Team:Shenzhen/Result/YAO.Channel
From 2012.igem.org
(Difference between revisions)
| Line 20: | Line 20: | ||
<div id="picshow"> | <div id="picshow"> | ||
| + | </div> | ||
| + | <div class="context"> | ||
| + | <h5>Construction of Signal Peptide System</h5> | ||
| + | <ul><p>Due to some inaccurate information in the iGEM website and our own mistakes, some errors accrued when we attempted to link the reporter gene with signal peptide.</p></ul> | ||
| + | <ul><p>The followings are the reason why the errors took place and the approach we applied to solved the problem.</p></ul> | ||
| + | <ul><li>Causes</li></ul> | ||
| + | <ul><p>The general adapter is:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAGA G[ATG ... TAA TAA] T ACTAGT A GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><p>And general adapter of fusion protein is:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAGA [ATG ... TAA TAA] ACTAGT A GCGGCCG CTGCAG </p></ul> | ||
| + | <ul><p>But our adapter of signal peptide is:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAG [ATG ... TAA TAA] T ACTAGT A GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><p>Bad news came when we attempted to link the General adapter with the General adapter fragment. The fusion protein is as following:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAGA G[part1] T ACTAGA G [part2]T ACTAGT A GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><p>As we can see, there are 8 bases at the fused locus, and in this case, frame shift mutation will occur.</p></ul> | ||
| + | <ul><p>We also produce fusion protein using two General adapter, the result is:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAGA [part1] ACTAGA [part2] ACTAGT A GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><p>There are 6 bases at the fused locus. No stop codon occurs, and it will not cause frame shift mutation.</p></ul> | ||
| + | <ul><p>If we use our adapter to link the adapter we designed, the locus is:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAG [part1] CTACTAGAG [part2] T ACTAGT A GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><p>In this case, no frame shift mutation occurs, and there are a lot of GFPs available if we select this adapter for use.</p></ul> | ||
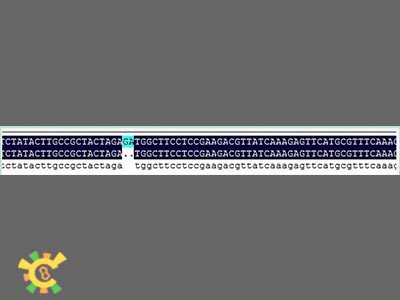
| + | <ul><p>However, when sequencing the fusion protein, we found the fusion locus had lost 2 bases thus causing frameshift mutation, and that the adapter of E1010 linked by backbone PSB1A3 became:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAG [ATG ... TAA TAA] T ACTAGT A GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><div class="figurep"> | ||
| + | [[File:result_yao_channel_p1.jpg]] | ||
| + | <p>Figure 1. Sequence and mapping result.</p></div></ul> | ||
| + | <ul><li>Solution</li></ul> | ||
| + | <ul> | ||
| + | <p>To correct this error, we designed some primers to perform the point mutation.</p> | ||
| + | </ul> | ||
| + | <div class="figurep"><ul><p>Table 1. Primers we used in this experiment</p></ul> | ||
| + | <ul><p> 5'-GTTTCTTCGAATTCGCGGCCGCTTCTAG</p></ul> | ||
| + | <table><tbody><tr><td>ZIM17-R</p></ul> | ||
| + | </td><td>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGATGTGCGATGAT</p></ul> | ||
| + | </td></tr><tr><td>FER-R</p></ul> | ||
| + | </td><td>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGGGCCATGCAGCT</p></ul> | ||
| + | </td></tr><tr><td>TIM21-R</p></ul> | ||
| + | </td><td>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGGGATTTTACTTG</p></ul> | ||
| + | </td></tr><tr><td>TOM22-R</p></ul> | ||
