Team:Osaka/Project
From 2012.igem.org
(→Radiation detection) |
|||
| (38 intermediate revisions not shown) | |||
| Line 4: | Line 4: | ||
== Project Details== | == Project Details== | ||
| - | <p> | + | <p> |
| - | + | It is still sharp in our memory that, on March 11, 2011, the Great East Japan Earthquake struck off the East coast of Japan's main island and triggered a series of events that led to the nation wide nuclear crisis. Moved by that accident in iGEM 2011, we have built a synthetic biological dosimeter to detect the radiation. In this year we further improved that "<b>Bio-dosimeter</b>". Our "<b>Bio-dosimeter</b>"consists of two points: <b>damage tolerance</b> and <b>radiation detection</b>. To introduce the tolerance to <i>E. coli</i>, we are trying to put in some radiation resistance genes from <i>Deinococcus radiodurans</i>. For the detection of the radiation, we are trying to connect the native DNA damage response system of <i>E. coli</i> to production of pigment lycopene as an optical reporter. Now, we are attempting to assess its tolerance to various types of DNA damage and to evaluate DNA damage detection more clearly. | |
| - | It is still sharp in our memory that, on March 11, 2011, the Great East Japan Earthquake struck off the coast of | + | |
</p> | </p> | ||
| Line 13: | Line 12: | ||
[[File:D.radiodurans.jpg|200px|right|''D.radiodurans'']] | [[File:D.radiodurans.jpg|200px|right|''D.radiodurans'']] | ||
<p> | <p> | ||
| - | The bacterium ''Deinococcus radiodurans'' shows remarkable resistance to a range of DNA damage caused by ionizing radiation, desiccation, UV radiation, oxidizing agents, and electrophilic mutagens. It is an | + | The bacterium ''Deinococcus radiodurans'' shows remarkable resistance to a range of DNA damage caused by ionizing radiation, desiccation, UV radiation, oxidizing agents, and electrophilic mutagens etc. It is an aerobic bacterium that is most famous for its extreme resistance to ionizing radiation,<i>D. radiodurans</i> is able not only to survive acute exposures to gamma radiation that exceed 15,000 Gy, but it can also grow continuously in the presence of continual exposure (60 Gy/hour) without any defectss on its growth rate or ability to express cloned genes. On the other hand, the enteric bacteria <i>E. coli</i> can withstand up to 200 Gy, where the exposure of just 5-10 Gy is lethal to a human being.</p> |
<p>We explored various genes from <i>D. radiodurans</i>, implicated in its remarkable DNA damage resistance. By BioBricking selected genes and transforming them into <i>E. coli</i>, we hoped to confer additional DNA damage tolerance to the host cells.</p> | <p>We explored various genes from <i>D. radiodurans</i>, implicated in its remarkable DNA damage resistance. By BioBricking selected genes and transforming them into <i>E. coli</i>, we hoped to confer additional DNA damage tolerance to the host cells.</p> | ||
| - | |||
==== Radiotolerance genes ==== | ==== Radiotolerance genes ==== | ||
| - | + | <i>D.radiodurans's</i> DNA repair system consist of many unique proteins. | |
[[File:DNA repair system.jpg|500px|center|''D.radiodurans'']] | [[File:DNA repair system.jpg|500px|center|''D.radiodurans'']] | ||
*'''PprI''' | *'''PprI''' | ||
