCR step 2
PCR steps 1.1 and 1.2 were purified using a PCR purification kit (Thermo). The primers used for this step were Rev2 and FW general on step 1.1 FW2 and Rev general on step 1.2
Result of step 2. The double band of step 2.2 is hard to explain, but the needed product is present.
9 August (Kees)
• PCR step 3
Template used was purified step 2.2 with primers: FW3 and Rev general
An overview of Steps 1.2-3.2. It is clear that the length increases as expected. The multiplicity of bands in the latter two steps could cause a problem in the ligating-pcr step 4
The nanodrop of step 3 yielded sufficient DNA to continue working (107.2 ng/µL). 10 August (Kees)
• PCR step 4
In this step, the amplicons of step 3 and 2.1 and mixed with the primers that anneal on the outside of the final construct. The overhang between the two parts should serve as a starting point for the amplification. So, for the PCR we used: 2 µL of step 2.1 (50 ng/µL) 1 µL of step 3 (100 ng/µL) With the primers: FW general Rev 3 (including a HIS-tag) This didn’t work out. Investigation of the primers showed a shift of 5aa in primer Rev 2. This error could inhibit the annealing and elongation of the overhang of the two parts. A new primer was ordered: Rev 2.2
11 August (Kees)
• PCR step 2 (with new Rev primer)
Step 2 was repeated using the new primer. The result was fully negative, including the positive control. Most like explanation is that one of the components was not added (FW primer, enzyme)
• Repeat PCR step 2
This time the step 2.2 did work.
• Step 4 attempt 2
The primers were used as on August 10, only the DNA has changed because of the different DNA resulting from step 2.
Because of repeats in the linker, and mainly, the k-coil there could be alternative annealing sites, causing a smear.
Hepatitis B inside modification
10 August
- 1st step of the 4 step reaction to fuse a coil to the C-terminal of the HepB core protein
-> 1st step seemed to work -> the gel shows a positive negative control
GFP modification
10 August
- 1st step of the 2 step reaction to fuse a coil to the N-terminal and a histag to the C-terminal of GFP
-> 1st step seemed to work
 "
"