Team:Wageningen UR/Journal/week20
From 2012.igem.org
Jeroenbosman (Talk | contribs) (→week 20: 10 September - 16 September) |
(→CCMV) |
||
| (15 intermediate revisions not shown) | |||
| Line 2: | Line 2: | ||
= week 20: 10 September - 16 September = | = week 20: 10 September - 16 September = | ||
| + | ==== CCMV ==== | ||
| + | <br/><br/> | ||
| + | ''10th September'' (Hugo) | ||
| + | <br/> | ||
| + | Colony PCR on transformant colonies of 07-09. | ||
| + | <br/> | ||
| + | Inoculated 10 mL of LB with cells from an IPTG_CCMV_NEG colony overnight. | ||
| + | <br/> | ||
| + | Inoculated 10 mL of LB with cells from an IPTG_CCMV_HIS colony overnight. | ||
| + | <br/><br/> | ||
| + | ''11th September'' (Hugo) | ||
| + | <br/> | ||
| + | Miniprepped the two cultures that grew overnight.<br/> | ||
| + | Colony PCR on additional colonies from the CCMV_COIL into pSB1C3 (RFP backbone) plate. | ||
| + | <br/> | ||
| + | Inoculated 10 mL of LB with cells from four CCMV_COIL colonies overnight. | ||
| + | <br/> | ||
| + | ''12th September'' (Hugo) | ||
| + | Miniprepped the four cultures that grew overnight. | ||
| + | <br/> | ||
| + | Digestion check on the 6 minipreps, followed by a digestion check using EcoRI + PstI (figure 1).<br/> | ||
| + | Lane 1: Marker<br/> | ||
| + | Lane 2: CCMV_COIL #4<br/> | ||
| + | Lane 3: CCMV_COIL #12<br/> | ||
| + | Lane 4: CCMV_COIL #25<br/> | ||
| + | Lane 5: CCMV_COIL #28<br/> | ||
| + | Lane 6: CCMV_HIS <br/> | ||
| + | Lane 7: IPTG_CCMV_NEG<br/> | ||
| + | Lane 8: Marker<br/> | ||
| + | <br/> | ||
| + | [[File:Digestion_check_12_09.PNG|400px|left|thumb|''Figure 1. Digestion check of plasmids before they were sent for sequencing.'']] | ||
| + | <br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/> | ||
| + | Sent all 6 miniprepped plasmids for sequencing (BaseClear). | ||
| + | <br/><br/> | ||
| + | ''13th September'' (Hugo) | ||
| + | <br/> | ||
| + | Kees did colony PCR on additional colonies from the CCMV_Delta26_HIS into pSB1C3 (RFP backbone). | ||
| + | <br/> | ||
| + | |||
| + | <br/><br/> | ||
| + | ''13th and 14th September'' (Robert) | ||
| + | <br><br> | ||
| + | Started PCR for the CCMV wt K-coil construct with the primers: | ||
| + | * primer 1 FW1 FL-CCMV wt (5' TGGTGGTGGTTCTGGTGGTGGTGGTTCTGCTGCTGCTCCATGGGTACAGTCGGAAC 3') | ||
| + | * primer 2 BW CCMV suffix | ||
| + | We took two different concentrations (30ng and 3ng) of the plasmid with the CCMV wt construct to avoid impurities. | ||
| + | <br><br> | ||
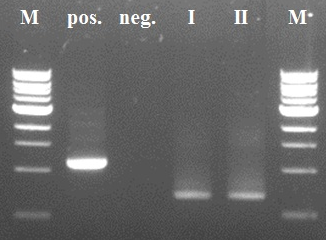
| + | [[File:CCMV wt K-coil PCR step1.jpg|400px|left|thumb|''Figure 2: PCR step I CCMV wt + K-coil'']][[File:PCR step II CCMV wt K-coil.jpg|400px|left|thumb|''Figure 3: PCR step II CCMV wt + K-coil'']] | ||
| + | <br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br> | ||
| + | Positive control of the first PCR step was negative due to a pipetting mistake (too less DNTP's). We took the PCR product of sample I for the second PCR step (after a PCR purification with a ThermoScientific kit), because sample II showed an impurity. In the second step the positive control worked properly as intented. We decided not try the third and fourth step, because of a mistake in primer design earlier. | ||
| + | <br><br> | ||
'''TuYV''' | '''TuYV''' | ||
| Line 35: | Line 86: | ||
''12 September'' | ''12 September'' | ||
| - | * Miniprep and digestion check of colonies 5.Sept. (Hephis tag in | + | * Miniprep and digestion check of colonies 5.Sept. (Hephis tag in pSB1C3) |
-> Restriction check shows inserts that might be too large (expected size: around 600bp) | -> Restriction check shows inserts that might be too large (expected size: around 600bp) | ||
(see picture ''restriction check 12 September'') | (see picture ''restriction check 12 September'') | ||
| - | * Sent samples for sequencing (HepB core protein + his tag in | + | * Sent samples for sequencing (HepB core protein + his tag in pSB1C3) |
| Line 106: | Line 157: | ||
''12 September'' | ''12 September'' | ||
