Team:Kyoto/Project
From 2012.igem.org
Contents |
Have you ever seen flower fairies? Probably no(some of you might come across them in your childhood), because they are imaginary creatures existing only in fairy tales. What if we can live with flower fairies? Their lovely powers to make flowers bloom would be profitable for us, including application to agriculture. That’s why we set our project to realize it with synthetic biology, Flower Fairy E.coli!
Our goal is to produce E.coli which can make flowers bloom as Flower Fairies. To make it possible, we focus on FT protein, the identity of Florigen.
There are four issues in order to create Flower Fairy E.coli. It is unclear whether E.coli(prokaryote) could produce functional FT properly because FT is usually produced in the plant cells(eukaryote). After produced, FT have to go through four walls, inner and outer membranes, a cell wall of the plant cells and a cell membrane of the plant. Even though FT could get inside of the cells, it is unknown whether FT protein transcribed in E.coli can activate shoot apex cells and bloom flowers.
We have to go through four steps for purpose of obtaining our goal-Flowering Fairy E.coli-
The four steps are composed of “EXPRESSION”,”SECRETION”,
”PENETRATION”, and ”ACTIVATION”
1.EXPRESSION
On the first step; EXPRESSION, E.coli produce florigen inside itself.
When we got FT gene, we had a difficulty in constructing iGEM parts. The first problem is that FT sequence had two cleavage sites of iGEM restriction enzymes. In order to eliminate cleavage sites of iGEM restriction enzymes, we performed Inverse PCR of plasmids with two kinds of primer which have mutation.
As a result, we could get mutated plasmids, which are not cleaved by iGEM restriction enzymes. In this way, we made FT gene available(Fig.1-2)
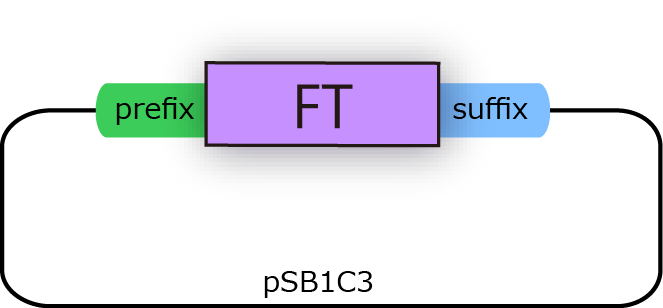
We constructed the plasmid shown in the Fig.1-3.
FT and His tagged FT are regulated by T7 promoter, BBa_I719005 and strong RBS, BBa_B0034.
In order to confirm the expression of FT protein, we performed western blotting using FT specific antibody and checked the place of the FT protein band.
As a result, FT and His:FT bands were observed at the expected molecular weight region.
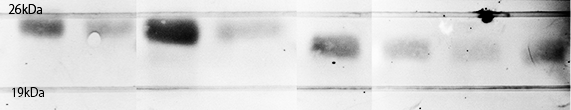
Each plasmid shown in Fig.1-3 was transformed into BL21(DE3). Cells were precultured overnight and diluted into fresh SOC medium. IPTG was added when OD600 was approx. 0.5, then cells were incubated for 4h or overnight at 20°C. 100µL of culture was used for SDS-PAGE. Lane1 : Cell lysate diluted into 100µL of sample buffer, not induced
Lane2 : Cell lysate diluted into 100µL of sample buffer, IPTG induced 4h.
Lane3 : Cell lysate diluted into 100µL of sample buffer, IPTG induced overnight
Lane4 : Cell lysate diluted into 50µL of sample buffer, IPTG induced overnight
Lane5 : Cell lysate diluted into 100µL of sample buffer, not induced
Lane6 : Cell lysate diluted into 100µL of sample buffer, IPTG induced 4h.
Lane7 : Cell lysate diluted into 100µL of sample buffer, IPTG induced overnight
Lane8 : Cell lysate diluted into 50µL of sample buffer, IPTG induced overnight
2.SECRETION
On the second step; SECRETION, E.coli secretes florigen outside the cell.
Even though our E.coli can produce FT protein, a big issue remains: how they can transport proteins to the outside of the cells? To make it possible, we tried to make Tat cassette and kil protein inducer. This cassette makes E.coli carry proteins with torA signal via Tat protein transportation pathway from cytoplasm to periplasm. Periplasm is a space between inner and outer membrane. Kil protein encourages proteins to move from periplasm to surroundings[8]. Our Secretion team made this protein secretion system and visualized and confirmed its function by using GFP.
