Team:Shenzhen/Result/YAO.Channel
From 2012.igem.org
(Difference between revisions)
| (15 intermediate revisions not shown) | |||
| Line 21: | Line 21: | ||
<div id="picshow"> | <div id="picshow"> | ||
</div> | </div> | ||
| + | |||
<div class="context"> | <div class="context"> | ||
| Line 26: | Line 27: | ||
<ul><p> Signal peptides are the tools we use to assemble YAO. Channel-Toc and Tic proteins, and to guide the protein into YAO matrix through YAO. Channel. We have designed several Signal Peptide BioBricks (shown in http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2012&group=Shenzhen). </p></ul> | <ul><p> Signal peptides are the tools we use to assemble YAO. Channel-Toc and Tic proteins, and to guide the protein into YAO matrix through YAO. Channel. We have designed several Signal Peptide BioBricks (shown in http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2012&group=Shenzhen). </p></ul> | ||
<ul><p> In total, there are 6 types of signal peptide involved in our project: signal peptide (SP) of Zim17, SP of ferredoxi, SP of Tim21, SP of Tom40, SP of Tom70, SP of Tom22.</p></ul> | <ul><p> In total, there are 6 types of signal peptide involved in our project: signal peptide (SP) of Zim17, SP of ferredoxi, SP of Tim21, SP of Tom40, SP of Tom70, SP of Tom22.</p></ul> | ||
| - | <ul><li> | + | <ul><li>1. SP of Zim17</li> |
| - | 1. SP of Zim17</li><p> | + | <p> Zim17 is a zinc finger protein located in mitochondrial matrix. The signal peptide of Zim17 is amino-terminal segments rich in positively charged residues that form two to three turns of a helix with amphipathic character, and it directs the proteins imported into mitochondrial matrix. The sequence can be found at http://www.uniprot.org/blast/?about=P42844[1-47]. We synthesized the sequence and linked it with RFP & YFP.</p> |
| - | Zim17 is a zinc finger protein located in mitochondrial matrix. The signal peptide of Zim17 is amino-terminal segments rich in positively charged residues that form two to three turns of a helix with amphipathic character, and it directs the proteins imported into mitochondrial matrix. The sequence can be found at http://www.uniprot.org/blast/?about=P42844[1-47]. We synthesized the sequence and linked it with RFP & YFP.</p><p> | + | <p>The document we mainly refer to is:</p> |
| - | The document we mainly refer to is:</p><p> | + | <p>Burri L, Vascotto K, Fredersdorf S, Tiedt R, Hall MN, Lithgow T. Zim17, a novel zinc finger protein essential for protein import into mitochondria. J. Biol. Chem. 2004;279:50243-50249. </p></ul> |
| - | Burri L, Vascotto K, Fredersdorf S, Tiedt R, Hall MN, Lithgow T. Zim17, a novel zinc finger protein essential for protein import into mitochondria. J. Biol. Chem. 2004;279:50243-50249. </p></ul> | + | <ul><li>2. SP of ferredoxi</li> |
| - | <ul><li> | + | <p> The signal peptide of ferredoxi, as a chloroplast-target transit peptide, has an α helix at its N-terminus, followed by an unstructured C-terminal domain of ~19 amino acids. The sequence can be found at http://www.uniprot.org/blast/?about=P07839%5b1-32%5d. We synthesized the sequence and linked it with RFP & YFP.</p> |
| - | 2. SP of ferredoxi</li><p> | + | <p>The document we mainly refer to is:</p> |
| - | The signal peptide of ferredoxi, as a chloroplast-target transit peptide, has an α helix at its N-terminus, followed by an unstructured C-terminal domain of ~19 amino acids. The sequence can be found at http://www.uniprot.org/blast/?about=P07839%5b1-32%5d. We synthesized the sequence and linked it with RFP & YFP.</p><p> | + | <p>Bruce, B. D. (2000). "Chloroplast transit peptides: structure, function and evolution." Trends in Cell Biology 10(10): 440-447. </p></ul> |
| - | The document we mainly refer to is:</p><p> | + | <ul> |
| - | Bruce, B. D. (2000). "Chloroplast transit peptides: structure, function and evolution." Trends in Cell Biology 10(10): 440-447. </p></ul> | + | <li>3. SP of Tim21</li> |
| - | <ul><li> | + | <p> Tim21 is one component of the translocase of the inner mitochondrial membrane (TIM complex), and the signal peptide of Tim21 contains 239 amino acids with a N-terminally single transmembrane segment showing the characteristics of a positively charged mitochondrial presequence with a predicted cleavage site after residue 42. The sequence can be found at http://www.uniprot.org/blast/?about=P53220[1-70]. We synthesized the sequence and linked it with RFP & YFP.</p> |
| - | 3. SP of Tim21</li><p> | + | <p>The document we mainly refer to is:</p> |