| - | * Miniprep and digestion check of colonies 5.Sept. (Hepinsidecoil in | + | * Miniprep and digestion check of colonies 5.Sept. (Hepinsidecoil in pSB1C3) |
-> Restriction check shows inserts that might be too large (expected size: around 600bp) | -> Restriction check shows inserts that might be too large (expected size: around 600bp) | ||
| Line 113: | Line 164: | ||
| - | * Sent samples for sequencing (HepB core protein + insidecoil in | + | * Sent samples for sequencing (HepB core protein + insidecoil in pSB1C3) |
| Line 186: | Line 237: | ||
[[File: GFP-coil fusion PCR step 2.png|center|500px|thumb|<p align="justify">''Figure 5: step 2 of the PCR reaction with 2 different primer sets and different template concentrations (primer set 1: forward primer GFP-coil 2nd step; reverse primer GFP primer adding a his tag – primer set 2: forward primer GFP-coil 2nd step; reverse primer sequencing primer (no his tag)) ''</p>]] | [[File: GFP-coil fusion PCR step 2.png|center|500px|thumb|<p align="justify">''Figure 5: step 2 of the PCR reaction with 2 different primer sets and different template concentrations (primer set 1: forward primer GFP-coil 2nd step; reverse primer GFP primer adding a his tag – primer set 2: forward primer GFP-coil 2nd step; reverse primer sequencing primer (no his tag)) ''</p>]] | ||
| - | -> the 2nd PCR reaction worked well with both template concentrations - the experiment will be continued with the samples containing less template | + | -> the 2nd PCR reaction worked well with both template concentrations - the experiment will be continued with the samples containing less template |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
---- | ---- | ||
[[https://2012.igem.org/Team:Wageningen_UR/Journal/week19 previous week]] [[https://2012.igem.org/Team:Wageningen_UR/Journal/week21 next week]] | [[https://2012.igem.org/Team:Wageningen_UR/Journal/week19 previous week]] [[https://2012.igem.org/Team:Wageningen_UR/Journal/week21 next week]] | ||
Latest revision as of 19:56, 26 September 2012
week 20: 10 September - 16 September
CCMV
10th September (Hugo)
Colony PCR on transformant colonies of 07-09.
Inoculated 10 mL of LB with cells from an IPTG_CCMV_NEG colony overnight.
Inoculated 10 mL of LB with cells from an IPTG_CCMV_HIS colony overnight.
11th September (Hugo)
Miniprepped the two cultures that grew overnight.
Colony PCR on additional colonies from the CCMV_COIL into pSB1C3 (RFP backbone) plate.
Inoculated 10 mL of LB with cells from four CCMV_COIL colonies overnight.
12th September (Hugo)
Miniprepped the four cultures that grew overnight.
Digestion check on the 6 minipreps, followed by a digestion check using EcoRI + PstI (figure 1).
Lane 1: Marker
Lane 2: CCMV_COIL #4
Lane 3: CCMV_COIL #12
Lane 4: CCMV_COIL #25
Lane 5: CCMV_COIL #28
Lane 6: CCMV_HIS
Lane 7: IPTG_CCMV_NEG
Lane 8: Marker
Sent all 6 miniprepped plasmids for sequencing (BaseClear).
13th September (Hugo)
Kees did colony PCR on additional colonies from the CCMV_Delta26_HIS into pSB1C3 (RFP backbone).
13th and 14th September (Robert)
Started PCR for the CCMV wt K-coil construct with the primers:
- primer 1 FW1 FL-CCMV wt (5' TGGTGGTGGTTCTGGTGGTGGTGGTTCTGCTGCTGCTCCATGGGTACAGTCGGAAC 3')
- primer 2 BW CCMV suffix
We took two different concentrations (30ng and 3ng) of the plasmid with the CCMV wt construct to avoid impurities.
Positive control of the first PCR step was negative due to a pipetting mistake (too less DNTP's). We took the PCR product of sample I for the second PCR step (after a PCR purification with a ThermoScientific kit), because sample II showed an impurity. In the second step the positive control worked properly as intented. We decided not try the third and fourth step, because of a mistake in primer design earlier.
TuYV
Another two colonies PCR with pJET sequencing primers were done to find out colonise containing TuYV coat protein with readthrough and coat protein with readthrough with his-tag. Finally another colony containing either the coat protein and readthrough or coat protein and readthrough with his-tag was identified.
All the colonies for colony PCR were minipreped.