Result 1: Modified TorA signal
The Twin Arginate Translocation pathway(Tat) is one of the secretion systems E.coli originally has. This system can carry proteins that have torA signal anino acid sequences at N terminal. TatA, TatB and TatC compose Tat complex on inner membrane. Tat complex recognizes torA signal peptide and then it transports proteins (with torA) from cytoplasm to periplasm with maintaining their folding. In short, proteins secreted via Tat pathway can keep active[].
In this experiment, we wanted to design a applicable TorA signal device to meet various needs and to check the function of signal sequence. TorA signal was, actually,submitted by Canbrige 2011(BBa_K233307). These signals, however, doesn't contain RBS so that you need to make RBS by yourself. In addition, old torA signal cause a stop codon between signal peptide and target CDS when you assemble them by standard or 3A assembly. For these reasons, all of other teams make an effort to combine TorA signal to targets, such as using Gibson assembly. That's too trouble!
To avoid to raise a stop codon in scar sequence between torA signal and a target protein CDS, we made Frameshift mutation twice on torA signal. As a result, you can make torA-fusion target protein by standard or 3A assembly. In addition to that, our torA signal has RBS. This can be an easy-to-use biobrick part. It can shorten the time of constructions because it only needs a promoter and a target protein.
We read the sequence data of our modified torA signal and confirmed our torA signal don't raise stop codon when it used in Standard or 3A assembly. Using green fluorescent protein (GFP) as a target protein, we observed the torA-GFP fusion-expressing cells(Fig.1-1,1-2). The torA-GFP fusion was successfully expressed. This means RBS in our torA signal worked.
Result 2: Construction of Tat cassette
TatABC composes a pathway from cytoplasm to periplasm.
Result 3: Assay of Kil
Kil makes holes in outer membrane and we expect that a protein goes through these holes. Needless to say, the function of outer membrane is essential for E.coli to survive. In other words, overexpression of Kil causes cell death. For this reason, we must find the suitable amount of expression.
We made the construct, lacp-RBS-Kil-double terminator, whose backbone is pSB3C5. After culturing for 18hr at 37℃, we eliminated the supernatant using a centrifuge, and diluted it until OD600=0.1. Then we dispensed it. The dispense volume was 3mL. We added 0/0.001/0.01/0.1/1mM IPTG to each. While culturing again at 37℃, we measured OD600. The table below shows the results.
| IPTG | 0 | 0.001mM | 0.01mM | 0.1mM | 1mM |
|---|---|---|---|---|---|
| 0.5h | 0.202 | 0.207 | 0.201 | 0.207 | 0.200 |
| 1h | 0.406 | 0.428 | 0.424 | 0.402 | 0.421 |
| 1.5h | 0.796 | 0.813 | 0.779 | 0.751 | 0.802 |
| 2h | 1.107 | 1.129 | 1.141 | 1.092 | 1.124 |
| 2.5h | 1.546 | 1.565 | 1.541 | 1.532 | 1.578 |
| 3h | 1.912 | 1.933 | 1.890 | 1.883 | 1.940 |
| 3.5h | 2.300 | 2.295 | 2.238 | 2.247 | 2.259 |
| 4h | 2.546 | 2.545 | 2.528 | 2.490 | 2.520 |
| 4.5h | 2.688 | 2.663 | 2.603 | 2.633 | 2.666 |
| 5h | 2.699 | 2.826 | 2.787 | 2.593 | 2.673 |
| 5.5h | 2.863 | 2.742 | 2.754 | 2.741 | 2.756 |
| 6h | 2.831 | 2.876 | 2.758 | 2.759 | 2.728 |
| 20h | 2.671 | 2.706 | 2.680 | 2.606 | 2.619 |
This result indicates that the expression of LacP-RBS-Kil-DT(pSB3C5) makes no effect on suvival of E.coli.
Detail of Our Secretion System
Our wonderful secretion system is constructed by tatABCD, Kil and another gene. Another gene is PspA (phage-shock protein A) gene. E.coli has it originally and this gene is expressed when their inner membrane is damaged. PspA maintains membrane potential and H+ concentration gradient between periplasm and cytoplasm.Our secretion system makes many holes in inner and outer membranes. In other words, E.coli which has our secretion system is under the membrane stress conditions. But by introducing pspA into our Flower Fairy E.coli, the E.coli comes to be able to maintain the vitality, though they have many holes in the membrane.