| - | Tim21 is one component of the translocase of the inner mitochondrial membrane (TIM complex), and the signal peptide of Tim21 contains 239 amino acids with a N-terminally single transmembrane segment showing the characteristics of a positively charged mitochondrial presequence with a predicted cleavage site after residue 42. The sequence can be found at http://www.uniprot.org/blast/?about=P53220[1-70]. We synthesized the sequence and linked it with RFP & YFP.</p><p> | + | <p>Chacinska, A., M. Lind, et al. (2005). "Mitochondrial Presequence Translocase: Switching between TOM Tethering and Motor Recruitment Involves Tim21 and Tim17." Cell 120(6): 817-829.</p> |
| - | The document we mainly refer to is:</p><p> | + | </ul> |
| - | Chacinska, A., M. Lind, et al. (2005). "Mitochondrial Presequence Translocase: Switching between TOM Tethering and Motor Recruitment Involves Tim21 and Tim17." Cell 120(6): 817-829.</p></ul> | + | |
<ul><li> | <ul><li> | ||
4. SP of Tom40</li><p> | 4. SP of Tom40</li><p> | ||
| Line 57: | Line 57: | ||
Plant Physiology July 2000 vol. 123 no. 3 811-816.</p></ul> | Plant Physiology July 2000 vol. 123 no. 3 811-816.</p></ul> | ||
</div> | </div> | ||
| - | |||
<div class="context"> | <div class="context"> | ||
| - | <h5>Signal Peptides | + | <h5>Construction of Signal Peptide System</h5> |
| - | <ul>< | + | <ul><p>Due to some inaccurate information in the iGEM website and our own mistakes, some errors accrued when we attempted to link the reporter gene with signal peptide.</p></ul> |
| - | <ul><p>Verify that the mitochondrial signal peptides can lead peptides into the mitochondria, or onto the mitochondrial membrane, while the chloroplast signal peptides can not.</p><ul> | + | <ul><p>The followings are the reason why the errors took place and the approach we applied to solved the problem.</p></ul> |
| - | <ul>< | + | <ul><li>Causes</li></ul> |
| - | <ul><p>GFP, Signal peptides (ZIM17, FER, TIM21, TOM22, TOM70, TOM20, TOM40), yeast strain.</p><ul> | + | <ul><p>The general adapter is:</p></ul> |
| - | <ul>< | + | <ul><p> GAATTC GCGGCCGC T TCTAGA G[ATG ... TAA TAA] T ACTAGT A GCGGCCG CTGCAG</p></ul> |
| + | <ul><p>And general adapter of fusion protein is:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAGA [ATG ... TAA TAA] ACTAGT A GCGGCCG CTGCAG </p></ul> | ||
| + | <ul><p>But our adapter of signal peptide is:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAG [ATG ... TAA TAA] T ACTAGT A GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><p>Bad news came when we attempted to link the General adapter with the General adapter fragment. The fusion protein is as following:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAGA G[part1] T ACTAGA G [part2]T ACTAGT A GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><p>As we can see, there are 8 bases at the fused locus, and in this case, frame shift mutation will occur.</p></ul> | ||
| + | <ul><p>We also produce fusion protein using two General adapter, the result is:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAGA [part1] ACTAGA [part2] ACTAGT A GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><p>There are 6 bases at the fused locus. No stop codon occurs, and it will not cause frame shift mutation.</p></ul> | ||
| + | <ul><p>If we use our adapter to link the adapter we designed, the locus is:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAG [part1] CTACTAGAG [part2] T ACTAGT A GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><p>In this case, no frame shift mutation occurs, and there are a lot of GFPs available if we select this adapter for use.</p></ul> | ||
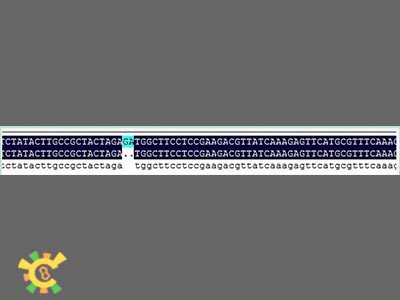
| + | <ul><p>However, when sequencing the fusion protein, we found the fusion locus had lost 2 bases thus causing frameshift mutation, and that the adapter of E1010 linked by backbone PSB1A3 became:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAG [ATG ... TAA TAA] T ACTAGT A GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><div class="figurep"> | ||
| + | [[File:result_yao_channel_p1.jpg]] | ||
| + | <p>Figure 1. Sequence and mapping result.</p></div></ul> | ||
| + | <ul><li>Solution</li></ul> | ||
| + | <ul> | ||
| + | <p>To correct this error, we designed some primers to perform the point mutation.</p> | ||
| + | </ul> | ||
| + | <div class="figurep"><ul><p>Table 1. Primers we used in this experiment</p></ul> | ||
| + | <table> | ||
| + | <tr><td><p>Common</p> | ||
| + | </td><td><p>5'-GTTTCTTCGAATTCGCGGCCGCTTCTAG</p> | ||
| + | </td></tr><tr><td><p>ZIM17-R</p> | ||
| + | </td><td><p>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGATGTGCGATGAT</p> | ||
