Team:TU-Eindhoven/LEC/LabTheory
From 2012.igem.org
| Line 14: | Line 14: | ||
<h3>Chassis</h3> | <h3>Chassis</h3> | ||
<p>Before this light emitting cell display project could start, it was necessary to decide on a suitable chassis. Candidates were E. coli and S. cerevisiae. Both are common model organisms that can be used for protein expression and are cheap to culture. To reach the goal of this project over expression of voltage-gated calcium channel was needed. As soon as it was found that CCH1-MID1, homologous to mammalian voltage-gated calcium channels, could be over expressed in S. cerevisiae, it was decided to use yeast as the chassis in our project. In the lab we had a strain called INVSc1 available which was compatiable with the plasmids we were planning to use.</p> | <p>Before this light emitting cell display project could start, it was necessary to decide on a suitable chassis. Candidates were E. coli and S. cerevisiae. Both are common model organisms that can be used for protein expression and are cheap to culture. To reach the goal of this project over expression of voltage-gated calcium channel was needed. As soon as it was found that CCH1-MID1, homologous to mammalian voltage-gated calcium channels, could be over expressed in S. cerevisiae, it was decided to use yeast as the chassis in our project. In the lab we had a strain called INVSc1 available which was compatiable with the plasmids we were planning to use.</p> | ||
| - | + | <br /> | |
| - | < | + | <h3>Plasmids and transformations</h3> |
<p>Since we choose yeast as our chassis we need to clone all required genes into shuttle vectors. A shuttle vector is a special type of vector that can be propagated both in yeast and in bacteria. Cloning can be done in fast growing E. coli while proteins can be expressed in yeast. There are low copy-number and high copy-number variants of shuttle vectors, the choice of which depends on the required amount of that protein in the cell. In this project we used three different shuttle vectors, each carrying one gene to be over expressed.</p> | <p>Since we choose yeast as our chassis we need to clone all required genes into shuttle vectors. A shuttle vector is a special type of vector that can be propagated both in yeast and in bacteria. Cloning can be done in fast growing E. coli while proteins can be expressed in yeast. There are low copy-number and high copy-number variants of shuttle vectors, the choice of which depends on the required amount of that protein in the cell. In this project we used three different shuttle vectors, each carrying one gene to be over expressed.</p> | ||
| Line 28: | Line 28: | ||
<p>Selection of successful transformants is done by auxotrophic markers, one for each plasmid. The yeast strain we use for transformations is deficient in the synthesis of certain amino acids. These usually have to be added to the medium for the yeast to survive. The uptake of a plasmid however, will restore the ability of the cell to synthesize the missing amino acid and therefore survive in the medium.</p> | <p>Selection of successful transformants is done by auxotrophic markers, one for each plasmid. The yeast strain we use for transformations is deficient in the synthesis of certain amino acids. These usually have to be added to the medium for the yeast to survive. The uptake of a plasmid however, will restore the ability of the cell to synthesize the missing amino acid and therefore survive in the medium.</p> | ||