| - | PprI, which is unique to ''D. radiodurans'', is | + | PprI, which is unique to ''D. radiodurans'', is regarded as the most important protein for the radiation tolerance. |
PprI can significantly and specifically induce the gene expression of recA and pprA and enhance the enzyme | PprI can significantly and specifically induce the gene expression of recA and pprA and enhance the enzyme | ||
activities of catalases. These results strongly suggest that PprI plays a crucial role in regulating multiple DNA repair and protection | activities of catalases. These results strongly suggest that PprI plays a crucial role in regulating multiple DNA repair and protection | ||
| Line 28: | Line 26: | ||
*'''PprA''' | *'''PprA''' | ||
| - | A pleiotropic protein promoting DNA repair, its role in radiation resistance of '' | + | A pleiotropic protein promoting DNA repair, its role in radiation resistance of ''D. radiodurans'' |
was demonstrated. | was demonstrated. | ||
PprA preferentially binds to double-stranded DNA carrying strand breaks, inhibits ''E. coli'' exonuclease III activity, and stimulates the DNA end-joining reaction catalysed by ATP-dependent and NAD-dependent DNA ligases. These | PprA preferentially binds to double-stranded DNA carrying strand breaks, inhibits ''E. coli'' exonuclease III activity, and stimulates the DNA end-joining reaction catalysed by ATP-dependent and NAD-dependent DNA ligases. These | ||
| Line 36: | Line 34: | ||
*'''PprM''' | *'''PprM''' | ||
PprM (a modulator of the PprI-dependent DNA damage response) is a homolog of cold shock protein (Csp). PprM regulates the induction of PprA but not that of RecA. PprM belongs in a distinct clade of a subfamily together with Csp homologs from ''D. geothermalis'' and ''Thermus thermophilus''. | PprM (a modulator of the PprI-dependent DNA damage response) is a homolog of cold shock protein (Csp). PprM regulates the induction of PprA but not that of RecA. PprM belongs in a distinct clade of a subfamily together with Csp homologs from ''D. geothermalis'' and ''Thermus thermophilus''. | ||
| - | PprM plays an important role in the induction of | + | PprM plays an important role in the induction of PprA and is involved in the unique radiation response mechanism controlled by PprI in ''D. radiodurans''. |
*'''RecA''' | *'''RecA''' | ||
| - | + | <i>D. radiodurans's</i> RecA protein has been characterized and its gene has been sequenced; its gene is homologous with <i>E. coli's</i> recA gene. <i>D. radiodurans's</i> recA mutants are highly sensitive to UV and ionizing radiation. It was repored by Carroll et al (1996) that ''E. coli'' recA did not complement an IR-sensitive <i>D. radiodurans</i> recA point-mutant (rec30) and that expression of <i>D. radiodurans</i> recA in <i>E. coli</i> was lethal. However recently, it was reported that <I>E. coli</I> recA can provide partial complementation to a <i>D. radiodurans</i> recA null mutant (Schlesinger, 2007). | |
| - | + | [[File:Effects of inoing radiation.png|350px|right|''D.radiodurans'']] | |
| - | [[File:Effects of inoing radiation.png|350px| | + | |
| - | When cells are exposed to | + | ==== The effects of ionizing radiation ==== |
