Team:Marburg SYNMIKRO/Results
From 2012.igem.org
(Created page with "{{:Team:Marburg_SYNMIKRO:Template:Header}} {{:Team:Marburg_SYNMIKRO:Template:Footer}}") |
(→Biobricks:) |
||
| (68 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{:Team:Marburg_SYNMIKRO:Template:Header}} | {{:Team:Marburg_SYNMIKRO:Template:Header}} | ||
| + | |||
| + | <html> | ||
| + | <div class="conboxb"> | ||
| + | </html> | ||
| + | === '''Biobricks:''' === | ||
| + | |||
| + | '''[http://partsregistry.org/Part:BBa_K866000 BBa_K866000] (CFP-Fusion Brick):''' | ||
| + | |||
| + | In order to evaluate putative recombination events in ''E. coli'', we decided to generate fluorescent fusion proteins, which localize at different compartments within a cell. | ||
| + | [http://partsregistry.org/Part:BBa_E0020 BBa_E0020] is a biobrick coding for the Cyan Fluorescent Protein (CFP). To make CFP suitable for fusion proteins, we used this biobrick as a template to create a CFP coding sequence (CDS) which can be cloned in-frame to other protein coding sequences via [http://partsregistry.org/Assembly:Standard_assembly Standard Assembly]. | ||
| + | |||
| + | Our novel CFP-fusion brick was constructed by site directed mutagenesis through polymerase chain reaction (PCR) using primer [https://2012.igem.org/Team:Marburg_SYNMIKRO/Primer MS_16] and [https://2012.igem.org/Team:Marburg_SYNMIKRO/Primer MS_17]. PCR products were separated by agarose gel electrophoresis, extracted from the gel and cloned into [http://partsregistry.org/Part:pSB1C3 pSB1C3] via [http://partsregistry.org/Help:Standards/Assembly/RFC10 Standard Assembly]. Positive clones were verified by [[Team:Marburg_SYNMIKRO/Notebook#27.08.2012|colony PCR]] and [http://partsregistry.org/cgi/partsdb/dna.cgi?part_name=BBa_K866000 sequencing]. The CFP fusion brick was verified by expression and localization in ''E. coli'' as fusion protein with different intracellular localization domains (A-modules) [[Team:Marburg_SYNMIKRO/Project#Localization domains|GroES and AmiC]] using [[Team:Marburg_SYNMIKRO/Results#Fluorescence microscopy:|fluorescence microscopy]]. | ||
| + | |||
| + | |||
| + | '''[http://partsregistry.org/Part:BBa_K866001 BBa_K866001] (mRFP-Fusion Brick):''' | ||
| + | |||
| + | In addition to the CFP-fusion protein, fusion-competent monomeric Red Fluorescent Protein (mRFP) was contructed from [http://partsregistry.org/Part:BBa_E1010 BBa_E1010] as template. Site directed mutagenesis was performed via PCR using primer [https://2012.igem.org/Team:Marburg_SYNMIKRO/Primer MS14] and [https://2012.igem.org/Team:Marburg_SYNMIKRO/Primer MS15]. PCR products were separated by agarose gel electrophoresis, extracted from the gel and cloned into [http://partsregistry.org/Part:pSB1C3 pSB1C3] via [http://partsregistry.org/Help:Standards/Assembly/RFC10 Standard Assembly]. Positive clones have been verified by [[Team:Marburg_SYNMIKRO/Notebook#27.08.2012|colony PCR]] and [http://partsregistry.org/cgi/partsdb/dna.cgi?part_name=BBa%20K866001 DNA sequencing]. | ||