| - | + | <br /> | |
<h3>Compatibility of yeast and device</h3> | <h3>Compatibility of yeast and device</h3> | ||
<p>The genetically modified S. cerevisiae cells will be put to the test in our home-made [[Team:TU-Eindhoven/LEC/Device|device]], designed for providing electrical stimuli to the yeast cells. These tests are done in a single bath filled with SC medium containing nutrients for our genetically modified yeast and additional free Ca<sup>2+</sup> ions. After electrical stimulation of yeast at different positions in the device, optical signals will be expected to be visible to the naked eye. In a more sensitive analysis, isolated protein will be characterized on a spectrophotometer.</p> | <p>The genetically modified S. cerevisiae cells will be put to the test in our home-made [[Team:TU-Eindhoven/LEC/Device|device]], designed for providing electrical stimuli to the yeast cells. These tests are done in a single bath filled with SC medium containing nutrients for our genetically modified yeast and additional free Ca<sup>2+</sup> ions. After electrical stimulation of yeast at different positions in the device, optical signals will be expected to be visible to the naked eye. In a more sensitive analysis, isolated protein will be characterized on a spectrophotometer.</p> | ||
| - | + | <br /> | |
<h3>GECOs</h3> | <h3>GECOs</h3> | ||
| Line 40: | Line 40: | ||
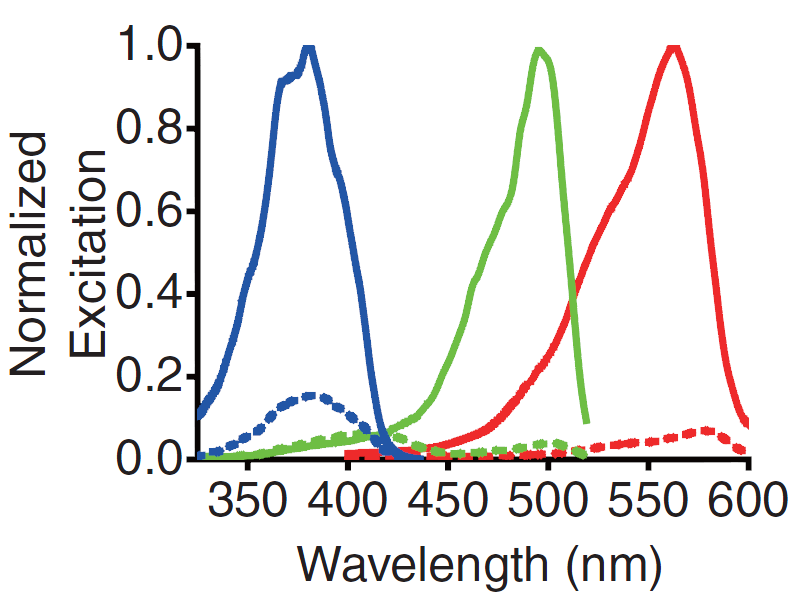
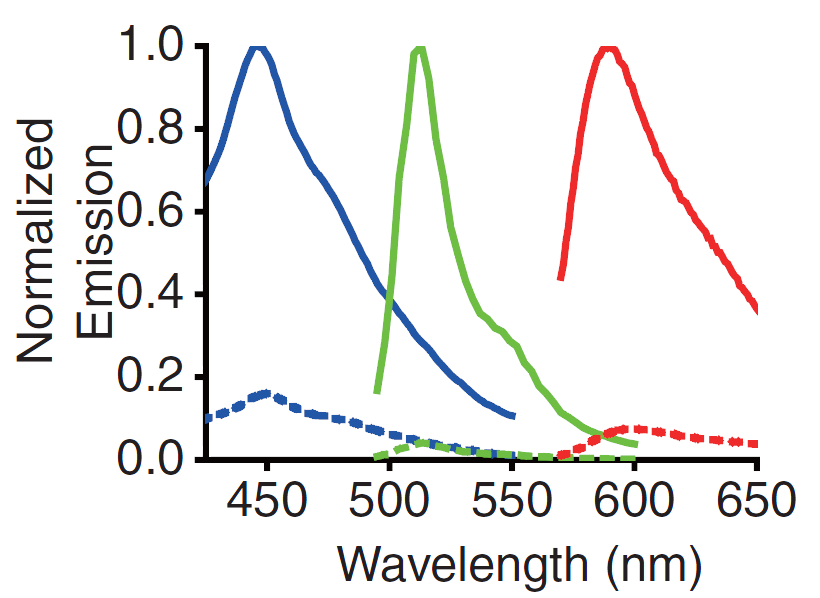
<p>A GECO is a protein which emits light in the presence of Ca<sup>2+</sup><html><a href="#ref_zhao" name="text_zhao"><sup>[2]</sup></a></html>. There are two important classes of genetically encoded Ca<sup>2+</sup> indicators. One is called the Forster Resonance Energy Transfer (FRET)-based cameleon type<html><a href="#ref_miyawaki" name="text_miyawaki"><sup>[4]</sup></a></html> and the other one is called the single Green Fluorescent Protein (GFP) type<html><a href="#ref_nakai" name="text_nakai"><sup>[5]</sup></a></html>. The GECO protein belongs to the single GFP type. Research has shown that Ca<sup>2+</sup> indicators targeted to the E. coli periplasm can be shifted toward the Ca<sup>2+</sup>-free or Ca<sup>2+</sup> -bound states by manipulation of the environmental Ca<sup>2+</sup> concentration<html><a href="#ref_zhao" name="text_zhao"><sup>[2]</sup></a></html>. Robert E. Campbell et al. named these Ca<sup>2+</sup> indicators GECOs. R-GECO, G-GECO and B-GECO emit red, green or blue light respectively, each with another excitation and emission spectrum (Fig. 2 and Fig. 3).