Construction
Tat secretion cassette with constitutive promoter(BBa_K797004)
This cassette allows E.coli to secrete proteins with torA signal. Wild type Tat protein secretion system is so weak that Kyoto 2012 constructs Tat cassette to reinforce the ability of transportation of Tat system. This part includes tatA, B and C proteins coding region and pspA (phage shock protein A). Tat A, B and C proteins are the main components of Tat complex where proteins with torA signal go through, and pspA can encourage protein secretion via Tat system. Kyoto 2012 suggests this new way of secretion and provides iGEMers with this cassette regulated by constitutive promoter.
We checked the sequence of tatABCD(BBa_K797000) and the sequence of pspA (BBa_K797001) individually, and then, we made TAT construction composed of constitutive promoter(BBa_J23107), tatABCD
(BBa_K797000),pspA(BBa_K797001) and double terminator
(BBa_B0015). This TAT secretion cassette is too long device to sequence, so that we performed electrophoresis of this cassette and confirmed the length of our parts.
Considering that the sequences of tatABCD and pspA are correct,and the length of TAT secretion casssette is correct, we declare that this construction of TAT secretion cassette has been completed.
3.PENETRATION
1. Cool system for penetration --R9 peptide--
On the third step; PENETRATION.
FT protein from Flower Fairy E.coli needs to enter into plant cells in order to induce plants to bloom.
From early stage, we sought various ways to penetrate FT protein into plants, but each way has serious problems.
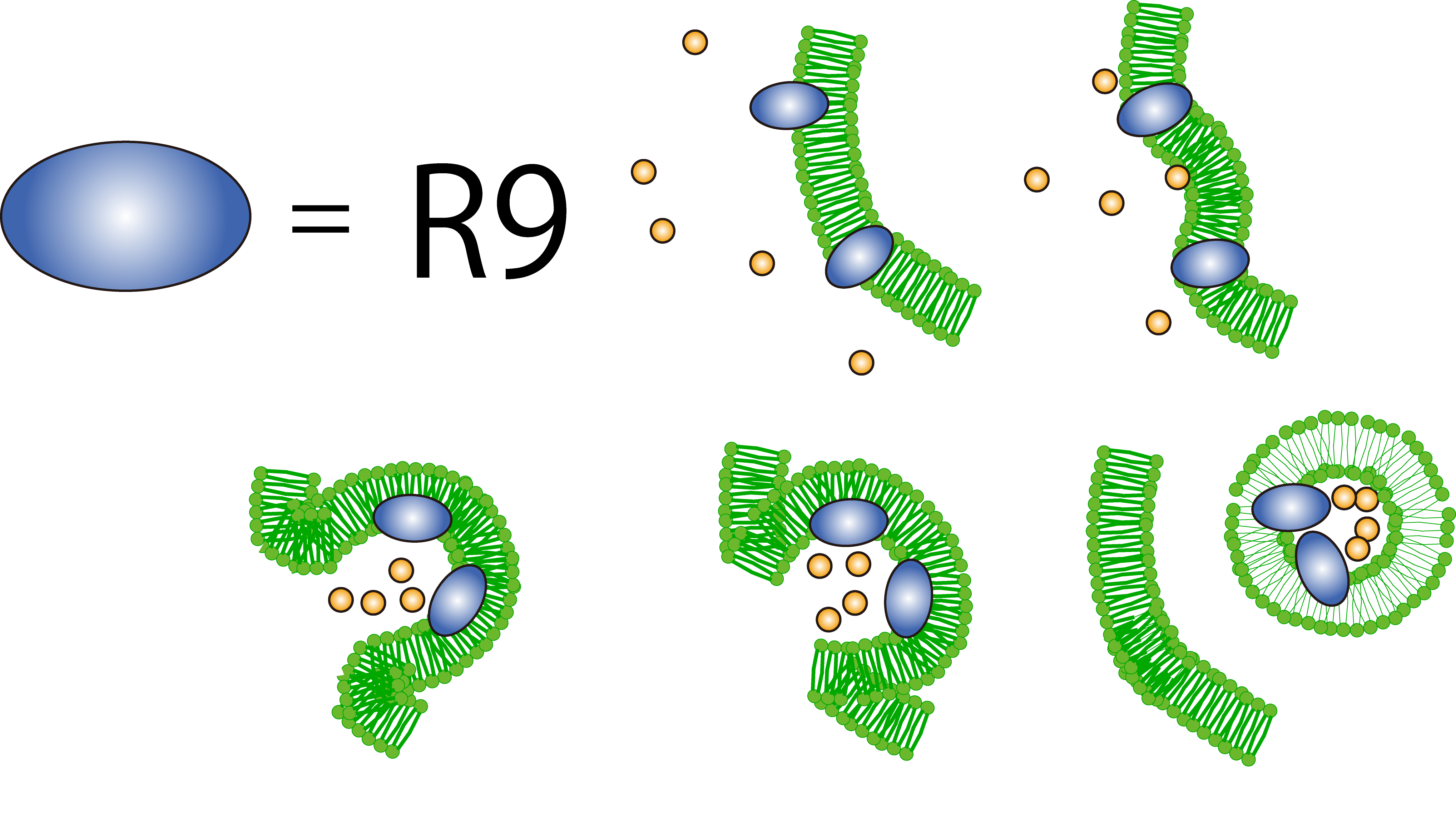
R9 peptide is thought to act on a cell membrane and causes macropinocytosis, a specific form of endocytosis. R9 peptide glues to cell membrane of plants because of hydrophobic.The cell responses to the stimulus and cause macropinocytosis. FT protein around an invaginating region of the cell is taken in the cell.