| + | |||
| + | |||
| + | '''[http://partsregistry.org/Part:BBa_K866002 BBa_K866002] (Gin DNA Invertase):''' | ||
| + | |||
| + | The G-segment Invertase (Gin) from [[Team:Marburg_SYNMIKRO/Project#Phage Mu|phage Mu]] is a site-specific DNA recombinase, which causes inversion of gene segments flanked by [[Team:Marburg_SYNMIKRO/Project#Gin recombination|gix sites]] as inverted repeats. If the gene segment is flanked by gix-sites as direct repeats Gin recombinase causes [[Team:Marburg_SYNMIKRO/Project#Site-specific recombination|deletions]] of the gene segment. Efficient recombination depends on a short DNA-segment which acts as recombinational [[Team:Marburg_SYNMIKRO/Project#Enhancer|Enhancer]]. | ||
| + | To facilitate expression of Gin the biobrick contains a ribosomal binding site (RBS) attached upstream of the Gin start codon. | ||
| + | |||
| + | We designed primer [https://2012.igem.org/Team:Marburg_SYNMIKRO/Primer MS11] and [https://2012.igem.org/Team:Marburg_SYNMIKRO/Primer MS12] to fuse the well characterized and strong RBS [http://partsregistry.org/Part:BBa_J61100 BBa_J61100] from the Anderson RBS-family during amplification of the Gin coding sequence. As template for PCR, chromosomal DNA from ''E. coli'' strain C600 Mucts62 (lysogenic for temperature sensitive phage Mu cts62) was used. PCR amplicons were separated via agarose gel electrophoresis and cloned into [http://partsregistry.org/Part:pSB1C3 pSB1C3] through [http://partsregistry.org/Help:Standards/Assembly/RFC10 Standard Assembly]. Positive clones were evaluated by [[Team:Marburg_SYNMIKRO/Notebook#20.09.2012|restriction enzyme digest]] and [http://partsregistry.org/cgi/partsdb/dna.cgi?part_name=BBa%20K866002 DNA sequencing]. | ||
| + | The [http://partsregistry.org/Part:BBa_K866002 Gin DNA Invertase Biobrick] can be used with promoters of different strength in order to evaluate the optimal level of Gin-expression to stimulate recombination events in ''E. coli''. | ||
| + | |||
| + | |||
| + | '''[http://partsregistry.org/Part:BBa_K866003 BBa_K866003] (SacB Suicide Gene):''' | ||
| + | |||
| + | To allow counter-selection against non-recombined plasmids, the suicide gene ''sacB'' was chosen. In the presence of sucrose in the medium, expression of the ''Bacillus subtilis sacB'' gene is toxic for gram negative ''E. coli'' cells. ''sacB'' codes for the enzyme levansucrase, which uses sucrose to produce the polymeric <html>levan (2,6-β-D-fructosyl)<sub>n</sub></html> in the periplasm, which disturbs cell metabolism and growth, leading to cell death. In our final [[Team:Marburg_SYNMIKRO/Project#Construct|recombination construct]], ''sacB'' will be located between the A and B-modules. Successful recombination will cause deletion of the suicide gene and thus cells that contain plasmids with positive recombination events can be selected on sucrose containing media. | ||