| + | When cells are exposed to Ionizing radiation, it can produce reactive oxygen and break chemical bonds. To induce such kinds of DNA damages, we used chemical agents such as mitomycine C and hydrogen peroxide. | ||
*'''Mitomycin C''' | *'''Mitomycin C''' | ||
| - | The DNA-damaging agent mitomycin C is known to introduce interstrand cross-links into duplex DNA. | + | The DNA-damaging agent mitomycin C is known to introduce interstrand cross-links into duplex DNA. Mitomycin C is reduced in cells and bonds with the specific base sequence. When the bonding mitomycin C is oxidized autonomically, a reactive oxygen is produced and it breaks DNA chains near the bond site. |
| - | + | ||
| + | *'''Hydrogen peroxide''' | ||
| + | Hydrogen peroxide is known as a highly reactive compound. Therefore, when hydrogen peroxide is introduced into a cell, it breaks not only DNA molecules but also various substances in the cell, such as cell membrane, cytoplasm. This is contrasting to the fact that MitomycinC specifically breaks DNA molecules. | ||
=== Radiation detection === | === Radiation detection === | ||
==== SOS response ==== | ==== SOS response ==== | ||
| - | [[File: | + | [[File:Sos response.png|left|sos responce|350px]] |
| - | <p> | + | <p>When DNA is significantly damaged (e.g. by exposure to UV radiation or chemicals), several DNA damage-related proteins are synthesized quickly. |
| - | This reaction to DNA damage is | + | This reaction to DNA damage is known as SOS response.</p> |
| - | <p>RecA | + | <p>RecA of <i>Escherichia coli</i> is a 38 kilodalton protein essential for repair and maintenance of DNA. RecA has multiple activities, all related to DNA repair. In the bacterial SOS response, it has a co-protease function in the autocatalytic cleavage of the LexA repressor and the λ repressor. lexA gene is expressed constitutively and prevents expression of damage-related proteins by binding to a SOS box as a repressor. RecA is activated by binding to single-stranded DNA, and the activated RecA then turns on LexA protease activity. Self-cleavage of LexA derepresses the expression of damage-related proteins enabling a response to be mounted. </p> |
| - | <p>We decided to employ the promoter of the | + | <p>We decided to employ the promoter of the recA gene of <i>D. radiodurans</i> ([http://partsregistry.org/wiki/index.php?title=Part:BBa_J22106 BBa J22106]) to detect DNA damage. While RecA is an inducer of SOS genes, it itself is an SOS gene that is auto-induced upon DNA damage. Expression of genes downstream of this promoter is induced by DNA damage.</p> |
<br> | <br> | ||
==== Lycopene biosynthesis ==== | ==== Lycopene biosynthesis ==== | ||
[[File:Lycopene biosynthesis.jpg|280px|right]] | [[File:Lycopene biosynthesis.jpg|280px|right]] | ||
| - | <p>Our Bio-dosimeter | + | <p>Our Bio-dosimeter needs some visible output to alert users to radioactivity (detected as DNA damage). In a previous iGEM project, "colrcoli", we attempted to use <I>E.coli</I> as a paint tool. To that end, we examined biosynthesis of carotenoid pigments as a way of producing color. Here, we attempted to use biosynthesis of the carotenoid lycopene as a reporter for DNA damage.</p> |