Plants by themselves practice CPP to transport biomolecules such as proteins inside the cell, in spite of their cell walls.
Then, we determined to cause penetration of FT protein by R9 coding region.
2. GFP antibody specificity check
==
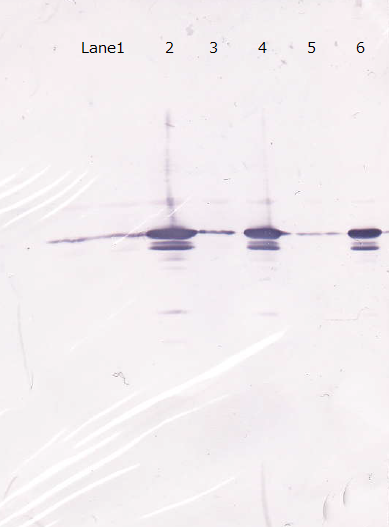
Lane1 : Cell lysate 10µL, not induced
Lane2 : Cell lysate 10µL, IPTG induced
Lane3 : Cell lysate 5µL, not induced
Lane4 : Cell lysate 5µL, IPTG induced
Lane5 : Cell lysate 2µL, not induced
Lane6 : Cell lysate 2µL, IPTG induced
First, we checked the specificity of anti GFP monoclonal antibody, because we must confirm if R9 peptide fusioned GFP is expressed in E.coli.
We used the existing GFP generator parts, [http://partsregistry.org/Part:BBa_I746915 BBa_I746915].
The parts is consist of T7 promoter 6-his tagged superfolder GFP.
Unfortunately, we used inappropriate molecular marker and could'nt confirm the molecular weights of samples.
Each samples induced with IPTG showed one main band and one extra band, and uninduced controls showed one bands.
The relation of intensity of bands between induced samples and controls is corresponded to the existence of IPTG.
From this result, we concluded that GFP antibody has enough specificity.
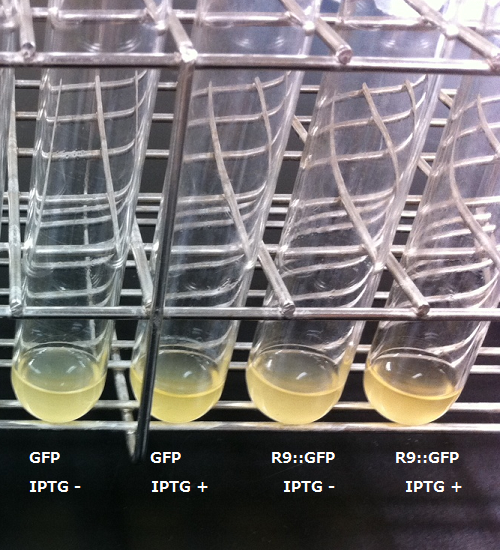
After the 4h of IPTG induction, we noticed that E.coli expressing R9::GFP were growing poorly.
Moreover, we couldn't get any bands of R9::GFP, as shown in the Fig.3-4.