| + | </td></tr><tr><td><p>FER-R</p> | ||
| + | </td><td><p>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGGGCCATGCAGCT</p> | ||
| + | </td></tr><tr><td><p>TIM21-R</p> | ||
| + | </td><td><p>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGGGATTTTACTTG</p> | ||
| + | </td></tr><tr><td><p>TOM22-R</p> | ||
| + | </td><td><p>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGCGGCAAGTATAG</p> | ||
| + | </td></tr><tr><td><p>TOM70-R</p> | ||
| + | </td><td><p>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGGTTGTAATAATA</p> | ||
| + | </td></tr><tr><td><p>TOM20-R</p> | ||
| + | </td><td><p>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGCCTGAATTGCGG</p> | ||
| + | </td></tr><tr><td><p>TOM40-R</p> | ||
| + | </td><td><p>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGCAATTGAGGAAG</p> | ||
| + | </td></tr></table></div> | ||
| + | <ul><p>COM-F is the forward primer of all signal peptides, and the other seven are the reverse primers of signal peptides. After our experimental adapters are as followings:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAG [part] CT ACTAG TA T A GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><p>We add three bases in the adapter. If we link the signal peptide at N-terminal, the fused locus is:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAG [part1] CTACTAGAG [part2]T ACTAGT A GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><p> In this case, there are nine bases at the locus, and there aren’t stop codon or frame shift mutations, and our adapter can be used.</p></ul> | ||
| + | <ul><p>After such treatment, final adapter is:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAGA G [part] CT ACTAG TA T A GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><p>If we add signal peptide at the N-terminal of adapter, fusion protein is:</p></ul> | ||
| + | <ul><p> GAATTC GCGGCCGC T TCTAGAG [part1] CTACTAGAG [part2] CT ACTAGT ATA GCGGCCG CTGCAG</p></ul> | ||
| + | <ul><li>Protocol</li></ul> | ||
| + | <ul><p>Step 1:Extraction plasmid</p></ul> | ||
| + | <ul><p> Zim17, FER, TIM21, TOM22, TOM70, TOM20, TOM50</p></ul> | ||
| + | <ul><p>Step 2:Point mutation</p></ul> | ||
| + | <ul><p> PCR system(take ZIM17 as example):</p></ul> | ||
| + | <ul><p> ZIM17 plasmid template 0.2ul</p></ul> | ||
| + | <ul><p> Ex Taq 0.5ul</p></ul> | ||
| + | <ul><p> 10 x Ex Taq Buffer 2.0ul</p></ul> | ||
| + | <ul><p> dNTP Mix 1.0ul</p></ul> | ||
| + | <ul><p> COM-Fv0.5ul</p></ul> | ||
| + | <ul><p> ZIM17-R 0.5ul</p></ul> | ||
| + | <ul><p> ddH2O 15.3ul</p></ul> | ||
| + | <ul><p> Reaction condition:</p></ul> | ||
| + | <ul><p> 94℃ 5min</p></ul> | ||
| + | <ul><p> 94℃v30s</p></ul> | ||
| + | <ul><p> 55℃ 30s 30 cycle</p></ul> | ||
| + | <ul><p> 72℃ 30s</p></ul> | ||
| + | <ul><p> 72℃ 10min</p></ul> | ||
| + | <ul><p> 12℃ ∞</p></ul> | ||
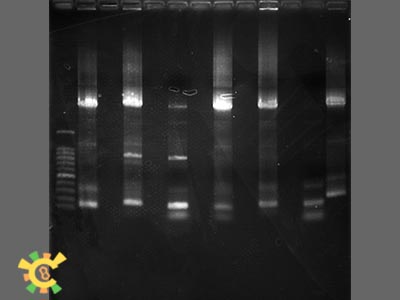
| + | <ul><p>Step 3:Add 4ul, 6 x loading Buffer, and Run agarose gel. The result are as followings:</p></ul> | ||
| + | <ul><div class="figurep"> | ||
| + | [[File:result_yao_channel_p2.jpg]] | ||
| + | <p>Figure 2. Result for point mutation. Lane 1=100bp Marker 2=Tom70 4=Tom40 6=Tom20 8=sZIM!& 10=FER 12=Tom22 13=Tim21</p></div></ul> | ||
| + | <ul><p>Step 4:gel extraction(use QIAquick Gel Extraction Kit)</p></ul> | ||
| + | <ul><p>Step 5:use NANODROP 2000 to measure concentration.</p></ul> | ||
| + | <ul><p>Step 6: enzyme digestion</p></ul> | ||
| + | <ul><p> enzyme digestion system:</p></ul> | ||
| + | <ul><p> target fragment 8ul</p></ul> | ||
| + | <ul><p> ECoRI 0.5ul</p></ul> | ||
| + | <ul><p> SpeI 0.5u</p></ul> | ||
| + | <ul><p> 10 x H Buffer 1ul</p></ul> | ||
| + | <ul><p> Reaction condition: 37℃ x 3h + 70℃ x 30min</p></ul> | ||
| + | </div> | ||
| + | <div class="context"> | ||
| + | <h5>Signal Peptides Validation Protocol</h5> | ||
| + | <ul><li>Purpose</li></ul> | ||
| + | <ul><p>Verify that the mitochondrial signal peptides can lead peptides into the mitochondria, or onto the mitochondrial membrane, while the chloroplast signal peptides can not.</p></ul> | ||
| + | <ul><li>Materials</li></ul> | ||
| + | <ul><p>GFP, Signal peptides (ZIM17, FER, TIM21, TOM22, TOM70, TOM20, TOM40), yeast strain.</p></ul> | ||
| + | <ul><li>Methods</li></ul> | ||
<ul><li>I. First Vector Construction</li></ul> | <ul><li>I. First Vector Construction</li></ul> | ||
| - | <ul><p>1. Synthesis of ZIM17, FER, TIM21, TOM22, TOM70, TOM20, TOM40.</p><ul> | + | <ul><p>1. Synthesis of ZIM17, FER, TIM21, TOM22, TOM70, TOM20, TOM40.</p></ul> |