| - | <p>Carotenoid is a family of natural pigments. Many plants such as fruits and vegetables contain these pigments. For example, tomato has lycopene(red), carrot has carotene(orange) | + | <p>Carotenoid is a family of natural pigments. Many plants such as fruits and vegetables contain these pigments. For example, tomato has lycopene (red), carrot has carotene (orange), and xanthophyll (yellow) is found in almost all plants.</p> |
| - | <p>Biosynthesis of carotenoid pigments starts from FPP(FARNESYL DIPHOSPHATE). FPP is formed from isopentenylpyrophosphate(IPP) and dimethylallylpyrophosphate(DMAPP), which can be biologically produced by two distinct pathways, the mevalonate and non-mevalonate pathways. In <i>E.coli</i>, FPP is formed through the non-mevalonate pathway. By the introduction of heterologous enzymatic genes colorless FPP is then converted to orange-red lycopene, which has a peak absorbance at 407nm that is easily measured. </p> | + | <p>Biosynthesis of carotenoid pigments starts from FPP (FARNESYL DIPHOSPHATE). FPP is formed from isopentenylpyrophosphate (IPP) and dimethylallylpyrophosphate (DMAPP), which can be biologically produced by two distinct pathways, the mevalonate and non-mevalonate pathways. In <i>E.coli</i>, FPP is formed through the non-mevalonate pathway. By the introduction of heterologous enzymatic genes colorless FPP is then converted to orange-red lycopene, which has a peak absorbance at 407nm that is easily measured. </p> |
<p>As shown in the figure on the right, the heterologous enzymes CrtE, CrtB, CrtI were introduced into <i>E. coli</i> to complete the biosynthesis of lycopene. We used a lycopene biosynthetic gene cluster provided by 2009 Cambridge (<partinfo>K274100</partinfo>).<br> | <p>As shown in the figure on the right, the heterologous enzymes CrtE, CrtB, CrtI were introduced into <i>E. coli</i> to complete the biosynthesis of lycopene. We used a lycopene biosynthetic gene cluster provided by 2009 Cambridge (<partinfo>K274100</partinfo>).<br> | ||
<br> | <br> | ||
| - | |||
=== References=== | === References=== | ||
| Line 77: | Line 75: | ||
<li>H. Lu et al., “''Deinococcus radiodurans'' PprI switches on DNA damage response and cellular survival networks after radiation damage,” Molecular & Cellular Proteomics: MCP, vol. 8, no. 3, pp. 481-494, Mar. 2009.</li> | <li>H. Lu et al., “''Deinococcus radiodurans'' PprI switches on DNA damage response and cellular survival networks after radiation damage,” Molecular & Cellular Proteomics: MCP, vol. 8, no. 3, pp. 481-494, Mar. 2009.</li> | ||
<li>H. Ohba, K. Satoh, H. Sghaier, T. Yanagisawa, and I. Narumi, “Identification of PprM: a modulator of the PprI-dependent DNA damage response in ''Deinococcus radiodurans'',” Extremophiles: Life Under Extreme Conditions, vol. 13, no. 3, pp. 471-479, May 2009.</li> | <li>H. Ohba, K. Satoh, H. Sghaier, T. Yanagisawa, and I. Narumi, “Identification of PprM: a modulator of the PprI-dependent DNA damage response in ''Deinococcus radiodurans'',” Extremophiles: Life Under Extreme Conditions, vol. 13, no. 3, pp. 471-479, May 2009.</li> | ||
| + | <li>Dusre L, Covey JM, Collins C, and Sinha BK, "DNA damage, cytotoxicity and free radical formation by mitomycin C in human cells." Chem Biol Interact. 1989;71(1):63-78 </li> | ||
