Team:Shenzhen/Result/YAO.Channel
From 2012.igem.org
(Difference between revisions)
| (One intermediate revision not shown) | |||
| Line 20: | Line 20: | ||
<div id="picshow"> | <div id="picshow"> | ||
| + | </div> | ||
| + | |||
| + | |||
| + | <div class="context"> | ||
| + | <h5>Construction of a Signal Peptide System</h5> | ||
| + | <ul><p> Signal peptides are the tools we use to assemble YAO. Channel-Toc and Tic proteins, and to guide the protein into YAO matrix through YAO. Channel. We have designed several Signal Peptide BioBricks (shown in http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2012&group=Shenzhen). </p></ul> | ||
| + | <ul><p> In total, there are 6 types of signal peptide involved in our project: signal peptide (SP) of Zim17, SP of ferredoxi, SP of Tim21, SP of Tom40, SP of Tom70, SP of Tom22.</p></ul> | ||
| + | <ul><li>1. SP of Zim17</li> | ||
| + | <p> Zim17 is a zinc finger protein located in mitochondrial matrix. The signal peptide of Zim17 is amino-terminal segments rich in positively charged residues that form two to three turns of a helix with amphipathic character, and it directs the proteins imported into mitochondrial matrix. The sequence can be found at http://www.uniprot.org/blast/?about=P42844[1-47]. We synthesized the sequence and linked it with RFP & YFP.</p> | ||
| + | <p>The document we mainly refer to is:</p> | ||
| + | <p>Burri L, Vascotto K, Fredersdorf S, Tiedt R, Hall MN, Lithgow T. Zim17, a novel zinc finger protein essential for protein import into mitochondria. J. Biol. Chem. 2004;279:50243-50249. </p></ul> | ||
| + | <ul><li>2. SP of ferredoxi</li> | ||
| + | <p> The signal peptide of ferredoxi, as a chloroplast-target transit peptide, has an α helix at its N-terminus, followed by an unstructured C-terminal domain of ~19 amino acids. The sequence can be found at http://www.uniprot.org/blast/?about=P07839%5b1-32%5d. We synthesized the sequence and linked it with RFP & YFP.</p> | ||
| + | <p>The document we mainly refer to is:</p> | ||
| + | <p>Bruce, B. D. (2000). "Chloroplast transit peptides: structure, function and evolution." Trends in Cell Biology 10(10): 440-447. </p></ul> | ||
| + | <ul> | ||
| + | <li>3. SP of Tim21</li> | ||
| + | <p> Tim21 is one component of the translocase of the inner mitochondrial membrane (TIM complex), and the signal peptide of Tim21 contains 239 amino acids with a N-terminally single transmembrane segment showing the characteristics of a positively charged mitochondrial presequence with a predicted cleavage site after residue 42. The sequence can be found at http://www.uniprot.org/blast/?about=P53220[1-70]. We synthesized the sequence and linked it with RFP & YFP.</p> | ||
| + | <p>The document we mainly refer to is:</p> | ||
| + | <p>Chacinska, A., M. Lind, et al. (2005). "Mitochondrial Presequence Translocase: Switching between TOM Tethering and Motor Recruitment Involves Tim21 and Tim17." Cell 120(6): 817-829.</p> | ||
| + | </ul> | ||
| + | <ul><li> | ||
| + | 4. SP of Tom40</li><p> | ||
| + | Tom40 is the core component of the translocase of the outer mitochondrial membrane (TOM complex), presenting an β-barrel structure. The signal peptide of Tom40 is a conserved β-signal motif, located near the C-terminus, The sequence can be found in the document BMC Genomics, 12, 79. We synthesized the sequence and linked it with RFP & YFP.</p><p> | ||
| + | The document we mainly refer to is:</p><p> | ||
| + | Imai, K., Fujita, N., Gromiha, M.M. and Horton, P. (2011) Eukaryote-wide sequence analysis of mitochondrial beta-barrel outer membrane proteins. BMC Genomics, 12, 79.</p></ul> | ||
| + | <ul><li> | ||
| + | 5. SP of Tom70 </li><p> | ||