<ul><p>2. The synthetic fragments are connected with PUC57 the carrier. Mix the strain which containing the carrier and 4 ml of LB+AMP liquid medium in 15 ml centrifuge tube, shake at 37 ℃, 220 rpm overnight.</p></ul> | <ul><p>2. The synthetic fragments are connected with PUC57 the carrier. Mix the strain which containing the carrier and 4 ml of LB+AMP liquid medium in 15 ml centrifuge tube, shake at 37 ℃, 220 rpm overnight.</p></ul> | ||
| - | <ul><p>3. Mix other two strains( BBa-E0030, BBa-E1010 containing biobrick) and 4 ml of LB+AMP liquid medium in 15 ml centrifuge tube, shake at 37 ℃, 220 rpm overnight.</p><ul> | + | <ul><p>3. Mix other two strains( BBa-E0030, BBa-E1010 containing biobrick) and 4 ml of LB+AMP liquid medium in 15 ml centrifuge tube, shake at 37 ℃, 220 rpm overnight.</p></ul> |
| - | <ul><p>4. Extract plasmid ( ZIM17, FER, TIM21, TOM22, TOM70, TOM20, TOM40, BBa-E0030, BBa-E1010).</p><ul> | + | <ul><p>4. Extract plasmid ( ZIM17, FER, TIM21, TOM22, TOM70, TOM20, TOM40, BBa-E0030, BBa-E1010).</p></ul> |
| - | <ul><p>5. Use NANODROP 2000 from Thermo Co. to measure plasmid concentration.</p><ul> | + | <ul><p>5. Use NANODROP 2000 from Thermo Co. to measure plasmid concentration.</p></ul> |
<ul><p>6. Digest: </p><p> | <ul><p>6. Digest: </p><p> | ||
| - | ZIM17, FER, TIM21, TOM22, TOM70, TOM20, TOM40 2-3ug respectively.</p><p> | + | ZIM17, FER, TIM21, TOM22, TOM70, TOM20, TOM40 2-3ug respectively.</p><p> |
| - | ECoRI 2ul</p><p> | + | ECoRI 2ul</p><p> |
| - | SpeI 2ul</p><p> | + | SpeI 2ul</p><p> |
| - | 10 x H Buffer 5ul</p><p> | + | 10 x H Buffer 5ul</p><p> |
| - | ddH2O 50ul</p><p> | + | ddH2O 50ul</p><p> |
| - | BBa-E0030, BBa-E1010 2-3ug respectively</p><p> | + | BBa-E0030, BBa-E1010 2-3ug respectively</p><p> |
| - | ECoRI 2ul</p><p> | + | ECoRI 2ul</p><p> |
| - | XbaI 2ul</p><p> | + | XbaI 2ul</p><p> |
| - | 10 x M Buffer 5ul</p><p> | + | 10 x M Buffer 5ul</p><p> |
| - | ddH2O 50ul</p><p> | + | ddH2O 50ul</p><p> - Reaction conditions:37℃ x 3h + 70℃ x 30min</p></ul> |
| - | Reaction conditions:37℃ x 3h + 70℃ x 30min</p><ul> | + | <ul><p>7. Run a 1% agarose gel 120V 45 min, and EB staining.</p></ul> |
| - | <ul><p>7. Run a 1% agarose gel 120V 45 min, and EB staining.</p><ul> | + | <ul><p>8. Gel Extraction( QIAquick Gel Extraction Kit).</p></ul> |
| - | <ul><p>8. Gel Extraction( QIAquick Gel Extraction Kit).</p><ul> | + | <ul><p>9. Measure the concentration of the recycle fragments.</p></ul> |
| - | <ul><p>9. Measure the concentration of the recycle fragments.</p><ul> | + | <ul><p>10. Ligation:</p><p> |
| - | <ul><p>10. Ligation:</p><p> | + | (1) ZIM17(ECoRI/SpeI) 6ul (2) ZIM17(ECoRI/SpeI) 6ul</p><p> |
| - | (1) ZIM17(ECoRI/SpeI) 6ul (2) ZIM17(ECoRI/SpeI) 6ul</p><p> | + | BBa-E0030(ECoRI/XbaI) 2ul BBa-E1010(ECoRI/XbaI) 2ul</p><p> |
| - | BBa-E0030(ECoRI/XbaI) 2ul BBa-E1010(ECoRI/XbaI) 2ul</p><p> | + | T4 ligase 1ul T4 ligase 1ul</p><p> |
| - | T4 ligase 1ul T4 ligase 1ul</p><p> | + | T4 ligase Buffer 1ul T4 ligase Buffer 1ul</p><p> |
| - | T4 ligase Buffer 1ul T4 ligase Buffer 1ul</p><p> | + | and the same to (3) ~ (14).</p><p> - Ligation condition: Room temperature, 3h.</p></ul> |
| - | and the same to (3) ~ (14).</p><p> | + | <ul><p>11. Transformation</p> |
| - | Ligation condition: Room temperature, 3h.</p><ul> | + | <p> (1) UV sterilization half an hour earlier, and melt competent cells (DH5a) with ice bath.</p><p> |
| - | <ul>< | + | (2) 10 μl ligation product + 50 μl of competent cells, with 30 min ice bath.</p><p> |
| - | <p>(1) UV sterilization half an hour earlier, and melt competent cells (DH5a) with ice bath.</p><p> | + | (3) 42 ℃ water bath heat shock 50s.</p><p> |
| - | (2) 10 μl ligation product + 50 μl of competent cells, with 30 min ice bath.</p><p> | + | (4) cool down with ice for 2 min.</p><p> |
| - | (3) 42 ℃ water bath heat shock 50s.</p><p> | + | (5) Adding 500μl LB liquid medium without antibiotics, 37 ℃ shake (225 rpm) 1 h, recovery.</p><p> |
| - | (4) cool down with ice for 2 min.</p><p> | + | (6) 3000 rpm × 5 min, discard 400μl supernatant and percussion of the remaining liquid and mix.</p><p> |
| - | (5) Adding 500μl LB liquid medium without antibiotics, 37 ℃ shake (225 rpm) 1 h, recovery.</p><p> | + | |
| - | (6) 3000 rpm × 5 min, discard 400μl supernatant and percussion of the remaining liquid and mix.</p><p> | + | |
(7) Overlay 60μl broth on solid LB medium with Kan resistance, 37 ℃, inverted overnight culture in incubator.</p></ul> | (7) Overlay 60μl broth on solid LB medium with Kan resistance, 37 ℃, inverted overnight culture in incubator.</p></ul> | ||
| - | <ul>< | + | <ul><p>12. Validation positive clones</p> |