| + | <li>Switala J, Loewen PC, "Diversity of properties among catalases." Arch Biochem Biophys. 2002 May 15;401(2):145-54. </li> | ||
| + | <li>K. Ueda, J. Morita, and T. Komano, "Phage inactivation and DNA scission activities of 7-N-(p-Hydroxyphenyl)Mitomycin C" The journal of antibiotics, Oct. 1982. | ||
| + | </li> | ||
</ol> | </ol> | ||
Latest revision as of 14:26, 21 October 2012
Project Details
It is still sharp in our memory that, on March 11, 2011, the Great East Japan Earthquake struck off the East coast of Japan's main island and triggered a series of events that led to the nation wide nuclear crisis. Moved by that accident in iGEM 2011, we have built a synthetic biological dosimeter to detect the radiation. In this year we further improved that "Bio-dosimeter". Our "Bio-dosimeter"consists of two points: damage tolerance and radiation detection. To introduce the tolerance to E. coli, we are trying to put in some radiation resistance genes from Deinococcus radiodurans. For the detection of the radiation, we are trying to connect the native DNA damage response system of E. coli to production of pigment lycopene as an optical reporter. Now, we are attempting to assess its tolerance to various types of DNA damage and to evaluate DNA damage detection more clearly.
Damage tolerance
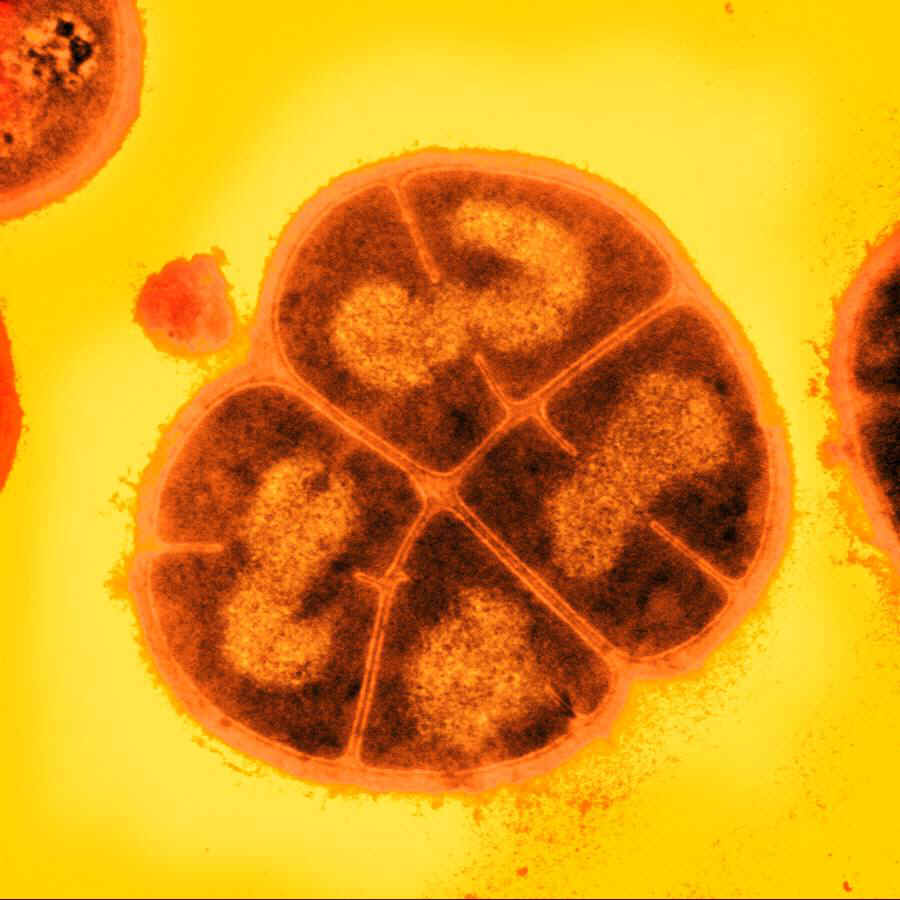
D. radiodurans
The bacterium Deinococcus radiodurans shows remarkable resistance to a range of DNA damage caused by ionizing radiation, desiccation, UV radiation, oxidizing agents, and electrophilic mutagens etc. It is an aerobic bacterium that is most famous for its extreme resistance to ionizing radiation,D. radiodurans is able not only to survive acute exposures to gamma radiation that exceed 15,000 Gy, but it can also grow continuously in the presence of continual exposure (60 Gy/hour) without any defectss on its growth rate or ability to express cloned genes. On the other hand, the enteric bacteria E. coli can withstand up to 200 Gy, where the exposure of just 5-10 Gy is lethal to a human being.
We explored various genes from D. radiodurans, implicated in its remarkable DNA damage resistance. By BioBricking selected genes and transforming them into E. coli, we hoped to confer additional DNA damage tolerance to the host cells.
Radiotolerance genes
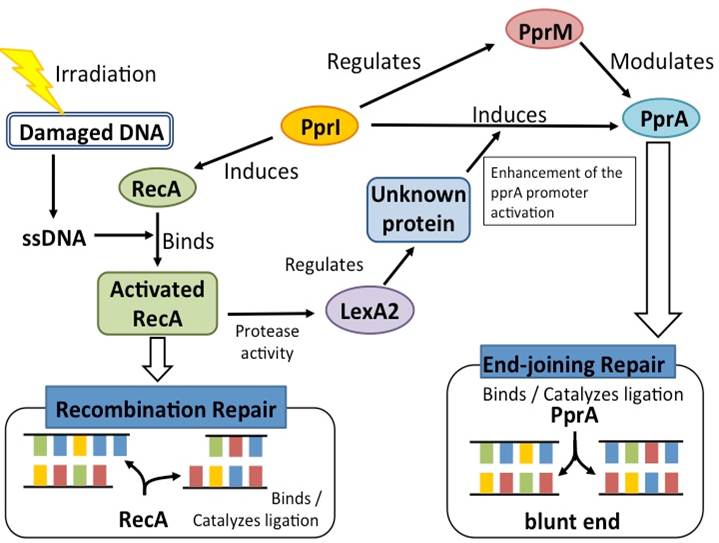
D.radiodurans's DNA repair system consist of many unique proteins.
- PprI
PprI, which is unique to D. radiodurans, is regarded as the most important protein for the radiation tolerance. PprI can significantly and specifically induce the gene expression of recA and pprA and enhance the enzyme activities of catalases. These results strongly suggest that PprI plays a crucial role in regulating multiple DNA repair and protection pathways in response to radiation stress.
- PprA
A pleiotropic protein promoting DNA repair, its role in radiation resistance of D. radiodurans was demonstrated. PprA preferentially binds to double-stranded DNA carrying strand breaks, inhibits E. coli exonuclease III activity, and stimulates the DNA end-joining reaction catalysed by ATP-dependent and NAD-dependent DNA ligases. These results suggest that D. radiodurans has a radiationinduced non-homologous end-joining repair mechanism in which PprA plays a critical role.