3. Separating R9 peptide and GFP
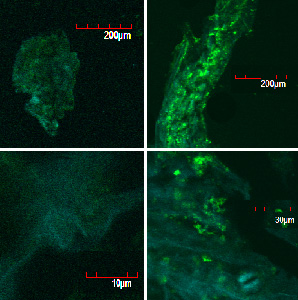
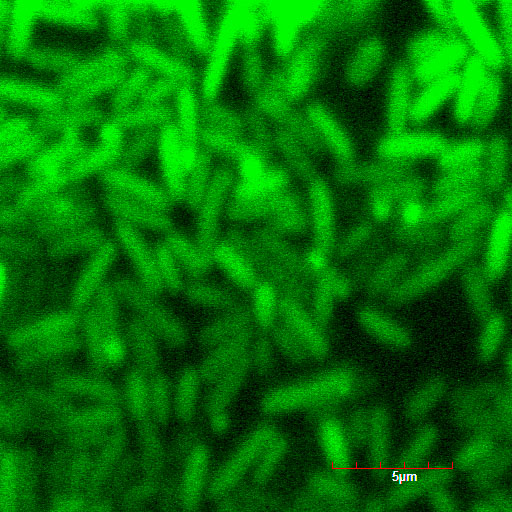
These pictures shows cells of Arabidopsis thaliana leaves soaked in a solution for five minutes, and Hoechst dyeing. Left side samples are soaked in only GFP, the right side samples in GFP and R9 peptide. From top to bottom, nuclei by Hoechst (10 times magnification), GFP fluorescence (10 times), nuclei by Hoechst (60 times), GFP fluorescence (60 times) (With confocal microscope Fluoview FU10i OLYMPUS)
When R9 and GFP were conected, they didn't work normally. By way of experiment, we separated R9::GFP into two segments and soaked plant cells into a sollution of them. After 5 minutes we washed cells by PBS in order to wash GFP away. Then we succeeded in penetration by getting the figure of GFP fluoresence.
The controls on the left were soaked in only GFP, and the samples on the right-hand side were soaked in GFP and R9. These two pictures show the action of R9 peptide. R9 peptide kept GFP in or around plant cells. This figure strongly suggests that R9 peptide works successfully and penetrates cell membrane with GFP.
Fig. Verification of R9 function with use of GFP.
4. Activation
On the final step,Activation. We verified whether FT normally worked in plant cells.
