Team:Tokyo Tech/Projects/PHAs/detail/index.htm
From 2012.igem.org
(→B.PHB production in cells and preparation before the confirmation with Nile blue A) |
(→Construction of pha-C1-A-B1 in Biobrick format) |
||
| Line 13: | Line 13: | ||
==Construction of pha-C1-A-B1 in Biobrick format== | ==Construction of pha-C1-A-B1 in Biobrick format== | ||
[[https://2012.igem.org/Team:Tokyo_Tech/Projects/PHAs/index.htm#Construction_of_phaC1-A-B1_in_Biobrick_format Back to "Construction of phaC1-A-B1 in Biobrick format"]] | [[https://2012.igem.org/Team:Tokyo_Tech/Projects/PHAs/index.htm#Construction_of_phaC1-A-B1_in_Biobrick_format Back to "Construction of phaC1-A-B1 in Biobrick format"]] | ||
| - | [[File:tokyotech PHA biobrick.png| | + | [[File:tokyotech PHA biobrick.png|300px|thumb|right|fig3,construction of phaC1-A-B1]] |
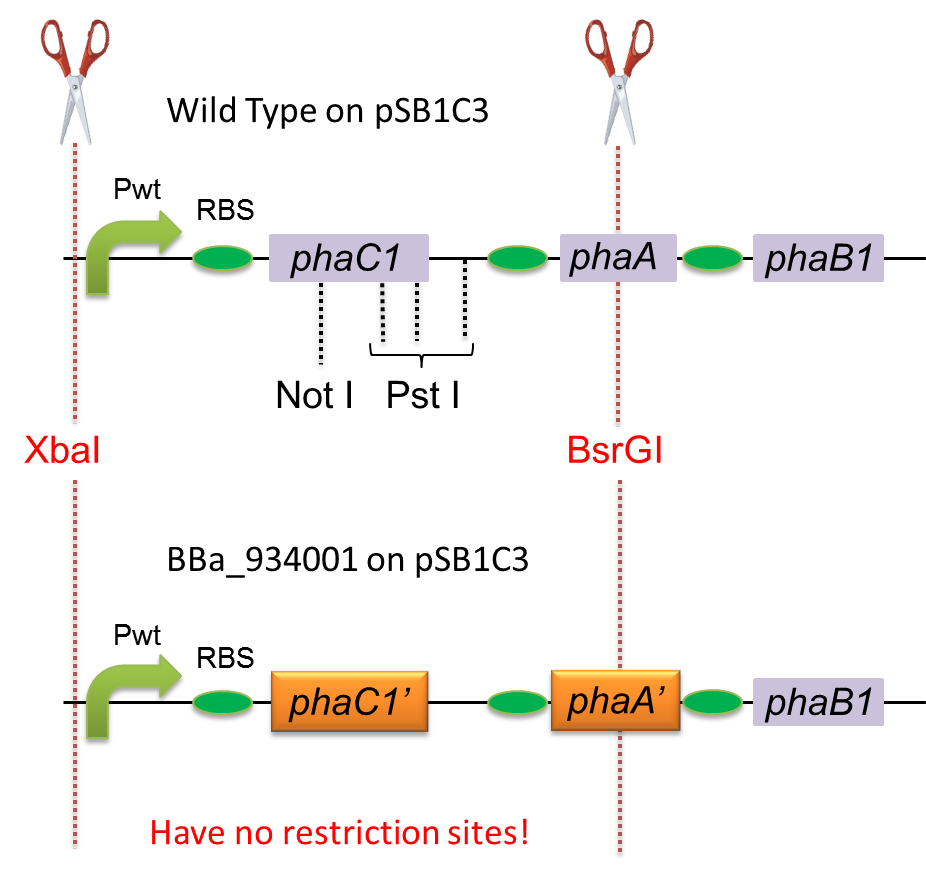
To construct a part that meets Biobrick format, we have modified the phaC1-A-B1 operon not to contain forbidden restriction enzyme sites. First, we cloned the wild type gene phaC1-A-B1 from R.eutropha H16 by using PCR and inserted the gene into pSB1C3. However, wild type phaC1-A-B1 gene sequence contains one NotI and three PstI recognition sites that are not allowed in Biobrick format. To get phaC1-A-B1 sequence without these recognition sites, we ordered the chemically synthesized DNA from IDT/MBL. In this chemically synthesized DNA, coding is optimized for E.coli. We used restriction enzyme XbaI (on pSB1C3) and BsrGI (on phaC1-A-B1) to insert sequence. That is to say, we got Poly[(R)-3-hydroxybutyrate] synthesizing gene in Biobrick format ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K934001 BBa_K934001]). | To construct a part that meets Biobrick format, we have modified the phaC1-A-B1 operon not to contain forbidden restriction enzyme sites. First, we cloned the wild type gene phaC1-A-B1 from R.eutropha H16 by using PCR and inserted the gene into pSB1C3. However, wild type phaC1-A-B1 gene sequence contains one NotI and three PstI recognition sites that are not allowed in Biobrick format. To get phaC1-A-B1 sequence without these recognition sites, we ordered the chemically synthesized DNA from IDT/MBL. In this chemically synthesized DNA, coding is optimized for E.coli. We used restriction enzyme XbaI (on pSB1C3) and BsrGI (on phaC1-A-B1) to insert sequence. That is to say, we got Poly[(R)-3-hydroxybutyrate] synthesizing gene in Biobrick format ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K934001 BBa_K934001]). | ||