TuYV constructs (PCR products) were digested, ligated into backbone from BBa_J04450 and transformed into MachI strain. In parallel, TuYV constructs (PCR products) were digested, ligated with vector containing the IPTG induced promoter again, and transformed into MachI.
The next day we saw very promising colonies and the colony PCR confirmed that we successfully put TuYV constructs into the chloramphenical backbone, but for adding them at the downstream of IPTG induced promoter, colony PCR revealed that there were only the backbones containing the IPTG induced promoter in the E.coli.
Polero
Cells containing the plasmid( IPTG induced promoter with polero coat protein) were induced and lysised by French Press.
Dialysis and purification were done with Polero VLPs buffer. Samples were ready to send for EM test.
Hepatitis B general
10 September
- 2nd try colony PCR of the transformations from 4. September (with sequencing primers)
-> 6 out of 9 samples show an insert of the right size (see picture Colony PCR 12 September)
12 September
- Miniprep and digestion check of colonies 5.Sept. (Hephis tag in pSB1C3)
-> Restriction check shows inserts that might be too large (expected size: around 600bp) (see picture restriction check 12 September)
- Sent samples for sequencing (HepB core protein + his tag in pSB1C3)
13 September
- digestion and ligation of
- HepB core protein behind an IPTG promoter in pSB1C3 backbone
- transformaion with DH5α
Hepatitis B outside modification
- 10 September (Kees)
• Colony PCR
The colonies were picked after which part of it was put in pcr-tubes for checking the content and part pas plated for continuation of the colony.
- 11 September (Kees)
• Reverse PCR coil modification Same was done as on 7 September, with the following modifications: - 25 pcr cycles instead of 15. This will yield more template, taking the risk of modifications. - Analyse pcr result on a agarose gel before further processing. - Ligate overnight at 16 degrees if gel shows linear band at about 3kb. (protocols: http://www.neb.com/nebecomm/products_intl/faqproductM0202.asp, http://www.idtdna.com/pages/docs/user-guides-and-protocols/mutagenesis-application-guide.pdf, p.32) - Digest with DpnI after ligation
- 12 September (Kees)
• Digestion and plating After digestion of the ligated sample and the non-ligated sample, the cells were transformed: - Template (positive control) - Ligated + digested - Digested only
- 13 September (Kees)
One colony grew on the DpnI+Ligated plate. This colony will be checked with colony pcr and inocculated to produce anything that is in there.
Hepatitis B inside modification
10 September
- 2nd try colony PCR of the transformations from 4. September (with sequencing primers)
-> one colony seems to have the correct insert in the pSB1C3 backbone -> no conclusion can be made about samples containing the Hep-inside coil fragment in the Bba_J04500 brick (see picture Colony PCR 12 September)
12 September
- Miniprep and digestion check of colonies 5.Sept. (Hepinsidecoil in pSB1C3)
-> Restriction check shows inserts that might be too large (expected size: around 600bp)
- Sent samples for sequencing (HepB core protein + insidecoil in pSB1C3)
13 September
- digestion and ligation of
- HepB - inside coil PCR fragment in Bba_J04500 - HepB -inside coil in pSB1C3 backbone into Bba_J04500
- transformaion with DH5α
GFP modification
10 September
- colony PCR of the transformations from 6. September (PCR step 1 in pJET) - using pJET sequencing primers
-> no band on the gel
11 September
- step 1 of the PCR reaction
- With different template concentrations (from 20ng until 0.2ng added to the mix) - With two different primer sets (both with the forward primer adding part of the coil + once with the GFP reverse primer adding a histag at the C-terminal and once with the sequencing reverse primer)
-> bands of the expected size visible (even the samples that contain a very low ammount of template). There is also a second band visible (decreasing in strenght with decreasing template concentration) -> something went wrong with the second set of reactions (the reactions where made in duplo)
-> this shows that the experiment can be continued with the sample containing only 0.2ng template DNA (so the risk of transforming with the original template is lowered)
- Of the GFP-coil 1st step (both primer sets) in pJET (samples with 0.2ng template in the PCR mixture)
- Transformation With DH5α using different amount of ligation mixture for the electro transformation (1, 3 and 5 µL)
-> more colonies where growing on the plates containing transformants transformed with 5µL
12 September
- Colony PCR of the transformations (GFP-coil step1 in pJET in DH5α)
-> the transformation was successful - there are colonies present that contain an insert with the expected size
13 September
- step 2 of the PCR reaction (continued with 2 different reverse primers) using a colony from the plate (tranformtation GFP-coil step 1 in pJET with DH5α) as template
-> the PCR reaction did not suceed - there where no bands visible on the gel
14 September
- miniprep of the transformants from 11 September
- step 2 of the PCR reaction using different dilutions of the miniprep as template (2 and 0.2 ng in the mix)
-> the 2nd PCR reaction worked well with both template concentrations - the experiment will be continued with the samples containing less template
 "
"