FT protein is derived from plant cells and it is capable of post-translational modification. E.coli cannot do post-translational modification, so FT protein derived from Flower Fairy E.coli may not work normally. As a final step, we tried to confirm whether FT protein by our E.coli led to flower formation.
How to Verify FT Function
[[|thumb|right|300px|Fig. 4-1 FT protein upregulates some flowering genes and they induce plants to bloom. Verification of upreguration of flowering genes ]]
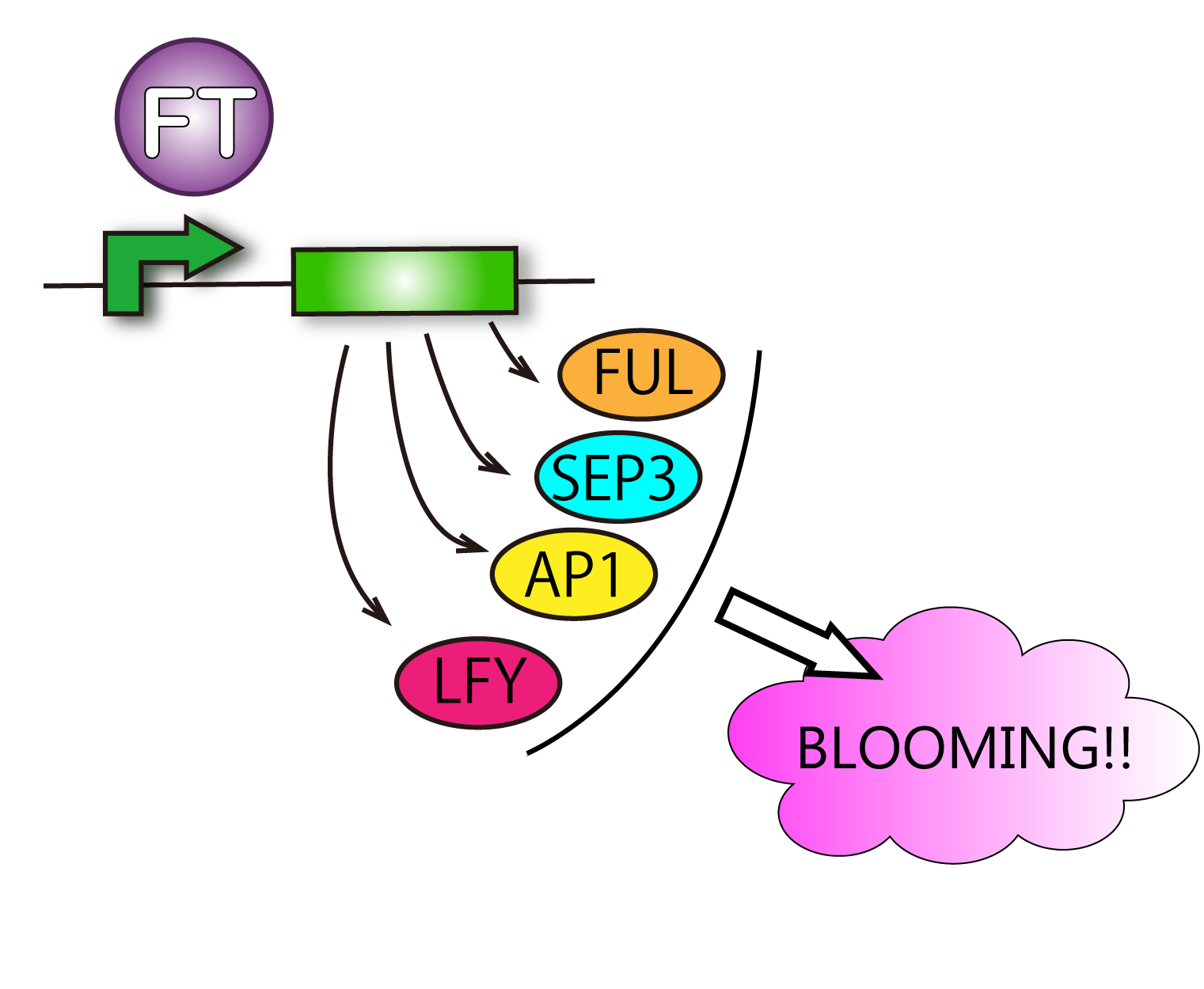
FT protein increases transcriptive activity of several proteins which lead to flower formation. For that reason it can be said that we have verified FT function when we have found rises of activities of the proteins.
Although such proteins activated by FT are various, we check APETALA 1(AP1), SEPALLATA3(SEP3) and FRUITFULL(FUL). This is because AP1 is the representative protein activated by FT, and SEP3 and FUL are activated in leaves. It is difficult for us to handle cells of tips of stems. So we focused on cells of leaves. Leaves' cells are easy to handle for us. We used RT-PCR because of chasing FT protein's function.
We did RT-PCR to compare mRNA expression of Arabidopsis leaves treated with/without FT.
Two types of samples were prepared, one is treated with FT and the other is treated with GFP as control.
GFP was used as control because its molecular weight(27kDa) is relatively similar to that of FT(20kDa.)
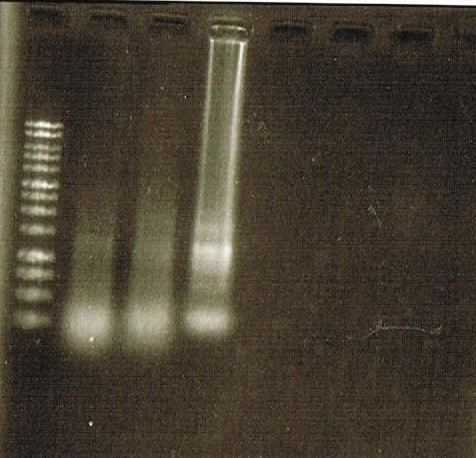
Fig.4-2 is the result of RT-PCR.