Fig3:construction of phaC1-A-B1 | Fig3:construction of phaC1-A-B1 | ||
Revision as of 18:37, 26 September 2012
Materials & Methods
Construction of pha-C1-A-B1 in Biobrick format
[Back to "Construction of phaC1-A-B1 in Biobrick format"]
To construct a part that meets Biobrick format, we have modified the phaC1-A-B1 operon not to contain forbidden restriction enzyme sites. First, we cloned the wild type gene phaC1-A-B1 from R.eutropha H16 by using PCR and inserted the gene into pSB1C3. However, wild type phaC1-A-B1 gene sequence contains one NotI and three PstI recognition sites that are not allowed in Biobrick format. To get phaC1-A-B1 sequence without these recognition sites, we ordered the chemically synthesized DNA from IDT/MBL. In this chemically synthesized DNA, coding is optimized for E.coli. We used restriction enzyme XbaI (on pSB1C3) and BsrGI (on phaC1-A-B1) to insert sequence. That is to say, we got Poly[(R)-3-hydroxybutyrate] synthesizing gene in Biobrick format (BBa_K934001). Fig3:construction of phaC1-A-B1
[Back to "Construction of phaC1-A-B1 in Biobrick format"]
Production trial of PHAs by past teams
[Back to "Construction of phaC1-A-B1 in Biobrick format"]
We introduce some attempts in the past iGEM to show how great our work is in iGEM.
| Part number | Description | States | experience |
| BBa_K338003 | PHA Synthase Composite, Part 1/2 | Planning | |
| BBa_K338004 | PHA Synthase Composite, Part 1/2 | Available | None |
They couldn’t prove that their engineered bacteria produced PHB according to the team wiki.
| Part number | Description | States | experience |
| BBa_K342001 | Pha C (poly-β-hydroxybutyrate polymerase) | Available | None |
They registered and submitted a part, which contains only phaC sequence, as worked. But the team wiki shows that the part doesn’t meet the standard which is required in iGEM because they couldn’t cut their part with Xba1.