| + | The ''sacB'' gene was amplified using primer [https://2012.igem.org/Team:Marburg_SYNMIKRO/Primer MS26] and [https://2012.igem.org/Team:Marburg_SYNMIKRO/Primer MS27] from the suicide plasmid pNTPS (kindly provided by K. Thormann, MPI Marburg) via PCR. PCR products were separated by agarose gel electrophoresis, extracted from the gel and cloned into [http://partsregistry.org/Part:pSB1C3 pSB1C3] via [http://partsregistry.org/Help:Standards/Assembly/RFC10 Standard Assembly]. Positive clones were verified by [[Team:Marburg_SYNMIKRO/Notebook#17.09.2012|restriction enzyme digest]] and [http://partsregistry.org/cgi/partsdb/dna.cgi?part_name=BBa%20K866003 DNA sequencing]. | ||
| + | [http://partsregistry.org/Part:BBa_K866003 BBa_K866003] can be fused to promoters of different strength and tested for optimal counter-selection through cell death on sucrose containing media. | ||
| + | |||
| + | <html> | ||
| + | </div> | ||
| + | </html> | ||
| + | <html> | ||
| + | <div class="conboxb"> | ||
| + | </html> | ||
| + | |||
| + | === '''Experiments:''' === | ||
| + | |||
| + | |||
| + | ==== Generation of test constructs: ==== | ||
| + | As a first step towards our complete [[Team:Marburg_SYNMIKRO/Project#Construct|recombination construct]], we generated the final recombination gene products i.e. the fusion of a complete A-module with a complete B-module connected by a ''gix'' recombination site. These putative recombination constructs would be the outcome if a recombination event would have occurred in the complete [[Team:Marburg_SYNMIKRO/Project#Construct|recombination construct]]. | ||
| + | For this purpose, all three cellular localization domains (A-Modules), [[Team:Marburg_SYNMIKRO/Project#Localization domains|HU, GroES and AmiC]] have been synthesized. The A modules each consist of the localization domain (AmiC, GroES or HU) preceded by a standard ribosome binding site (RBS) [http://partsregistry.org/Part:BBa_J61100 BBa_J61100] and the strong constitutive promotor [http://partsregistry.org/Part:BBa_J23108 BBa_J23108]. Downstream of the localization domains, a [[Team:Marburg_SYNMIKRO/Project#Gin recombination|''gix'' site]] is located. Between promotor, RBS, localization domain and the gix site, the [http://partsregistry.org/Help:Standards/Assembly/RFC10 scar sequence] from standard assembly was included to test for the correct expression of the reading frame. The complete A-modules are flanked by the biobrick standard assembly [http://partsregistry.org/Help:Standards/Assembly/RFC10 RFC10] prefix and suffix. | ||
| + | |||
| + | The complete A-modules coding for AmiC and GroES were cloned N-terminally to | ||