- PprM
PprM (a modulator of the PprI-dependent DNA damage response) is a homolog of cold shock protein (Csp). PprM regulates the induction of PprA but not that of RecA. PprM belongs in a distinct clade of a subfamily together with Csp homologs from D. geothermalis and Thermus thermophilus. PprM plays an important role in the induction of PprA and is involved in the unique radiation response mechanism controlled by PprI in D. radiodurans.
- RecA
D. radiodurans's RecA protein has been characterized and its gene has been sequenced; its gene is homologous with E. coli's recA gene. D. radiodurans's recA mutants are highly sensitive to UV and ionizing radiation. It was repored by Carroll et al (1996) that E. coli recA did not complement an IR-sensitive D. radiodurans recA point-mutant (rec30) and that expression of D. radiodurans recA in E. coli was lethal. However recently, it was reported that E. coli recA can provide partial complementation to a D. radiodurans recA null mutant (Schlesinger, 2007).
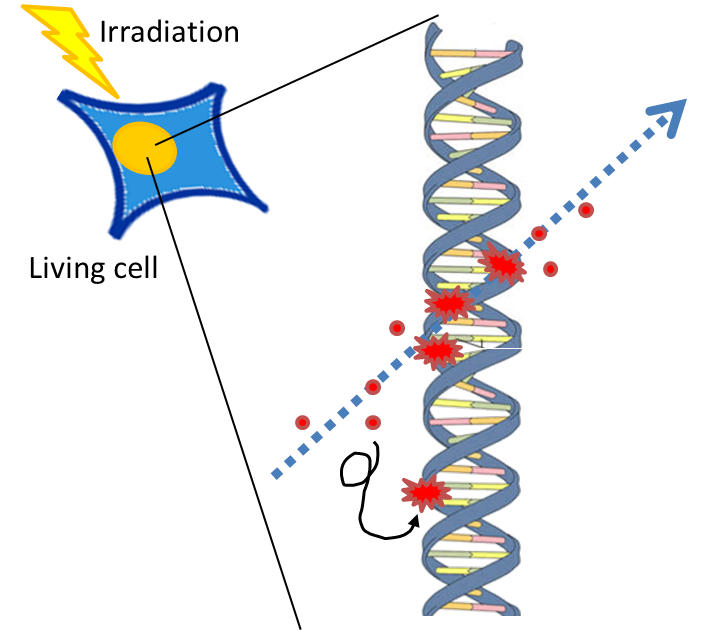
The effects of ionizing radiation
When cells are exposed to Ionizing radiation, it can produce reactive oxygen and break chemical bonds. To induce such kinds of DNA damages, we used chemical agents such as mitomycine C and hydrogen peroxide.
- Mitomycin C
The DNA-damaging agent mitomycin C is known to introduce interstrand cross-links into duplex DNA. Mitomycin C is reduced in cells and bonds with the specific base sequence. When the bonding mitomycin C is oxidized autonomically, a reactive oxygen is produced and it breaks DNA chains near the bond site.
- Hydrogen peroxide
Hydrogen peroxide is known as a highly reactive compound. Therefore, when hydrogen peroxide is introduced into a cell, it breaks not only DNA molecules but also various substances in the cell, such as cell membrane, cytoplasm. This is contrasting to the fact that MitomycinC specifically breaks DNA molecules.