| - | <p>dNTP mix (2.5mM) 1μl</p><p> | + | <p> dNTP mix (2.5mM) 1μl</p><p> |
| - | PCR 10×buffer 2μl</p><p> | + | PCR 10×buffer 2μl</p><p> |
| - | VF 0.5μl</p><p> | + | VF 0.5μl</p><p> |
| - | VR 0.5μl</p><p> | + | VR 0.5μl</p><p> |
| - | Ex Taq 0.5μl</p><p> | + | Ex Taq 0.5μl</p><p> |
| - | H2O 15.5μl</p><p> | + | H2O 15.5μl</p><p> |
| - | Primer sequence: </p><p> | + | Primer sequence: </p><p> |
| - | VF: TGCCACCTGACGTCTAAGAA</p><p> | + | VF: TGCCACCTGACGTCTAAGAA</p><p> |
| - | VR: ATTACCGCCTTTGAGTGAGC</p><p> | + | VR: ATTACCGCCTTTGAGTGAGC</p><p> |
| - | 94℃ 3 min</p><p> | + | 94℃ 3 min</p><p> |
| - | 94℃ 30s</p><p> | + | 94℃ 30s</p><p> |
| - | 55℃ 30s 30 cycles</p><p> | + | 55℃ 30s 30 cycles</p><p> |
| - | 72℃ 1 min</p><p> | + | 72℃ 1 min</p><p> |
| - | 72℃ 10 min</p><p> | + | 72℃ 10 min</p><p> |
| - | 12℃ ∞</p><p> | + | 12℃ ∞</p><p> |
| - | 1% to 1.5% agarose gel electrophoresis, voltage <130, time ≈ 40 min.</p><p> | + | 1% to 1.5% agarose gel electrophoresis, voltage <130, time ≈ 40 min.</p><p> - Result:</p></ul> |
| - | Result:</p></ul> | + | <ul><p>13. Sequencing</p><p> |
| - | <ul>< | + | Pick single colony of PCR positive clones to sequencing</p></ul> |
| - | Pick single colony of PCR positive clones to sequencing</p><ul> | + | <ul><p>14. Named the correct vectors( for convenient description)</p><p> |
| - | <ul>< | + | ZIM17-E0030, ZIM17-E1010, FER-E0030, FER-E1010, TIM2-E0030, TIM2-E1010, TOM22-E0030, TOM22-E1010, TOM70-E0030, TOM70-E1010, TOM20-E0030, TOM20-E1010, Tom40-E0030, Tom40-E1010</p></ul> |
| - | ZIM17-E0030, ZIM17-E1010, FER-E0030, | + | |
| - | + | ||
| - | TOM70-E1010, TOM20-E0030, | + | |
| - | TOM20-E1010, Tom40-E0030, Tom40-E1010</p></ul> | + | |
<ul><li>II. Second Vector Construction</li></ul> | <ul><li>II. Second Vector Construction</li></ul> | ||
| - | 1. Mix the strain which containing following carrier and 4 ml of LB+AMP liquid medium in 15 ml | + | <ul><p>1. Mix the strain which containing following carrier and 4 ml of LB+AMP liquid medium in 15 ml centrifuge tube, shake at 37 ℃, 220 rpm overnight.</p><p> |
| - | centrifuge tube, shake at 37 ℃, 220 rpm overnight. | + | ZIM17-E0030, ZIM17-E1010, FER-E0030, FER-E1010, TIM2-E0030, TIM2-E1010, TOM22-E0030, TOM22-E1010, TOM70-E0030, TOM70-E1010, TOM20-E0030, TOM20-E1010, Tom40-E0030, Tom40-E1010(kan resistant), T-gz-GAL1(Amp resistant).</p></ul> |
| - | ZIM17-E0030, ZIM17-E1010, FER-E0030, | + | <ul><p>2. Extract plasmid</p></ul> |
| - | + | <ul><p>3. Measure plasmid concentration</p></ul> | |
| - | TOM70-E1010, TOM20-E0030, TOM20-E1010, Tom40-E0030, Tom40-E1010(kan | + | <ul><p>4. Digestion </p><p> |
| - | resistant), T-gz-GAL1(Amp resistant). | + | ECoRI and XbaI:</p><p> |
| - | 2. Extract plasmid | + | ZIM17-E0030, ZIM17-E1010, FER-E0030, FER-E1010, TIM2-E0030, TIM2-E1010, TOM22-E0030, TOM22-E1010, TOM70-E0030, TOM70-E1010, TOM20-E0030, TOM20-E1010, Tom40-E0030, Tom40-E1010</p><p> |
| - | 3. Measure plasmid concentration | + | ECoRI and SpeI:</p><p> |
| - | 4. Digestion | + | T-gz-GAL1</p></ul> |
| - | ECoRI and XbaI: | + | <ul><p>5. Gel electrophoresis, recycle and measure plasmid concentration</p></ul> |
| - | ZIM17-E0030, ZIM17-E1010, FER-E0030, FER- | + | <ul><p>6. Ligation </p><p> |
| - | E1010, TIM2-E0030, TIM2-E1010, TOM22-E0030, TOM22-E1010, TOM70-E0030, | + | ZIM17-E0030, ZIM17-E1010, FER-E0030, FER-E1010, TIM2-E0030, TIM2-E1010, TOM22-E0030, TOM22-E1010, TOM70-E0030, TOM70-E1010, TOM20-E0030, TOM20-E1010, Tom40-E0030, Tom40-E1010 2ul </p><p> |
| - | TOM70-E1010, TOM20-E0030, TOM20-E1010, Tom40-E0030, Tom40-E1010 | + | respectively ligating with T-gz-GAL1 6ul.</p></ul> |
| - | ECoRI and SpeI: | + | <ul><p>7. Transformation</p></ul> |
| - | T-gz-GAL1 | + | <ul><p>8. PCR</p></ul> |
| - | 5. Gel electrophoresis, recycle and measure plasmid concentration | + | <ul><p>9. Shake culture</p></ul> |
| - | 6. Ligation | + | <ul><p>10. Sequencing</p></ul> |
| - | ZIM17-E0030, ZIM17-E1010, FER-E0030, | + | <ul><p>11. Named the correct vectors( for convenient description) </p><p> |
| - | + | GAL-ZIM17-E0030, GAL-ZIM17-E1010, GAL-FER-E0030, GAL-FER-E1010, GAL-TIM2-E0030, GAL-TIM2-E1010, GAL-TOM22-E0030, GAL-TOM22-E1010, GAL-TOM70-E0030, GAL-TOM70-E1010, GAL-TOM20-E0030, GALTOM20-E1010, GAL-Tom40-E0030, GAL-Tom40-E1010.</p></ul> | |
| - | TOM70-E1010, TOM20-E0030, TOM20-E1010, Tom40-E0030, Tom40-E1010 2ul | + | |
| - | respectively ligating with T-gz-GAL1 6ul. | + | |
| - | 7. Transformation | + | |
| - | 8. PCR | + | |
| - | 9. Shake culture | + | |
| - | 10. Sequencing | + | |
| - | 11. Named the correct vectors( for convenient description) | + | |