| + | our fluorescent fusion protein sequences [http://partsregistry.org/Part:BBa_K866000 BBa_K866000] (CFP-Fusion Brick), [http://partsregistry.org/Part:BBa_K866001 BBa_K866001] (mRFP-Fusion brick as well as the biobrick [http://partsregistry.org/Part:BBa_K125500 BBa_K125500] (GFP-Fusion Brick) via standard assembly. Verification of fluorescence was performed via fluorescence microscopy. | ||
| + | |||
| + | ==== Fluorescence microscopy: ==== | ||
| + | In order to screen for fluorescent ''E. coli'' harboring our putative test constructs, positively selected colonies through antibiotic resistance have been subjected to fluorescence microscopy. | ||
| + | For this purpose, colonies growing on LB-Agar supplemented with antibiotics for [http://partsregistry.org/Part:BBa_K866000 BBa_K866000] (CFP-Fusion Brick), [http://partsregistry.org/Part:BBa_K866001 BBa_K866001] (mRFP-Fusion brick) and [http://partsregistry.org/Part:BBa_K125500 BBa_K125500] (GFP-Fusion Brick), respectively, were cultured over-night in LB-medium with appropriate antibiotics. | ||
| + | Small aliquots (10 µL) of overnight cultures were added onto 2% agarose-pads casted on microscope slides, covered by cover-slips and used for fluorescence microscopy. | ||
| + | |||
| + | Altogether, 6 different constructs have been analyzed for fluorescence, being AmiC-GFP, AmiC-CFP, AmiC-mRFP, GroES-GFP, GroES-CFP and GroES-mRFP. | ||
| + | As can be seen in Figure 1, fluorescence was observed in all cells carrying a fusion construct. Especially in cells expressing GroES-GFP and AmiC-GFP (Fig. 1 E and B), intense fluorescence was detectable at cell poles for GroES and in the periplasm for AmiC. Similar localizations were also observed for GroES-mRFP and AmiC-mRFP (Fig. 1 D and A). For GroES-CFP and AmiC-CFP (Fig. 1 F and C), fluorescence is detectable, yet for AmiC-CFP no precise localization could be determined. | ||
| + | |||
| + | As a result, our test constructs representing a putative recombination event of our planned [[Team:Marburg_SYNMIKRO/Project#Construct|recombination construct]] were successfully expressed in ''E.coli''. Furthermore, GFP- and mRFP fused A-modules show enhanced fluorescence at cell poles for GroES and in the periplasm for AmiC. | ||
| + | |||
| + | |||
| + | |||
| + | [[File:Syn_8.jpg|center|600px]] | ||
| + | |||
| + | |||
| + | [[File:Syn_MR_Kontrolle_(P-RBS).jpg|center|600px]] | ||
| + | |||
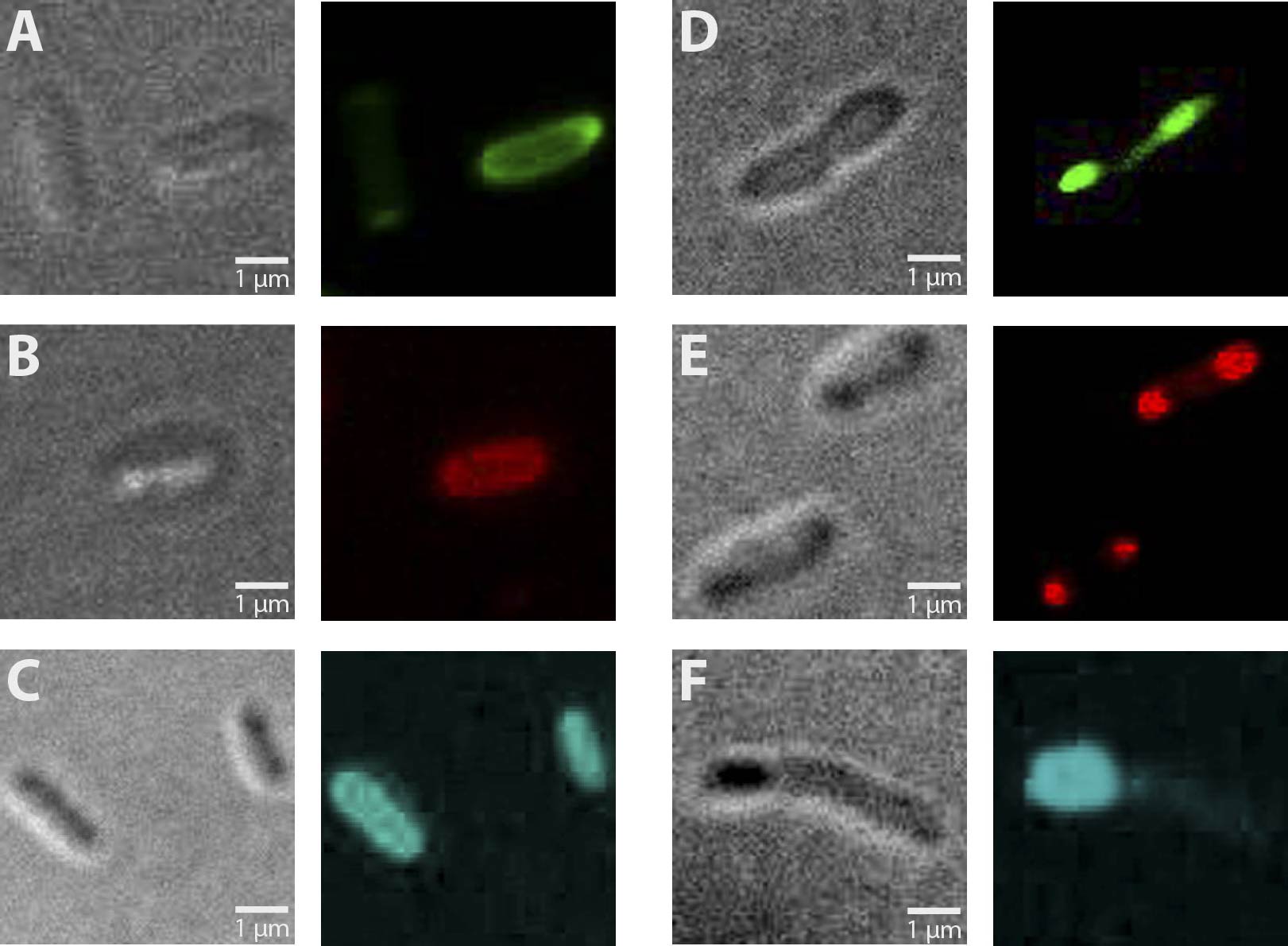
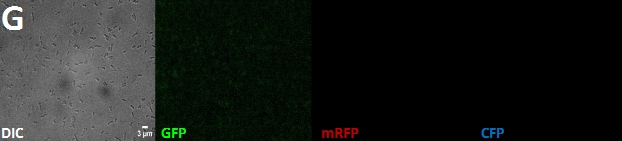
| + | '''Fig. 1''': Fluorescent ''E. coli'' cells harbouring the fusion constructs of [[Team:Marburg_SYNMIKRO/Project#Construct|A- and B-modules]]. '''A''': AmiC-mRFP. '''B''': AmiC-GFP. '''C''': AmiC-CFP. '''D''': GroES-mRFP. '''E''': GroES-GFP. '''F''': GroES-CFP. '''G''': ''E. coli'' harboring a Promotor-RBS construct which serves as a non-fluorescent control. All images were taken by fluorescence microscopy at a magnification of x100. | ||
| + | |||
| + | <html> | ||