Radiation detection
SOS response
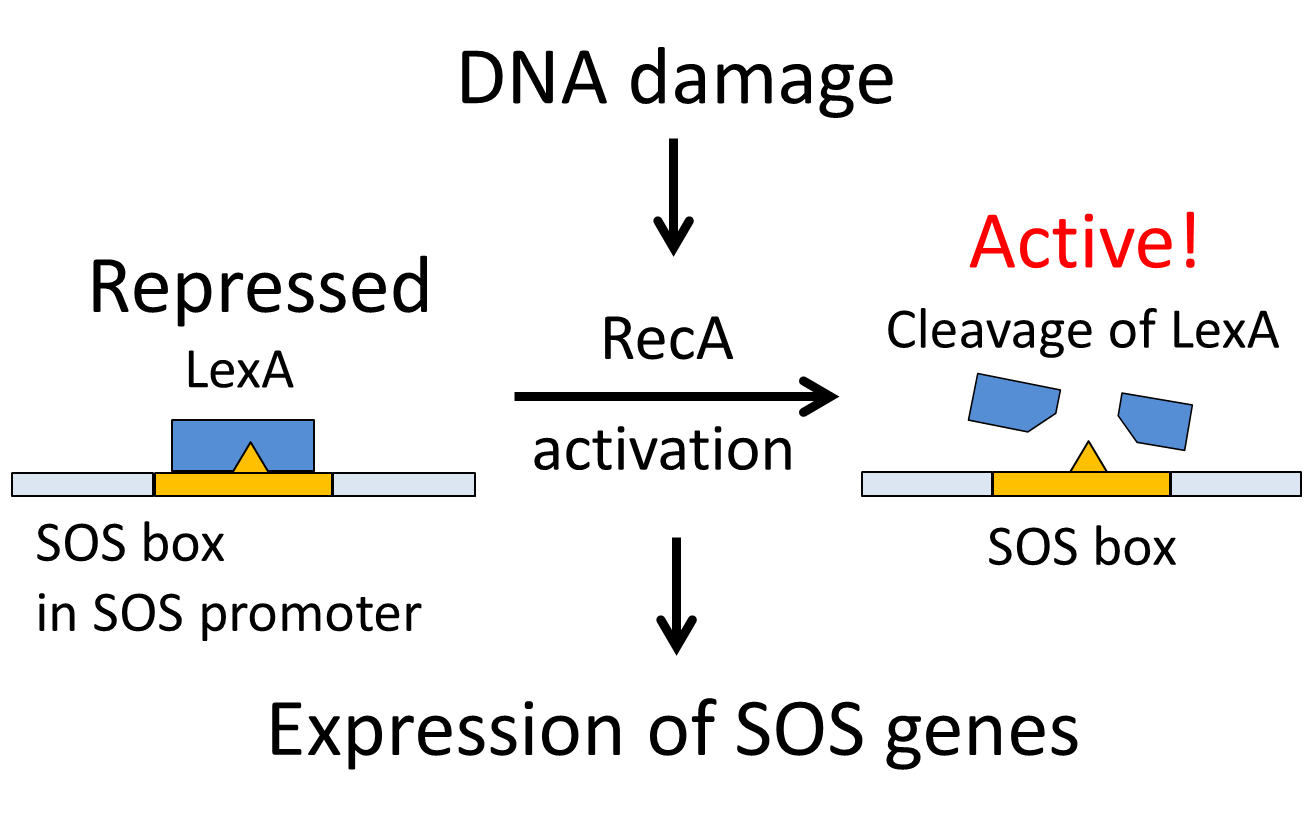
When DNA is significantly damaged (e.g. by exposure to UV radiation or chemicals), several DNA damage-related proteins are synthesized quickly. This reaction to DNA damage is known as SOS response.
RecA of Escherichia coli is a 38 kilodalton protein essential for repair and maintenance of DNA. RecA has multiple activities, all related to DNA repair. In the bacterial SOS response, it has a co-protease function in the autocatalytic cleavage of the LexA repressor and the λ repressor. lexA gene is expressed constitutively and prevents expression of damage-related proteins by binding to a SOS box as a repressor. RecA is activated by binding to single-stranded DNA, and the activated RecA then turns on LexA protease activity. Self-cleavage of LexA derepresses the expression of damage-related proteins enabling a response to be mounted.
We decided to employ the promoter of the recA gene of D. radiodurans ([http://partsregistry.org/wiki/index.php?title=Part:BBa_J22106 BBa J22106]) to detect DNA damage. While RecA is an inducer of SOS genes, it itself is an SOS gene that is auto-induced upon DNA damage. Expression of genes downstream of this promoter is induced by DNA damage.
Lycopene biosynthesis
Our Bio-dosimeter needs some visible output to alert users to radioactivity (detected as DNA damage). In a previous iGEM project, "colrcoli", we attempted to use E.coli as a paint tool. To that end, we examined biosynthesis of carotenoid pigments as a way of producing color. Here, we attempted to use biosynthesis of the carotenoid lycopene as a reporter for DNA damage.