</p> | <p>A GECO is a protein which emits light in the presence of Ca<sup>2+</sup><html><a href="#ref_zhao" name="text_zhao"><sup>[2]</sup></a></html>. There are two important classes of genetically encoded Ca<sup>2+</sup> indicators. One is called the Forster Resonance Energy Transfer (FRET)-based cameleon type<html><a href="#ref_miyawaki" name="text_miyawaki"><sup>[4]</sup></a></html> and the other one is called the single Green Fluorescent Protein (GFP) type<html><a href="#ref_nakai" name="text_nakai"><sup>[5]</sup></a></html>. The GECO protein belongs to the single GFP type. Research has shown that Ca<sup>2+</sup> indicators targeted to the E. coli periplasm can be shifted toward the Ca<sup>2+</sup>-free or Ca<sup>2+</sup> -bound states by manipulation of the environmental Ca<sup>2+</sup> concentration<html><a href="#ref_zhao" name="text_zhao"><sup>[2]</sup></a></html>. Robert E. Campbell et al. named these Ca<sup>2+</sup> indicators GECOs. R-GECO, G-GECO and B-GECO emit red, green or blue light respectively, each with another excitation and emission spectrum (Fig. 2 and Fig. 3).</p> | ||
| - | + | <br /> | |
<h3>References</h3> | <h3>References</h3> | ||
Revision as of 12:05, 26 September 2012

Design challenge
The aim of this project is to design and produce a new multi-color display in which genetically engineered cells function as pixels analogous to how a flat panel display works. In the lab we will make living cells that emit light in response to an electric stimulus. This can be achieved by genetic modification of yeast cells, through the introduction of fluorescent calcium sensors and calcium channels.
The plasma membrane of the brewer's yeast Saccharomyces cerevisiae contains the CCH1-MID1 channel that is homologous to mammalian voltage-gated calcium channels (VGCCs). It is hypothesized that upon depolarization of the plasma membrane, calcium ions selectively enter the cytoplasm through these channels [1]. Light will be emitted through the fluorescence of the GECO protein [2], a calcium sensor that is expressed from a genetically engineered plasmid. When the calcium concentration is high, the GECO protein will fluoresce, but when it is very low, the protein will not fluoresce. Electrical stimulation of the cell will allow calcium to enter into the cytoplasm through the calcium channels and the GECO proteins will start to fluoresce. After a while the calcium concentration will drop to homeostatic levels through active transport of calcium ions by the yeast's vacuole and the fluorescence will cease.
Challenges in the laboratory can be found in creating yeast cells with both GECO proteins and a sufficient number of calcium channels incorporated. The important design choices regarding the biological work are motivated below.
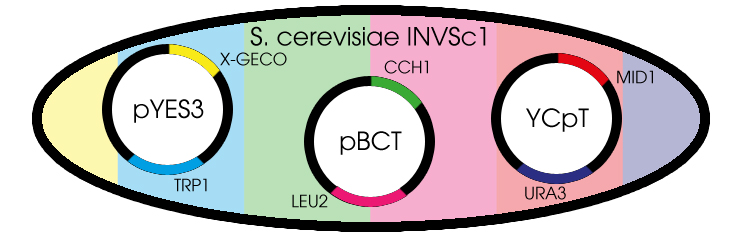
Fig. 1 Yeast cell with the needed plasmids
Chassis
Before this light emitting cell display project could start, it was necessary to decide on a suitable chassis. Candidates were E. coli and S. cerevisiae. Both are common model organisms that can be used for protein expression and are cheap to culture. To reach the goal of this project over expression of voltage-gated calcium channel was needed. As soon as it was found that CCH1-MID1, homologous to mammalian voltage-gated calcium channels, could be over expressed in S. cerevisiae, it was decided to use yeast as the chassis in our project. In the lab we had a strain called INVSc1 available which was compatiable with the plasmids we were planning to use.