| + | </div> | ||
| + | </html> | ||
| + | |||
{{:Team:Marburg_SYNMIKRO:Template:Footer}} | {{:Team:Marburg_SYNMIKRO:Template:Footer}} | ||
Latest revision as of 00:22, 27 September 2012

Contents |
Biobricks:
BBa_K866000 (CFP-Fusion Brick):
In order to evaluate putative recombination events in E. coli, we decided to generate fluorescent fusion proteins, which localize at different compartments within a cell. BBa_E0020 is a biobrick coding for the Cyan Fluorescent Protein (CFP). To make CFP suitable for fusion proteins, we used this biobrick as a template to create a CFP coding sequence (CDS) which can be cloned in-frame to other protein coding sequences via Standard Assembly.
Our novel CFP-fusion brick was constructed by site directed mutagenesis through polymerase chain reaction (PCR) using primer MS_16 and MS_17. PCR products were separated by agarose gel electrophoresis, extracted from the gel and cloned into pSB1C3 via Standard Assembly. Positive clones were verified by colony PCR and sequencing. The CFP fusion brick was verified by expression and localization in E. coli as fusion protein with different intracellular localization domains (A-modules) GroES and AmiC using fluorescence microscopy.
BBa_K866001 (mRFP-Fusion Brick):
In addition to the CFP-fusion protein, fusion-competent monomeric Red Fluorescent Protein (mRFP) was contructed from BBa_E1010 as template. Site directed mutagenesis was performed via PCR using primer MS14 and MS15. PCR products were separated by agarose gel electrophoresis, extracted from the gel and cloned into pSB1C3 via Standard Assembly. Positive clones have been verified by colony PCR and DNA sequencing.
BBa_K866002 (Gin DNA Invertase):
The G-segment Invertase (Gin) from phage Mu is a site-specific DNA recombinase, which causes inversion of gene segments flanked by gix sites as inverted repeats. If the gene segment is flanked by gix-sites as direct repeats Gin recombinase causes deletions of the gene segment. Efficient recombination depends on a short DNA-segment which acts as recombinational Enhancer. To facilitate expression of Gin the biobrick contains a ribosomal binding site (RBS) attached upstream of the Gin start codon.
We designed primer MS11 and MS12 to fuse the well characterized and strong RBS BBa_J61100 from the Anderson RBS-family during amplification of the Gin coding sequence. As template for PCR, chromosomal DNA from E. coli strain C600 Mucts62 (lysogenic for temperature sensitive phage Mu cts62) was used. PCR amplicons were separated via agarose gel electrophoresis and cloned into pSB1C3 through Standard Assembly. Positive clones were evaluated by restriction enzyme digest and DNA sequencing. The Gin DNA Invertase Biobrick can be used with promoters of different strength in order to evaluate the optimal level of Gin-expression to stimulate recombination events in E. coli.