Carotenoid is a family of natural pigments. Many plants such as fruits and vegetables contain these pigments. For example, tomato has lycopene (red), carrot has carotene (orange), and xanthophyll (yellow) is found in almost all plants.
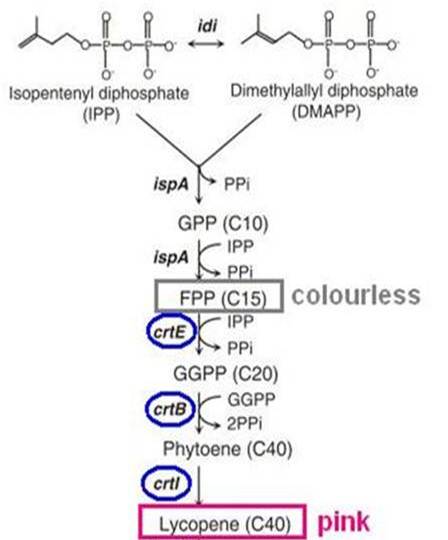
Biosynthesis of carotenoid pigments starts from FPP (FARNESYL DIPHOSPHATE). FPP is formed from isopentenylpyrophosphate (IPP) and dimethylallylpyrophosphate (DMAPP), which can be biologically produced by two distinct pathways, the mevalonate and non-mevalonate pathways. In E.coli, FPP is formed through the non-mevalonate pathway. By the introduction of heterologous enzymatic genes colorless FPP is then converted to orange-red lycopene, which has a peak absorbance at 407nm that is easily measured.
As shown in the figure on the right, the heterologous enzymes CrtE, CrtB, CrtI were introduced into E. coli to complete the biosynthesis of lycopene. We used a lycopene biosynthetic gene cluster provided by 2009 Cambridge (<partinfo>K274100</partinfo>).
References
- [1] Y. Hua et al., “PprI: a general switch responsible for extreme radioresistance of Deinococcus radiodurans,” Biochemical and Biophysical Research Communications, vol. 306, no. 2, pp. 354-360, Jun. 2003.
- G. Gao, B. Tian, L. Liu, D. Sheng, B. Shen, and Y. Hua, “Expression of Deinococcus radiodurans PprI enhances the radioresistance of Escherichia coli,” DNA Repair, vol. 2, no. 12, pp. 1419-1427, Dec. 2003.
- I. Narumi, K. Satoh, S. Cui, T. Funayama, S. Kitayama, and H. Watanabe, “PprA: a novel protein from Deinococcus radiodurans that stimulates DNA ligation,” Molecular Microbiology, vol. 54, no. 1, pp. 278-285, Oct. 2004.
- S. Kota and H. S. Misra, “PprA: A protein implicated in radioresistance of Deinococcus radiodurans stimulates catalase activity in Escherichia coli,” Applied Microbiology and Biotechnology, vol. 72, no. 4, pp. 790-796, Oct. 2006.
- H. Lu et al., “Deinococcus radiodurans PprI switches on DNA damage response and cellular survival networks after radiation damage,” Molecular & Cellular Proteomics: MCP, vol. 8, no. 3, pp. 481-494, Mar. 2009.
- H. Ohba, K. Satoh, H. Sghaier, T. Yanagisawa, and I. Narumi, “Identification of PprM: a modulator of the PprI-dependent DNA damage response in Deinococcus radiodurans,” Extremophiles: Life Under Extreme Conditions, vol. 13, no. 3, pp. 471-479, May 2009.
- Dusre L, Covey JM, Collins C, and Sinha BK, "DNA damage, cytotoxicity and free radical formation by mitomycin C in human cells." Chem Biol Interact. 1989;71(1):63-78
- Switala J, Loewen PC, "Diversity of properties among catalases." Arch Biochem Biophys. 2002 May 15;401(2):145-54.
- K. Ueda, J. Morita, and T. Komano, "Phage inactivation and DNA scission activities of 7-N-(p-Hydroxyphenyl)Mitomycin C" The journal of antibiotics, Oct. 1982.
 "
"