| - | GAL-ZIM17-E0030, GAL-ZIM17-E1010, GAL-FER-E0030, | + | |
| - | GAL-FER-E1010, GAL-TIM2-E0030, GAL-TIM2-E1010, GAL-TOM22-E0030, | + | |
| - | GAL-TOM22-E1010, GAL-TOM70-E0030, GAL-TOM70-E1010, GAL-TOM20-E0030, GALTOM20-E1010, GAL-Tom40-E0030, GAL-Tom40-E1010. | + | |
<ul><li>III. Third Vector Construction</li></ul> | <ul><li>III. Third Vector Construction</li></ul> | ||
| - | 1. Mix the strain which containing following carrier and 4 ml of LB+AMP liquid medium in 15 ml | + | <ul><p>1. Mix the strain which containing following carrier and 4 ml of LB+AMP liquid medium in 15 ml centrifuge tube, shake at 37 ℃, 220 rpm overnight.</p><p> |
| - | centrifuge tube, shake at 37 ℃, 220 rpm overnight. | + | GAL-ZIM17-E0030, GAL-ZIM17-E1010, GAL-FER-E0030, GAL-FER-E1010, GAL-TIM2-E0030, GAL-TIM2-E1010, GAL-TOM22-E0030, GAL-TOM22-E1010, GAL-TOM70-E0030, GAL-TOM70-E1010, GAL-TOM20-E0030, GAL-TOM20-E1010, GAL-Tom40-E0030, GAL-Tom40-E1010(kan resistant)</p><p> |
| - | GAL-ZIM17-E0030, GAL-ZIM17-E1010, GAL-FER-E0030, | + | BBa-J63002(Amp resistant, yeast ADH terminator)</p></ul> |
| - | GAL-FER-E1010, GAL-TIM2-E0030, GAL-TIM2-E1010, GAL-TOM22-E0030, | + | <ul><p>2. Extract plasmid</p></ul> |
| - | GAL-TOM22-E1010, GAL-TOM70-E0030, GAL-TOM70-E1010, GAL-TOM20-E0030, | + | <ul><p>3. Measure plasmid concentration</p></ul> |
| - | GAL-TOM20-E1010, GAL-Tom40-E0030, GAL-Tom40-E1010(kan resistant) | + | <ul><p>4. Digestion SpeI and PstI with 10X H Buffer:</p><p> |
| - | BBa-J63002(Amp resistant, yeast ADH terminator) | + | GAL-ZIM17-E0030, GAL-ZIM17-E1010, GAL-FER-E0030, GAL-FER-E1010, GAL-TIM2-E0030, GAL-TIM2-E1010, GAL-TOM22-E0030, GAL-TOM22-E1010, GAL-TOM70-E0030, GAL-TOM70-E1010, GAL-TOM20-E0030, GAL-TOM20-E1010, GAL-Tom40-E0030, GAL-Tom40-E1010</p><p> |
| - | 2. Extract plasmid | + | XbaI and PstI with 10X H Buffer: </p><p> |
| - | 3. Measure plasmid concentration | + | BBa-J63002</p></ul> |
| - | 4. Digestion SpeI and PstI with 10X H Buffer: | + | <ul><p>5. Gel electrophoresis, recycle and measure plasmid concentration</p></ul> |
| - | GAL-ZIM17-E0030, GAL-ZIM17-E1010, GAL-FER-E0030, | + | <ul><p>6. Ligation </p><p> |
| - | GAL-FER-E1010, GAL-TIM2-E0030, GAL-TIM2-E1010, GAL-TOM22-E0030, | + | GAL-ZIM17-E0030, GAL-ZIM17-E1010, GAL-FER-E0030, GAL-FER-E1010, GAL-TIM2-E0030, GAL-TIM2-E1010, GAL-TOM22-E0030, GAL-TOM22-E1010, GAL-TOM70-E0030, GAL-TOM70-E1010, GAL-TOM20-E0030, GAL-TOM20-E1010, GAL-Tom40-E0030, GAL-Tom40-E1010 2ul respectively ligating with BBa-J63002 6ul.</p></ul> |
| - | GAL-TOM22-E1010, GAL-TOM70-E0030, GAL-TOM70-E1010, GAL-TOM20-E0030, | + | <ul><p>7. Transformation</p></ul> |
| - | GAL-TOM20-E1010, GAL-Tom40-E0030, GAL-Tom40-E1010 | + | <ul><p>8. PCR</p></ul> |
| - | XbaI and PstI with 10X H Buffer: | + | <ul><p>9. Shake culture</p></ul> |
| - | BBa-J63002 | + | <ul><p>10. Sequencing</p></ul> |
| - | 5. Gel electrophoresis, recycle and measure plasmid concentration | + | <ul><p>11. Named the correct vectors( for convenient description) </p><p> |
| - | 6. Ligation | + | GAL-ZIM17-E0030-J63002, GAL-ZIM17-E1010-J63002, GAL-FER-E0030-J63002, GAL-FER-E1010-J63002, GAL-TIM2-E0030-J63002, GAL-TIM2-E1010-J63002, GAL-TOM22-E0030-J63002, GAL-TOM22-E1010-J63002, GALTOM70-E0030-J63002, GAL-TOM70-E1010-J63002, GAL-TOM20-E0030-J63002, GALTOM20-E1010-J63002, GAL-Tom40-E0030-J63002, GAL-Tom40-E1010-J63002</p></ul> |
| - | GAL-ZIM17-E0030, GAL-ZIM17-E1010, GAL-FER-E0030, | + | |
| - | GAL-FER-E1010, GAL-TIM2-E0030, GAL-TIM2-E1010, GAL-TOM22-E0030, | + | |
| - | GAL-TOM22-E1010, GAL-TOM70-E0030, GAL-TOM70-E1010, GAL-TOM20-E0030, | + | |
| - | GAL-TOM20-E1010, GAL-Tom40-E0030, GAL-Tom40-E1010 | + | |
| - | + | ||
| - | 7. Transformation | + | |
| - | 8. PCR | + | |
| - | 9. Shake culture | + | |
| - | 10. Sequencing | + | |
| - | 11. Named the correct vectors( for convenient description) | + | |
| - | GAL-ZIM17-E0030-J63002, GAL-ZIM17-E1010-J63002, GAL- | + | |
| - | + | ||
| - | GAL-TIM2-E1010-J63002, GAL-TOM22-E0030-J63002, GAL-TOM22-E1010-J63002, GALTOM70-E0030-J63002, GAL-TOM70-E1010-J63002, GAL-TOM20-E0030-J63002, GALTOM20-E1010-J63002, GAL-Tom40-E0030-J63002, GAL-Tom40-E1010-J63002 | + | |
<ul><li>IV. Construction of yeast expression vector</li></ul> | <ul><li>IV. Construction of yeast expression vector</li></ul> | ||
| - | 1. Mix the strain which containing following carrier and 4 ml of LB+AMP liquid medium in 15 ml | + | <ul><p>1. Mix the strain which containing following carrier and 4 ml of LB+AMP liquid medium in 15 ml centrifuge tube, shake at 37 ℃, 220 rpm overnight</p><p> |