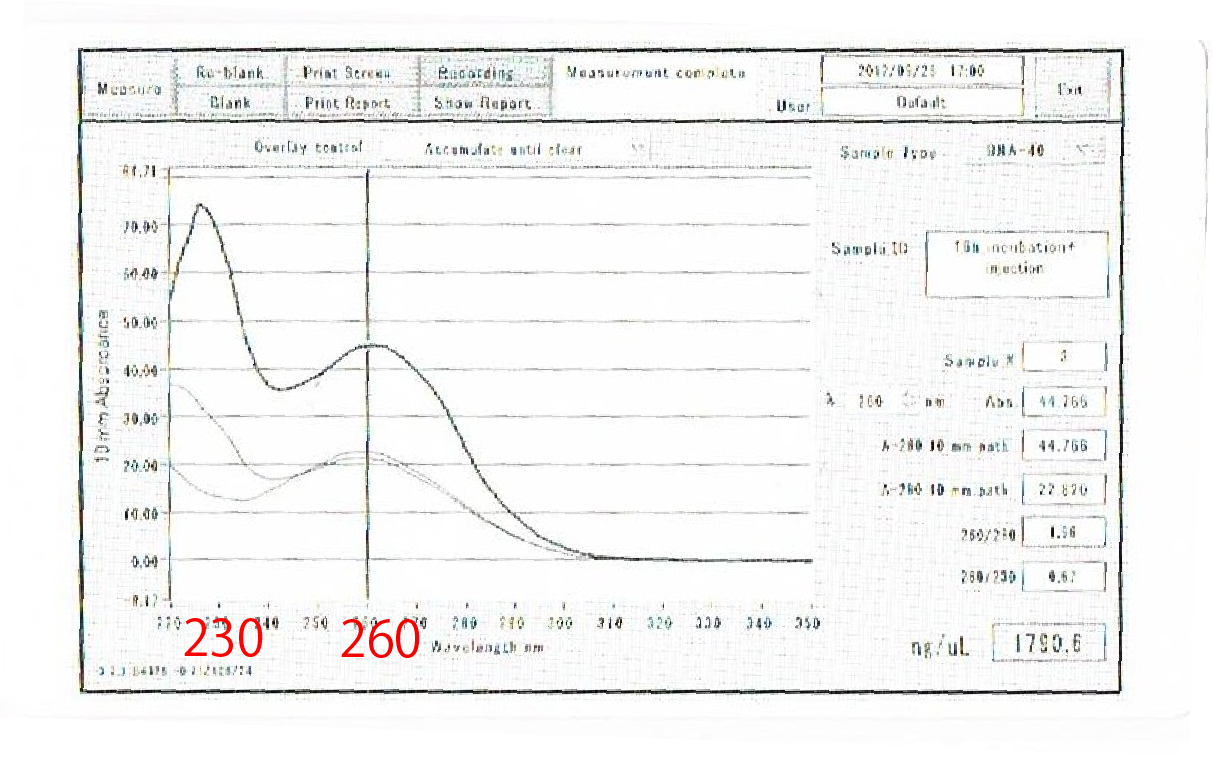
30mg of leaves of Arabidopsis thaliana before bolting were used for one sample. FT or GFP protein and R9 peptide were diluted in PBS (pH7.4), 50ug/L and 500ug/uL each. Leaves were soaked into FT-R9 or GFP-R9 solution for 5min. and incubated for 16hr. in PBS(pH7.4.) After incubation, leaves were freezed with liquid nitrogen and glinded immediately.
Total RNA was extracted by phenol-chloroform extraction. cDNA was synthesized by reverse transcription and used as templates of RT-PCR. TUBULIN was used for internal control of mRNA expression.
Lane1:TUBULIN (GFP-R9 treated) amplicon 61bp
Lane2:TUBULIN (FT-R9 treated)
Lane3:FUL (GFP-R9 treated) amplicon 132bp
Lane4:FUL (FT-R9 treated)
Lane5:SEP3 (GFP-R9 treated) amplicon 87bp
Lane6:SEP3 (FT-R9 treated)
Lane7:AP1 (GFP-R9 treated) amplicon 958bp
From this results, though we could not confirm the function of FT, upregulation of FUL, SEP3, and AP1
but we could amplify each gene successfully.
One possible reason of this failure is the poor quality of extracted RNA, resulted from 16h incubation
after harvesting leaves.
We compared the total RNA bu electrophoresis, shown in fig.4-3.
Achivement
To mutate and standardize FT sequence as a iGEM part.
To confirm expression of FT protein in E.coli.
Future Works
We noticed only flowering and florigen in this time but there are many many other plant hormones. We made translocation pathway from E.coli into plant cells, so we will be able to introduce plant hormones into plant cells if E.coli can make them. It means we can control plant growth in any stage through genetically engineered E.coli. In the future that is not so far, we will be able to meddle in plants' growth――germinating, elongation, flowering, and fructification. We human will finally accomplish a technology that control plants perfectly.
Moreover, R9 peptide functions not only plant cell. R9 peptide works on animal cell similarly. It means that we found a pathway into any kinds of cells. R9 peptide tag enables us to introduce proteins into any cells, so we will be able to controll all living cells using this technology.