| Part number | Description | States | experience |
| BBa_K282000 | phaAB | Available | None |
| Part number | Description | States | experience |
| BBa_K089001 | phaA gene | Planning | |
| BBa_K089002 | phaB gene | Planning | |
| BBa_K089003 | phaC gene | Planning |
| Part number | Description | States | experience |
| BBa_K125801 | RBS-phaA | Available | None |
| BBa_K125802 | RBS-phaB | Planning | |
| BBa_K125803 | RBS-phaC | Available | None |
| BBa_K125804 | RBS-phaE | Available | None |
| BBa_K125501 | phaA BioPlastic polyhydroxybutyrate synthesis pathway (origin PCC6803 slr1994) | Available | None |
| BBa_K125502 | phaB BioPlastic polyhydroxybutyrate synthesis pathway (origin PCC6803 slr1994) | Available | None |
| BBa_K125503 | phaC BioPlastic polyhydroxybutyrate synthesis pathway (origin PCC6803 slr1994) | Available | None |
| BBa_K125504 | phaE BioPlastic polyhydroxybutyrate synthesis pathway (origin PCC6803 slr1994) | Available | None |
| Part number | Description | States | experience |
| BBa_K156031 | RBS + phaA + double terminator | Available | None |
| BBa_K156033 | RBS + phaB1 + double terminator | Available | None |
| BBa_K156034 | RBS + phaC1 + double terminator | Available | None |
| BBa_K156012 | phaA (acetyl-CoA acetyltransferase) | Available | None |
| BBa_K156013 | phaB1 (acetyacetyl-CoA reductase) | Available | None |
| BBa_K156014 | phaC1 (Poly(3-hydroxybutyrate) polymerase) | Available | None |
| BBa_K156021 | Promoter + RBS + phaA + double terminator | Available | None |
| BBa_K156018 | Promoter + RBS + phaA | Planning | |
| BBa_K156019 | Promoter + RBS + phaB1 | Planning | |
| BBa_K156020 | Promoter + RBS + phaC1 | Planning | |
| BBa_K156022 | Promoter + RBS + phaB1 + double terminator | Planning | |
| BBa_K156023 | Promoter + RBS + phaC1 + double terminator | Planning |
[Back to "Construction of phaC1-A-B1 in Biobrick format"]
Protocol
A .PHB production on colonies and preparation before confirmation with Nile red under UV
[Back to "(i)Confirmation of PHB synthesized on colonies"]
1 Preparation of LB agar medium plate containing Nile red and Glucose
1.1 Autoclave a LB agar(final 40g/L) solution at 120 ° C
1.2 After the autoclave, add Chloramphenicol(final 25ug/ml), Nile red and glucose(final 20g/L) to the solution when it cools down.
1.3 Make LB agar medium plates with the mixture.
2 Transformation of pSB1C1 plasmid with phaC1-A-B1 into strain JM109
2.1 Thaw the competent cells JM109 at 4° C
2.2 Add the target DNA 3ul into 1.5ml tube, then add in 50ul the thawed competent cells.
2.3 Put the tube into ice for 15mins
2.4 42° C,30secs, heatshock
2.5 Add 160ul into the tube
2.6 Culture the transformant at 37° C for 30mins
2.7 Paint the transformant on LB agar medium with a large cone rod.
[Back to "(i)Confirmation of PHB synthesized on colonies"]
B.PHB production in cells and preparation before the confirmation with Nile blue A
[Back to "(ii)Confirmation of PHB accumulated in cells"]
1 Production of PHB
1.1 Acquire one colony of the transformed strains (JM109) with a platinum loop
1.2 Culture the colony in LB solution for 16hrs at 37 ° C
1.3 Measure LB medium (final 4%) and add it to each Erlenmeyer flask inside clean bench.
1.4 Add distilled water(final 95%) to each Erlenmeyer flask and cover the flasks with four-folded aluminum foil.
1.5 Set all flasks into autoclave
1.6 Add 1/1000 Chloramphenicol(final 25ug/ml) and glucose solution (50%) (final 20g/L) after the medium is completely cooled.
1.7 Add the solution of cultured cells into each flasks and shaking culture at 37 ° C for 96 hours.
2 Preparation before the confirmation (with Nile blue A) under fluorescent microscope
2.1 Collection of PHBs in JM109
2.1.1 Weigh empty 50ml falcon tube without lid and make a record.
2.1.2 Add some culture solution into each tube.
2.1.3 Set the tubes into centrifuge and make sure that the label faces outside.
2.1.4 4 ° C, 5000G, 10mins in centrifuge.
2.1.5 Remove the supernatant with electric pipettor then add culture solution and set in centrifuge again.
2.1.6 After adding all the culture solution and setting in centrifuge, remove the supernatant and add water, set in centrifuge again.
2.1.7 Remove the supernatant and add a little amount of water
2.1.8 Cover the tubes with double layers of parafilms and fully freeze them.
2.2 Freeze drying (lyophilization)
2.2.1 Poke several holes on the tubes’ parafilm with toothpick.
2.2.2 Set the tubes on the freeze drying machine.
2.2.3 Freeze dry for 3 days.
2.3 Stain PHB accumulated dried cells with Nile blue A before observation
2.3.1 Acquire dried cells after freeze drying
2.3.2 Put a small amount of cells on the slide glass
2.3.3 Add water on the cells and heat the slide glass immobilize the cells
2.3.4 Stain the cells with 1% Nile blue A solution (water) for 8 minutes
2.3.5 Wash excess Nile blue A with 8% acetic acid solution
 "
"