| + | Tom70 is also one component of TOM complex, but only contain a single transmembrane segment at their N terminus. The signal peptide of Tom70 is the transmembrane domain comprising a hydrophilic, positively charged segment. The sequence can be found in http://www.uniprot.org/blast/?about=P07213[11-30]. We synthesized the sequence and linked it with RFP & YFP.</p><p> | ||
| + | The document we mainly refer to is:</p><p> | ||
| + | Rapaport, D. (2003). "Finding the right organelle." EMBO Rep 4(10): 948-952. </p></ul> | ||
| + | <ul><li> | ||
| + | 6. SP of Tom22</li><p> | ||
| + | Tom22 is similar with Tom70, but it’s a tail-anchored protein. The signal peptide of Tom22 is also similar with the one of Tom70: moderately hydrophobic, relatively short, with positive charges at the flanking regions. The sequence can be found in http://www.uniprot.org/blast/?about=P49334[98-119]. We synthesized the sequence and linked it with RFP & YFP.</p><p> | ||
| + | The document we mainly refer to is:</p><p> | ||
| + | Plant Physiology July 2000 vol. 123 no. 3 811-816.</p></ul> | ||
</div> | </div> | ||
<div class="context"> | <div class="context"> | ||
| Line 51: | Line 87: | ||
</ul> | </ul> | ||
<div class="figurep"><ul><p>Table 1. Primers we used in this experiment</p></ul> | <div class="figurep"><ul><p>Table 1. Primers we used in this experiment</p></ul> | ||
| - | < | + | <table> |
| - | < | + | <tr><td><p>Common</p> |
| + | </td><td><p>5'-GTTTCTTCGAATTCGCGGCCGCTTCTAG</p> | ||
| + | </td></tr><tr><td><p>ZIM17-R</p> | ||
</td><td><p>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGATGTGCGATGAT</p> | </td><td><p>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGATGTGCGATGAT</p> | ||
</td></tr><tr><td><p>FER-R</p> | </td></tr><tr><td><p>FER-R</p> | ||
| Line 66: | Line 104: | ||
</td></tr><tr><td><p>TOM40-R</p> | </td></tr><tr><td><p>TOM40-R</p> | ||
</td><td><p>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGCAATTGAGGAAG</p> | </td><td><p>5'-GTTTCTTCCTGCAGCGGCCGCTATACTAGTAGCAATTGAGGAAG</p> | ||
| - | </td></tr | + | </td></tr></table></div> |
<ul><p>COM-F is the forward primer of all signal peptides, and the other seven are the reverse primers of signal peptides. After our experimental adapters are as followings:</p></ul> | <ul><p>COM-F is the forward primer of all signal peptides, and the other seven are the reverse primers of signal peptides. After our experimental adapters are as followings:</p></ul> | ||
<ul><p> GAATTC GCGGCCGC T TCTAG [part] CT ACTAG TA T A GCGGCCG CTGCAG</p></ul> | <ul><p> GAATTC GCGGCCGC T TCTAG [part] CT ACTAG TA T A GCGGCCG CTGCAG</p></ul> | ||
| Line 98: | Line 136: | ||
<ul><div class="figurep"> | <ul><div class="figurep"> | ||
[[File:result_yao_channel_p2.jpg]] | [[File:result_yao_channel_p2.jpg]] | ||
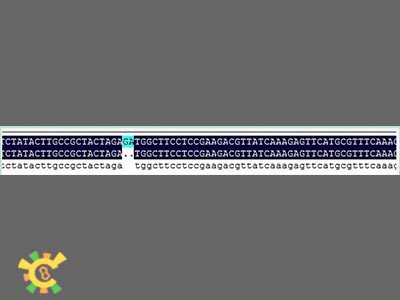
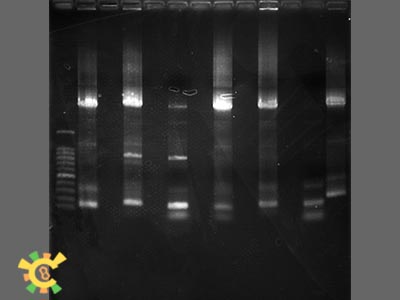
| - | <p>Figure 2. Result for point mutation. | + | <p>Figure 2. Result for point mutation. Lane 1=100bp Marker 2=Tom70 4=Tom40 6=Tom20 8=sZIM!& 10=FER 12=Tom22 13=Tim21</p></div></ul> |
| - | + | ||
<ul><p>Step 4:gel extraction(use QIAquick Gel Extraction Kit)</p></ul> | <ul><p>Step 4:gel extraction(use QIAquick Gel Extraction Kit)</p></ul> | ||
<ul><p>Step 5:use NANODROP 2000 to measure concentration.</p></ul> | <ul><p>Step 5:use NANODROP 2000 to measure concentration.</p></ul> | ||
| Line 110: | Line 147: | ||
<ul><p> Reaction condition: 37℃ x 3h + 70℃ x 30min</p></ul> | <ul><p> Reaction condition: 37℃ x 3h + 70℃ x 30min</p></ul> | ||
</div> | </div> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
<div class="context"> | <div class="context"> | ||
<h5>Signal Peptides Validation Protocol</h5> | <h5>Signal Peptides Validation Protocol</h5> | ||
Latest revision as of 07:47, 26 September 2012
 "
"