[1]Microsugar Chang et al. (2005)"Cellular internalization of fluorescent proteins via arginine-rich intracellular delivery peptide in plant cells" Plant Cell Physiol, 46(3), 482–488
[2]Paula Teper-Bamnolker and Alon Samach1 (2005) "The flowering integrator FT regulates SEPALLATA3 and
FRUITFULL accumulation in Arabidopsis leaves" The Plant Cell, 17, 2661–2675
[3]Philip A. Wigge et al. "Integration of spatial and temporal information during floral induction in Arabidopsis
[4]Sara Trabulo et al.(2010). "Cell-penetrating peptides—mechanisms of cellular uptake and generation of delivery
systems" Pharmaceuticals, 3, 961-993
[5]Unnamalai N, Kang BG, Lee. (2004) "Cationic oligopeptide-mediated delivery of dsRNA for post-transcriptional gene silencing in plant cells." FEBS Lett 21;566(1-3):307-10.
[6]Tracy Palmer and Ben C. Berks.(2012) "The twin-arginine translocation (Tat) protein export pathway" Nat Rev Microbiol, 10(7), 483-96
[7]Choi JH, Lee SY.(2004) "Secretory and extracellular production of recombinant proteins using Escherichia coli" Appl Microbiol Biotechnol, 64(5), 625-35
[8]Miksch G, Fiedler E, Dobrowolski P, Friehs K.(1997) "The kil gene of the ColE1 plasmid of Escherichia coli cntrolled by a growth-phase-dependent promoter mediates the secretion of a heterologous periplasmic protein during the stationary phase" Arch Microbiol, 167(2-3), 143-50
[9]Seibel BA, Walsh PJ.(2002) "Trimethylamine oxide accumulation in marine animals: relationship to acylglycerol storage" J Exp Biol, 205(Pt 3), 297-306
[10]Thomas JD, Daniel RA, Errington J, Robinson C.(2001) "Export of active green fluorescent protein to the periplasm by the twin-arginine translocase (Tat) pathway in Escherichia coli." Mol Microbiol, 39(1), 47-53
[11]Galán JE, Collmer A.(1999) "Type III secretion machines: bacterial devices for protein delivery into host cells." Science, 284(5418), 1322-8
[12]Suit JL, Luria SE.(1988) "Expression of the kil gene of the ColE1 plasmid in Escherichia coli Kilr mutants causes release of periplasmic enzymes and of colicin without cell death." J Bacteriol, 170(10), 4963-6
[13]DeLisa MP, Lee P, Palmer T, Georgiou G.(2004) "Phage shock protein PspA of Escherichia coli relieves saturation of protein export via the Tat pathway." J Bacteriol, 186(2), 366-73

BioBrick is useful for us because we can look for required BioBrick parts from its registory and recombine genes easily. If we want to introduce many parts into one plasmid, however, we have to repeat the process; restrict enzyme digestion and ligation. It takes us too much time and sometimes we lose enough time for other experiments.
We want to reduce the time required for the recombination of genes and get enough time for verification of the expression and the effect of genes.
Golden Gate assembly is the one of the ways to make it possible.
Some teams like 2011 WHU-China have used this assembly. But they didn't intend to spread Golden Gate Assembly through other iGEM teams. So, we tried to make it easier to use this assembly. And we created plasmid backbone parts [http://partsregistry.org/Part:BBa_K797013 "BBa_K797013"].
What's Golden Gate Assembly
Golden Gate Assembly is developed by Carola Engler, Ramona Gruetzner, Romy Kandzia and Sylvestre Marillonnet.
This method enables us introduce plural gene segments into one plasmid all at once.
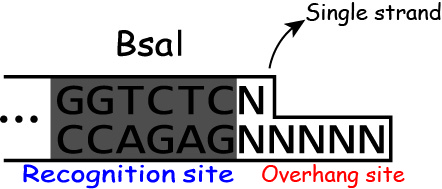
Golden Gate Assembly uses the feature of restrict enzyme "BsaI".
BsaI recognizes the sequence "GGTCTC" and cuts DNA like the figure.
And BsaI activity is independent of the sequences of the downstream of the
recognition site.
Restrict enzyme digestion and ligation are completed by just one PCR because once DNA is cut and ligated irreversibly, the recognition site of BsaI disappears.
Read more about [http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0005553 Golden Gate Assembly].
Before we started assembly, we had to make proper gene segments for Golden Gate assembly. The DNA segments we created had four base pair for ligation, one base pair spacer, BsaI recognition sites and four base pair at the both ends.
We amplified plasmid backbone(psB1K3).
After amplification, we assembled these DNA segments referring to the protocol.
Other iGEM teams can use this backbone plasmid for their Golden Gate assembly.
 "
"