| - | centrifuge tube, shake at 37 ℃, 220 rpm overnight | + | GAL-ZIM17-E0030-J63002, GAL-ZIM17-E1010-J63002, GAL-FER-E0030-J63002, GAL-FER-E1010-J63002, GAL-TIM2-E0030-J63002, GAL-TIM2-E1010-J63002, GAL-TOM22-E0030-J63002, GAL-TOM22-E1010-J63002, GALTOM70-E0030-J63002, GAL-TOM70-E1010-J63002, GAL-TOM20-E0030-J63002, GALTOM20-E1010-J63002, GAL-Tom40-E0030-J63002, GAL-Tom40-E1010-J63002(kan resistant)</p><p> |
| - | GAL-ZIM17-E0030-J63002, GAL-ZIM17-E1010-J63002, GAL- | + | YEP352(Amp resistant, yeast ADH terminator)</p></ul> |
| - | + | <ul><p>2. Extract plasmid</p></ul> | |
| - | GAL-TIM2-E1010-J63002, GAL-TOM22-E0030-J63002, GAL-TOM22-E1010-J63002, GALTOM70-E0030-J63002, GAL-TOM70-E1010-J63002, GAL-TOM20-E0030-J63002, GALTOM20-E1010-J63002, GAL-Tom40-E0030-J63002, GAL-Tom40-E1010-J63002(kan | + | <ul><p>3. Measure plasmid concentration</p></ul> |
| - | resistant) | + | <ul><p>4. Digestion </p><p> |
| - | YEP352(Amp resistant, yeast ADH terminator) | + | XbaI and PstI:</p><p> |
| - | 2. Extract plasmid | + | GAL-ZIM17-E0030-J63002, GAL-ZIM17-E1010-J63002, GAL-FER-E0030-J63002, GAL-FER-E1010-J63002, GAL-TIM2-E0030-J63002, GAL-TIM2-E1010-J63002, GAL-TOM22-E0030-J63002, GAL-TOM22-E1010-J63002, GALTOM70-E0030-J63002, GAL-TOM70-E1010-J63002, GAL-TOM20-E0030-J63002, GAL-TOM20-E1010-J63002, GAL-Tom40-E0030-J63002, GAL-Tom40-E1010-J63002, YEP352</p></ul> |
| - | 3. Measure plasmid concentration | + | <ul><p>5. Gel electrophoresis, recycle and measure plasmid concentration</p></ul> |
| - | 4. Digestion | + | <ul><p>6. Ligation </p><p> |
| - | XbaI and PstI: | + | GAL-ZIM17-E0030-J63002, GAL-ZIM17-E1010-J63002, GAL-FER-E0030-J63002, GAL-FER-E1010-J63002, GAL-TIM2-E0030-J63002, GAL-TIM2-E1010-J63002, GAL-TOM22-E0030-J63002, GAL-TOM22-E1010-J63002, GAL-TOM70-E0030-J63002, GAL-TOM70-E1010-J63002, GAL-TOM20-E0030-J63002, GAL-TOM20-E1010-J63002, GAL-Tom40-E0030-J63002, GAL-Tom40-E1010-J63002 6ul respectively ligating with YEP532 2ul.</p></ul> |
| - | GAL-ZIM17-E0030-J63002, GAL-ZIM17-E1010-J63002, GAL- | + | <ul><p>7. Transformation</p></ul> |
| - | + | <ul><p>8. PCR</p></ul> | |
| - | GAL-TIM2-E1010-J63002, GAL-TOM22-E0030-J63002, GAL-TOM22-E1010-J63002, GALTOM70-E0030-J63002, GAL-TOM70-E1010-J63002, GAL-TOM20-E0030-J63002, GAL-TOM20-E1010-J63002, GAL-Tom40-E0030-J63002, GAL-Tom40-E1010-J63002, YEP352 | + | <ul><p>9. Shake culture</p></ul> |
| - | 5. Gel electrophoresis, recycle and measure plasmid concentration | + | <ul><p>10. Sequencing</p></ul> |
| - | 6. Ligation | + | <ul><p>11. Named the correct vectors( for convenient description)</p><p> |
| - | GAL-ZIM17-E0030-J63002, GAL-ZIM17-E1010-J63002, GAL- | + | YEP-GAL-ZIM17-E0030-J63002, YEP-GAL-ZIM17-E1010-J63002, YEPGAL-FER-E0030-J63002, YEP-GAL-FER-E1010-J63002, YEP-GALTIM2-E0030-J63002, YEP-GAL-TIM2-E1010-J63002, YEP-GAL-TOM22-E0030-J63002, YEP-GAL-TOM22-E1010-J63002, YEP-GAL-TOM70-E0030-J63002, YEP-GAL-TOM70-E1010-J63002, YEP-GAL-TOM20-E0030-J63002, YEP-GAL-TOM20-E1010-J63002, YEPGAL-Tom40-E0030-J63002, YEP-GAL-Tom40-E1010-J63002</p></ul> |
| - | + | ||
| - | GAL-TIM2-E1010-J63002, GAL-TOM22-E0030-J63002, GAL-TOM22-E1010-J63002, | + | |
| - | GAL-TOM70-E0030-J63002, GAL-TOM70-E1010-J63002, GAL-TOM20-E0030-J63002, | + | |
| - | GAL-TOM20-E1010-J63002, GAL-Tom40-E0030-J63002, GAL-Tom40-E1010-J63002 6ul | + | |
| - | respectively ligating with YEP532 2ul. | + | |
| - | 7. Transformation | + | |
| - | 8. PCR | + | |
| - | 9. Shake culture | + | |
| - | 10. Sequencing | + | |
| - | 11. Named the correct vectors( for convenient description) | + | |
| - | YEP-GAL-ZIM17-E0030-J63002, YEP-GAL-ZIM17-E1010-J63002, YEPGAL-FER-E0030-J63002, YEP-GAL-FER-E1010-J63002, YEP-GALTIM2-E0030-J63002, YEP-GAL-TIM2-E1010-J63002, YEP-GAL-TOM22-E0030-J63002, | + | |
| - | YEP-GAL-TOM22-E1010-J63002, YEP-GAL-TOM70-E0030-J63002, YEP-GAL-TOM70- | + | |
| - | E1010-J63002, YEP-GAL-TOM20-E0030-J63002, YEP-GAL-TOM20-E1010-J63002, YEPGAL-Tom40-E0030-J63002, YEP-GAL-Tom40-E1010-J63002 | + | |
<ul><li>V. Yeast transformation</li></ul> | <ul><li>V. Yeast transformation</li></ul> | ||
| - | 1. Extract plasmid | + | <ul><p>1. Extract plasmid</p><p> |
| - | YEP-GAL-ZIM17-E0030-J63002, YEP-GAL-ZIM17-E1010-J63002, YEPGAL-FER-E0030-J63002, YEP-GAL-FER-E1010-J63002, YEP-GALTIM2-E0030-J63002, YEP-GAL-TIM2- | + | YEP-GAL-ZIM17-E0030-J63002, YEP-GAL-ZIM17-E1010-J63002, YEPGAL-FER-E0030-J63002, YEP-GAL-FER-E1010-J63002, YEP-GALTIM2-E0030-J63002, YEP-GAL-TIM2-1010-J63002, YEP-GAL-TOM22-E0030-J63002, </p><p> |