BBa_K866003 (SacB Suicide Gene):
To allow counter-selection against non-recombined plasmids, the suicide gene sacB was chosen. In the presence of sucrose in the medium, expression of the Bacillus subtilis sacB gene is toxic for gram negative E. coli cells. sacB codes for the enzyme levansucrase, which uses sucrose to produce the polymeric levan (2,6-β-D-fructosyl)n in the periplasm, which disturbs cell metabolism and growth, leading to cell death. In our final recombination construct, sacB will be located between the A and B-modules. Successful recombination will cause deletion of the suicide gene and thus cells that contain plasmids with positive recombination events can be selected on sucrose containing media. The sacB gene was amplified using primer MS26 and MS27 from the suicide plasmid pNTPS (kindly provided by K. Thormann, MPI Marburg) via PCR. PCR products were separated by agarose gel electrophoresis, extracted from the gel and cloned into pSB1C3 via Standard Assembly. Positive clones were verified by restriction enzyme digest and DNA sequencing. BBa_K866003 can be fused to promoters of different strength and tested for optimal counter-selection through cell death on sucrose containing media.
Experiments:
Generation of test constructs:
As a first step towards our complete recombination construct, we generated the final recombination gene products i.e. the fusion of a complete A-module with a complete B-module connected by a gix recombination site. These putative recombination constructs would be the outcome if a recombination event would have occurred in the complete recombination construct. For this purpose, all three cellular localization domains (A-Modules), HU, GroES and AmiC have been synthesized. The A modules each consist of the localization domain (AmiC, GroES or HU) preceded by a standard ribosome binding site (RBS) BBa_J61100 and the strong constitutive promotor BBa_J23108. Downstream of the localization domains, a gix site is located. Between promotor, RBS, localization domain and the gix site, the scar sequence from standard assembly was included to test for the correct expression of the reading frame. The complete A-modules are flanked by the biobrick standard assembly RFC10 prefix and suffix.
The complete A-modules coding for AmiC and GroES were cloned N-terminally to our fluorescent fusion protein sequences BBa_K866000 (CFP-Fusion Brick), BBa_K866001 (mRFP-Fusion brick as well as the biobrick BBa_K125500 (GFP-Fusion Brick) via standard assembly. Verification of fluorescence was performed via fluorescence microscopy.
Fluorescence microscopy:
In order to screen for fluorescent E. coli harboring our putative test constructs, positively selected colonies through antibiotic resistance have been subjected to fluorescence microscopy. For this purpose, colonies growing on LB-Agar supplemented with antibiotics for BBa_K866000 (CFP-Fusion Brick), BBa_K866001 (mRFP-Fusion brick) and BBa_K125500 (GFP-Fusion Brick), respectively, were cultured over-night in LB-medium with appropriate antibiotics. Small aliquots (10 µL) of overnight cultures were added onto 2% agarose-pads casted on microscope slides, covered by cover-slips and used for fluorescence microscopy.
Altogether, 6 different constructs have been analyzed for fluorescence, being AmiC-GFP, AmiC-CFP, AmiC-mRFP, GroES-GFP, GroES-CFP and GroES-mRFP. As can be seen in Figure 1, fluorescence was observed in all cells carrying a fusion construct. Especially in cells expressing GroES-GFP and AmiC-GFP (Fig. 1 E and B), intense fluorescence was detectable at cell poles for GroES and in the periplasm for AmiC. Similar localizations were also observed for GroES-mRFP and AmiC-mRFP (Fig. 1 D and A). For GroES-CFP and AmiC-CFP (Fig. 1 F and C), fluorescence is detectable, yet for AmiC-CFP no precise localization could be determined.
As a result, our test constructs representing a putative recombination event of our planned recombination construct were successfully expressed in E.coli. Furthermore, GFP- and mRFP fused A-modules show enhanced fluorescence at cell poles for GroES and in the periplasm for AmiC.
Fig. 1: Fluorescent E. coli cells harbouring the fusion constructs of A- and B-modules. A: AmiC-mRFP. B: AmiC-GFP. C: AmiC-CFP. D: GroES-mRFP. E: GroES-GFP. F: GroES-CFP. G: E. coli harboring a Promotor-RBS construct which serves as a non-fluorescent control. All images were taken by fluorescence microscopy at a magnification of x100.
 "
"