Plasmids and transformations
Since we choose yeast as our chassis we need to clone all required genes into shuttle vectors. A shuttle vector is a special type of vector that can be propagated both in yeast and in bacteria. Cloning can be done in fast growing E. coli while proteins can be expressed in yeast. There are low copy-number and high copy-number variants of shuttle vectors, the choice of which depends on the required amount of that protein in the cell. In this project we used three different shuttle vectors, each carrying one gene to be over expressed.
The proteins Cch1 and Mid1 together constitute a high-affinity Ca2+-channel. Both proteins are brought to over expression on a low copy-number shuttle vector. We are thankful to H. Iida & K. Iida for their donation of the shuttle vectors pBCT-CCH1H and YCpT-MID1 which they constructed in earlier research, and which were suited for use in our yeast.
The GECO proteins that were engineered in the group of Robert E. Campbell (Zhao et al., 2011[2]) are available from Addgene.org, a non-profit plasmid sharing service. Via PCR we cloned the GECOs from the shipping plasmid into a pYES3 shuttle vector acquired from Invitrogen (Fig. 1).
Transformation of S. cerevisiae cells can be done with a straightforward heat shock. To obtain a cell containing three different plasmids, three successive transformations are needed. Since we have three different GECOs, in the end we will have three different variants, all containing the CCH1-MID1 calcium channel but each with a different color of GECO.
The order in which plasmids are introduced can be varied. The route requiring the least transformations introduces CCH1 and MID1 first, then the GECOs. However, trying other routes in parallel provides a backup in case certain transformations fail.
Selection of successful transformants is done by auxotrophic markers, one for each plasmid. The yeast strain we use for transformations is deficient in the synthesis of certain amino acids. These usually have to be added to the medium for the yeast to survive. The uptake of a plasmid however, will restore the ability of the cell to synthesize the missing amino acid and therefore survive in the medium.
Compatibility of yeast and device
The genetically modified S. cerevisiae cells will be put to the test in our home-made device, designed for providing electrical stimuli to the yeast cells. These tests are done in a single bath filled with SC medium containing nutrients for our genetically modified yeast and additional free Ca2+ ions. After electrical stimulation of yeast at different positions in the device, optical signals will be expected to be visible to the naked eye. In a more sensitive analysis, isolated protein will be characterized on a spectrophotometer.
GECOs
Real-time imaging of biochemical events inside living cells is important for understanding the molecular basis of physiological processes and diseases[3]. Genetically encoded sensors based on fluorescent proteins (FPs) are frequently used for molecular recognition. In this iGEM project we use fluorescent proteins to provide the light in our display.
A GECO is a protein which emits light in the presence of Ca2+[2]. There are two important classes of genetically encoded Ca2+ indicators. One is called the Forster Resonance Energy Transfer (FRET)-based cameleon type[4] and the other one is called the single Green Fluorescent Protein (GFP) type[5]. The GECO protein belongs to the single GFP type. Research has shown that Ca2+ indicators targeted to the E. coli periplasm can be shifted toward the Ca2+-free or Ca2+ -bound states by manipulation of the environmental Ca2+ concentration[2]. Robert E. Campbell et al. named these Ca2+ indicators GECOs. R-GECO, G-GECO and B-GECO emit red, green or blue light respectively, each with another excitation and emission spectrum (Fig. 2 and Fig. 3).
References
- [1] Iida, K. et al. 2007, Iida, K. et al. ''Essential, Completely Conserved Glycine Residue in the Domain III S2S3 Linker of Voltage-gated Calcium Channel α1 Subunits in Yeast and Mammals'' in ''Journal of Biological Chemistry'' August 31 2007, Vol. 282, issue 35 pp. 25659-25667, DOI: 10.1074/jbc.M703757200
- [2] Zhao et al. 2011, Zhao, Y. et al. ''An Expanded Palette of Genetically Encoded Ca2+ Indicators'' in ''Science'' 30 September 2011, Vol. 333 no. 6051 pp. 1888-1891, DOI: 10.1126/science.1208592
- [3] Laurens Lindenburg and Maarten Merkx, ‘Colorful Calcium Sensors’, 2012
- [4] A. Miyawaki et al., Nature 338, 1997
- [5] J. Nakai, M. Ohkura, K. Imoto, Nat. Biotechnol. (2001)
 "
"