| + | </td><td>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGCGGCAAGTATAG</p></ul> | ||
| + | </td></tr><tr><td>TOM70-R</p></ul> | ||
| + | </td><td>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGGTTGTAATAATA</p></ul> | ||
| + | </td></tr><tr><td>TOM20-R</p></ul> | ||
| + | </td><td>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGCCTGAATTGCGG</p></ul> | ||
| + | </td></tr><tr><td>TOM40-R</p></ul> | ||
| + | </td><td>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGCAATTGAGGAAG</p></ul> | ||
| + | </td></tr></tbody></table></div> | ||
| + | <ul><p>COM-F is the forward primer of all signal peptides, and the other seven are the reverse primers of signal peptides. After our experimental adapters are as followings:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAG [part] CT ACTAG TA T A GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><p>We add three bases in the adapter. If we link the signal peptide at N-terminal, the fused locus is:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAG [part1] CTACTAGAG [part2]T ACTAGT A GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><p> In this case, there are nine bases at the locus, and there aren’t stop codon or frame shift mutations, and our adapter can be used.</p></ul> | ||
| + | <ul><p>After such treatment, final adapter is:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAGA G [part] CT ACTAG TA T A GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><p>If we add signal peptide at the N-terminal of adapter, fusion protein is:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAGAG [part1] CTACTAGAG [part2] CT ACTAGT ATA GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><li>Protocol</li></ul> | ||
| + | <ul><p>Step 1:Extraction plasmid</p></ul> | ||
| + | <ul><p> Zim17, FER, TIM21, TOM22, TOM70, TOM20, TOM50</p></ul> | ||
| + | <ul><p>Step 2:Point mutation</p></ul> | ||
| + | <ul><p> PCR system(take ZIM17 as example):</p></ul> | ||
| + | <ul><p> ZIM17 plasmid template 0.2ul</p></ul> | ||
| + | <ul><p> Ex Taq 0.5ul</p></ul> | ||
| + | <ul><p> 10 x Ex Taq Buffer 2.0ul</p></ul> | ||
| + | <ul><p> dNTP Mix 1.0ul</p></ul> | ||
| + | <ul><p> COM-Fv0.5ul</p></ul> | ||
| + | <ul><p> ZIM17-R 0.5ul</p></ul> | ||
| + | <ul><p> ddH2O 15.3ul</p></ul> | ||
| + | <ul><p> Reaction condition:</p></ul> | ||
| + | <ul><p> 94℃ 5min</p></ul> | ||
| + | <ul><p> 94℃v30s</p></ul> | ||
| + | <ul><p> 55℃ 30s 30 cycle</p></ul> | ||
| + | <ul><p> 72℃ 30s</p></ul> | ||
| + | <ul><p> 72℃ 10min</p></ul> | ||
| + | <ul><p> 12℃ ∞</p></ul> | ||
| + | <ul><p>Step 3:Add 4ul, 6 x loading Buffer, and Run agarose gel. The result are as followings:</p></ul> | ||
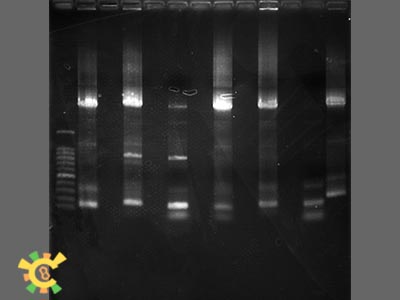
| + | <ul><div class="figurep"> | ||
| + | [[File:result_yao_channel_p2.jpg]] | ||
| + | <p>Figure 2. Result for point mutation.</p></div></ul> | ||
| + | <ul><p>Lane 1.=100bp Marker 2=Tom70 4=Tom40 6=Tom20 8=sZIM!& 10=FER 12=Tom22 13=Tim21</p></ul> | ||
| + | <ul><p>Step 4:gel extraction(use QIAquick Gel Extraction Kit)</p></ul> | ||
| + | <ul><p>Step 5:use NANODROP 2000 to measure concentration.</p></ul> | ||
| + | <ul><p>Step 6: enzyme digestion</p></ul> | ||
| + | <ul><p> enzyme digestion system:</p></ul> | ||
| + | <ul><p> target fragment 8ul</p></ul> | ||
| + | <ul><p> ECoRI 0.5ul</p></ul> | ||
| + | <ul><p> SpeI 0.5u</p></ul> | ||
| + | <ul><p> 10 x H Buffer 1ul</p></ul> | ||
| + | <ul><p> Reaction condition: 37℃ x 3h + 70℃ x 30min</p></ul> | ||
</div> | </div> | ||
Revision as of 07:38, 26 September 2012
 "
"