Team:Tsinghua-A/Wetlab/Part1
From 2012.igem.org
(Difference between revisions)
| Line 31: | Line 31: | ||
<p>In prokaryotic cells, the system consists of three circuits. One induces flip, one flips and the other expresses. According to the properties of the promoter pBAD, we build the first circuit, using arabinose as an inducement to generate the recombinase Cre. After the content of Cre increasing, loxP sites flip, causing the expression of supD tRNA and T7ptag. T7ptag and supD tRNA act as an AND gate which can active T7 promoter by <a href="https://2009.igem.org/Team:PKU_Beijing" style="color:blue;">(iGEM 2009 PKU_Beijing.)</a> Details and truth tables are as follows.</p></br></br> | <p>In prokaryotic cells, the system consists of three circuits. One induces flip, one flips and the other expresses. According to the properties of the promoter pBAD, we build the first circuit, using arabinose as an inducement to generate the recombinase Cre. After the content of Cre increasing, loxP sites flip, causing the expression of supD tRNA and T7ptag. T7ptag and supD tRNA act as an AND gate which can active T7 promoter by <a href="https://2009.igem.org/Team:PKU_Beijing" style="color:blue;">(iGEM 2009 PKU_Beijing.)</a> Details and truth tables are as follows.</p></br></br> | ||
<img src="https://static.igem.org/mediawiki/2012/2/20/THU-AWetlab1.png"/> | <img src="https://static.igem.org/mediawiki/2012/2/20/THU-AWetlab1.png"/> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
<table border="1"> | <table border="1"> | ||
<col width="100"> | <col width="100"> | ||
| Line 75: | Line 70: | ||
<td> 1 </td> | <td> 1 </td> | ||
<td> 1 </td> | <td> 1 </td> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | </br></br> | + | <p style="margin-left:300px;"><b>Before Reverse</b></p> |
| + | </br> | ||
| + | <img src="https://static.igem.org/mediawiki/2012/9/97/THU-AWetlab2.png"/> | ||
| + | |||
| + | </br> | ||
<table border="1"> | <table border="1"> | ||
| Line 151: | Line 135: | ||
</tr> | </tr> | ||
</table> | </table> | ||
| - | + | <p style="margin-left:300px;"><b>After Reverse</b></p> | |
</div> | </div> | ||
<a href="https://2012.igem.org/Team:Tsinghua-A/Wetlab" style="margin-left:40px;font-size:20px;">Return</a> | <a href="https://2012.igem.org/Team:Tsinghua-A/Wetlab" style="margin-left:40px;font-size:20px;">Return</a> | ||
Revision as of 17:54, 26 September 2012

Tsinghua-A::Wetlab::Prokaryotic cells

Construction in Prokaryotic cells
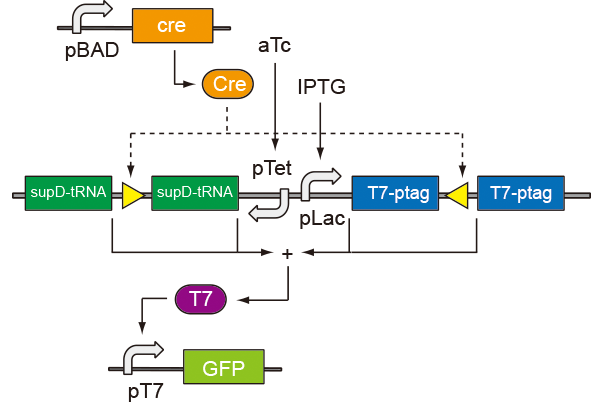
In prokaryotic cells, the system consists of three circuits. One induces flip, one flips and the other expresses. According to the properties of the promoter pBAD, we build the first circuit, using arabinose as an inducement to generate the recombinase Cre. After the content of Cre increasing, loxP sites flip, causing the expression of supD tRNA and T7ptag. T7ptag and supD tRNA act as an AND gate which can active T7 promoter by (iGEM 2009 PKU_Beijing.) Details and truth tables are as follows.
| RSL | Dox | miR-21 | miR-FF4 | Cyan | mKate | EYFP | |||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0 | 0 | 0 | 0 | 0 | 1 | |||||||||||||||||||||||||||||||
| 1 | 0 | 1 | 1 | 0 | 0 | 0 | |||||||||||||||||||||||||||||||
| 0 | 1 | 1 |
| RSL | Dox | miR-21 | miR-FF4 | Cyan | mKate | EYFP |
|---|---|---|---|---|---|---|
| 0 | 0 | 0 | 0 | 0 | 0 | 1 |
| 1 | 0 | 1 | 0 | 1 | 0 | 1 |
| 0 | 1 | 0 | 1 | 0 | 1 | 1 |
| 1 | 1 | 1 | 1 | 1 | 1 | 0 |
After Reverse
Return "
"