| - | YEP-GAL-TOM22-E1010-J63002, YEP-GAL-TOM70-E0030-J63002, YEP-GAL-TOM70- | + | YEP-GAL-TOM22-E1010-J63002, YEP-GAL-TOM70-E0030-J63002, YEP-GAL-TOM70-E1010-J63002, YEP-GAL-TOM20-E0030-J63002, YEP-GAL-TOM20-E1010-J63002, YEPGAL-Tom40-E0030-J63002, YEP-GAL-Tom40-E1010-J63002</p></ul> |
| - | E1010-J63002, YEP-GAL-TOM20-E0030-J63002, YEP-GAL-TOM20-E1010-J63002, YEPGAL-Tom40-E0030-J63002, YEP-GAL-Tom40-E1010-J63002 | + | <ul><p>2. Preparation of yeast competent</p><p> |
| - | 2. Preparation of yeast competent | + | (1) Streak a YPDA agar plate with a small portion of AH109 or Y187. Incubate the plate upside down at 30°C until colonies appear(~3days). Yeast strains can be stored for up to 1 month at 4°C on YPDA medium in culture plates.</p><p> |
| - | (1) | + | (2) Prepare 1.1X TE/LiAc Solution.</p><p> |
| - | Y187. Incubate the plate upside down at 30°C until colonies appear(~3days). Yeast | + | (3) Prepare YPDA liquid medium (Yeast Protocols Handbook).</p> |
| - | strains can be stored for up to 1 month at 4°C on YPDA medium in culture | + | <p> |
| - | plates | + | (4) Inoculate one colony(<4 weeks old, 2–3mm in diameter) into 3ml of YPDA medium in a sterile, 15-ml centrifuge tube.</p><p> |
| - | (2) | + | (5) Incubate at 30°C with shaking for 8hr.</p><p> |
| - | (3) | + | (6) Transfer 5µl of the culture to a 250-ml flask containing 50ml of YPDA.</p><p> |
| - | (4) | + | (7) Incubate at 30°C with shaking at 230–250 rpm for 16–20hr.The OD600 should reach 0.15–0.3.</p><p> |
| - | medium in a sterile, 15-ml centrifuge tube. | + | (8) Centrifuge the cells at 700 x g for 5min at room temperature.</p><p> |
| - | (5) | + | (9) Discard the supernatant and resuspend the cell pellet in 100 ml of YPDA.</p><p> |
| - | (6) | + | (10) Incubate at 30°C for 3–5hr(OD600=0.4~0.5).</p><p> |
| - | (7) | + | (11) Centrifuge the cells at 700 x g for 5min at room temperature.</p><p> |
| - | 0.15–0.3. | + | (12) Discard the supernatant and resuspend the cell pellet in 60ml of sterile, deionized H2O.</p><p> |
| - | (8) | + | (13) Centrifuge the cells at 700 x g for 5min at room temperature.</p><p> |
| - | (9) | + | (14) Discard the supernatant and resuspend the cells in 3 ml of 1.1 X TE/LiAc Solution.</p><p> |
| - | (11) | + | (15) Split the resuspension between two 1.5-ml microcentrifuge tubes (1.5 ml pertube).</p><p> |
| - | (12) | + | (16) Centrifuge each tube at high speed for 15 sec.</p><p> |
| - | deionized H2O. | + | (17) Discard the supernatant and resuspend each pellet in 600 µl of 1.1 X TE/LiAc Solution.</p></ul> |
| - | (13) | + | <ul><p>3. Transformation</p><p> |
| - | (14) | + | (1) 1.5ml centrifuge:</p><p> |
| - | (15) | + | plasmid 0.2ug</p><p> |
| - | + | Sperm DNA*(10 mg/ml) 0.1mg</p><p> | |
| - | (16) | + | Note: Sperm DNA: When freshly prepared water bath boil for 20 minutes, and immediately inserted into the ice bath and stored at -20 °C</p><p> |
| - | (17) | + | (2) Add 100 ul with 1 × TE / LiAc resuspended yeast competent cells per tube, mixed with shaking;</p><p> |
| - | Solution. | + | (3) Each added 600ul PEG / LiAc, shaken vigorously (to improve the conversion efficiency), 30 °C, 200 rpm shaking culture 30 min; </p><p> |
| - | 3. Transformation | + | (4) Each adding 70 ul DMSO, slowly inverted mix (not oscillating), 42 °C water bath thermal shock 15 min, insert into the ice bath to cool 1-2 min; </p><p> |
| - | (1) | + | (5) 14,000rpm × 5 sec, discard supernatant, resuspend cells with 0.1 ml 1×TE, overlay on SD/-Ura solid medium, 30°C inverted culture 3 days in incubator.</p></ul> |
| - | plasmid | + | <ul><p>4. Check out</p><p> |
| - | Sperm DNA*(10 mg/ml) | + | Draw the colonies grown on the said plate in the galactose-containing SD/-Ura solid plates, until the colonies grow, single colonies were picked, and microscopic examination under a fluorescence microscope and record the results.</p></ul> |
| - | + | </div> | |
| - | into the ice bath and stored at -20 °C | + | |
| - | (2) | + | |
| - | shaking; | + | |
| - | (3) | + | |
| - | 30 °C, 200 rpm shaking culture 30 min; | + | |
| - | (4) | + | |
| - | shock 15 min, insert into the ice bath to cool 1-2 min; | + | |
| - | (5) | + | |
| - | Ura solid medium, 30°C inverted culture 3 days in incubator. | + | |
| - | 4. Check out | + | |
| - | Draw the colonies grown on the said plate in the galactose-containing SD/-Ura solid plates, | + | |
| - | until the colonies grow, single colonies were picked, and microscopic examination under a | + | |
| - | fluorescence microscope and record the results. | + | |
</div> | </div> | ||
{{:Team:Shenzhen/Temp/gallery.htm}} | {{:Team:Shenzhen/Temp/gallery.htm}} | ||
Latest revision as of 07:47, 26 September 2012
 "
"