Team:Shenzhen/Result/YAO.Channel
From 2012.igem.org
-
S
BioBricks
- Summary
- YAO.Genome
- YAO.Channel
- YAO.Sensor
- YAO.Suicider
-
p
Notebook
- Team History
- YAO.Genome
- YAO.Channel
- YAO.Sensor
- YAO.Suicider
- YAO.Factory
-
e
Practices



- Causes
- Solution
- Protocol
- 1. SP of Zim17
- 2. SP of ferredoxi
- 3. SP of Tim21
- 4. SP of Tom40
- 5. SP of Tom70
- 6. SP of Tom22
- Purpose
- Materials
- Methods
- I. First Vector Construction
- II. Second Vector Construction
- III. Third Vector Construction
- IV. Construction of yeast expression vector
- V. Yeast transformation
Construction of Signal Peptide System
Due to some inaccurate information in the iGEM website and our own mistakes, some errors accrued when we attempted to link the reporter gene with signal peptide.
The followings are the reason why the errors took place and the approach we applied to solved the problem.
The general adapter is:
GAATTC GCGGCCGC T TCTAGA G[ATG ... TAA TAA] T ACTAGT A GCGGCCG CTGCAG
And general adapter of fusion protein is:
GAATTC GCGGCCGC T TCTAGA [ATG ... TAA TAA] ACTAGT A GCGGCCG CTGCAG
But our adapter of signal peptide is:
GAATTC GCGGCCGC T TCTAG [ATG ... TAA TAA] T ACTAGT A GCGGCCG CTGCAG
Bad news came when we attempted to link the General adapter with the General adapter fragment. The fusion protein is as following:
GAATTC GCGGCCGC T TCTAGA G[part1] T ACTAGA G [part2]T ACTAGT A GCGGCCG CTGCAG
As we can see, there are 8 bases at the fused locus, and in this case, frame shift mutation will occur.
We also produce fusion protein using two General adapter, the result is:
GAATTC GCGGCCGC T TCTAGA [part1] ACTAGA [part2] ACTAGT A GCGGCCG CTGCAG
There are 6 bases at the fused locus. No stop codon occurs, and it will not cause frame shift mutation.
If we use our adapter to link the adapter we designed, the locus is:
GAATTC GCGGCCGC T TCTAG [part1] CTACTAGAG [part2] T ACTAGT A GCGGCCG CTGCAG
In this case, no frame shift mutation occurs, and there are a lot of GFPs available if we select this adapter for use.
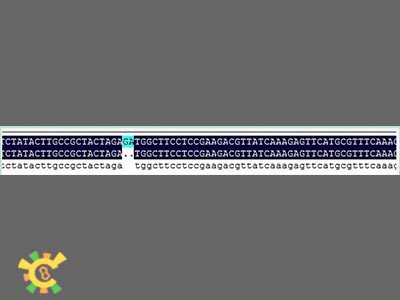
However, when sequencing the fusion protein, we found the fusion locus had lost 2 bases thus causing frameshift mutation, and that the adapter of E1010 linked by backbone PSB1A3 became:
GAATTC GCGGCCGC T TCTAG [ATG ... TAA TAA] T ACTAGT A GCGGCCG CTGCAG
To correct this error, we designed some primers to perform the point mutation.
Table 1. Primers we used in this experiment
5'-GTTTCTTCGAATTCGCGGCCGCTTCTAG
| ZIM17-R</p></ul> | <td>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGATGTGCGATGAT</p></ul> </td>
 "
"