Team:Kyoto/Project
From 2012.igem.org
(→Plasmid backbone (psB1K3)) |
|||
| (355 intermediate revisions not shown) | |||
| Line 18: | Line 18: | ||
<div id="kyoto-tab-contents"> | <div id="kyoto-tab-contents"> | ||
<div id="kyoto-tab-Florigen" class="displayOn"> | <div id="kyoto-tab-Florigen" class="displayOn"> | ||
| - | + | {{Kyoto/Project/FlowerFairy}} | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | { | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
</div> | </div> | ||
<div id="kyoto-tab-GoldenGate"> | <div id="kyoto-tab-GoldenGate"> | ||
| - | + | {{Kyoto/Project/GoldenGate}} | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
</div> | </div> | ||
</div> | </div> | ||
{{Kyoto/footer}} | {{Kyoto/footer}} | ||
Latest revision as of 10:26, 9 October 2012
Realizing Flower Fairy in real world
Have you ever seen flower fairies? Probably the answer is "NO", (though some of you might have come across them in your childhood,) because they are imaginary creatures which exist only in fairy tales. Don’t you think it would be wonderful if you could live with flower fairies? In addition, their lovely power to make flowers bloom would be profitable for us in many ways, such as application to agriculture. This is why we have set our project for realizing Flower Fairy E.coli with synthetic biology!!
In order to realize our dream, we focused on FT protein, known as Florigen.This protein is a kind of plant hormones. First of all, FT protein is produced in leaves, and then move to the shoot apex and bloom flowers. Therefore, FT protein is the key to our project.
Our Goal is to induce flower formation just by putting Flower Fairy E.coli on leaves!
When you want to use our Flower Fairy E.coli, all you have to do is just put them on plant leaves! When you spread Flower Fairy E.coli, FT protein is secreted and penetrate the cell membrane of a plant, and the plant starts blooming.
We had to go through four steps in order to achieve our goal――Flower Fairy E.coli.
These four steps are “EXPRESSION”,”SECRETION”,
”PENETRATION”, and ”ACTIVATION”
In each step, we have some problems to be attacked.
“EXPRESSION”; It is unclear whether E.coli can express FT protein, because FT protein is derived from plant cells .
”SECRETION”; After produced, ''E.coli'' have to secrete FT protein outside of themselves.
”PENETRATION”; FT protein has to penetrate into plant cells.
”ACTIVATION”; Even if FT protein could get into the cells, it is unclear whether FT protein produced by E.coli can activate genes in shoot apex cells and induce flower formation.
1.EXPRESSION
The first step is EXPRESSION.
We had to make E.coli produce FT protein.
However, in the natural environment, wildE.coli don't have FT gene. Therefore we tried to make a new BioBrick part including FT gene and introduce it into E.coli.
Modifying FT gene for Biobrick
FT gene is derived from Arabidopsis thaliana, a model plant. Professor Araki in Kyoto University kindly gave us FT cDNA in TOPO blunt end 2(Invitrogen). FT cDNA has two cleavage sites of iGEM restriction enzymes, EcoR1 and Pst1 (Fig.1-1 A), therefore we modified FT gene sequence by Inverse PCR with primers containing two base mismatches between primer and cDNA(Fig.1-1 B). As a result, we could get mutated FT cDNA, which are not cleaved by iGEM restriction enzymes, and then we added prefix and suffix to FT. Finally, we could make new BioBrick parts of FT (Fig.1-1 C).

A : Necessity for mutation of FT cDNA. FT sequence had two cleavage sites of iGEM restriction enzymes.
B : Inverse PCR Method
Inverse PCR is a measure of mutating FT gene segment. Inverse direction of primers and designing those including mismatch residue leads to mutation of the original plasmid.
C : Standardization of FT cDNA. We had mutated and added prefix and suffix to FT.
Confirming expression of FT
We constructed the plasmid shown in the Fig.1-2. One was FT gene with T7 promoter ([http://partsregistry.org/Part:BBa_I719005 BBa_I719005]) and 6His tag, and the other was FT gene only with T7 promoter. T7 promoter is a strong promotor regulated by T7 polymerase. In our strain of E.coli, the expression of T7 polymerase can be induced by IPTG. 6His tag, which is used in later steps, enabled us to purify FT protein from E.coli using affinity chromatography.
We needed to confirm that E. coli can express FT protein, because there was a possibility that E.coli can not translate FT or FT is unstable or toxic in E.coli.
In order to confirm the expression of FT protein, we performed Western blotting using anti-FT antibody.
As a result, we observed FT and 6 His:FT bands at the expected molecular weight region(Fig.1-2).
So we successfuly confirmed the FT expression!
We succeeded in confirming the mutation and the expression of FT and 6 His:FT protein in E.coli!
2.SECRETION
The second step is SECRETION. We want E.coli to secrete FT protein to outside of the cell.
Now our E.coli can produce FT protein, but a big issue still remains:
how they can transport proteins to the outside of the cells? Many people might think cell lysis is the best way.
But in our project lysis is not very good, because it would possibly cause all fairies' deaths. We hoped to see the continuous effect of Flower Fairy E.coli, not just temporary. So, we adopted a method other than lysis which enables E.coli to transport FT protein continuously.
Proteins are required to go through two membranes
E.coli have two membranes: inner membrane and outer membrane. To transport FT protein to outside of E.coli, FT protein has to pass through the two membranes. So we searched for secretion systems to make FT protein go through these membranes.
TorA signal enables proteins to go through the inner membrane via Tat pathway
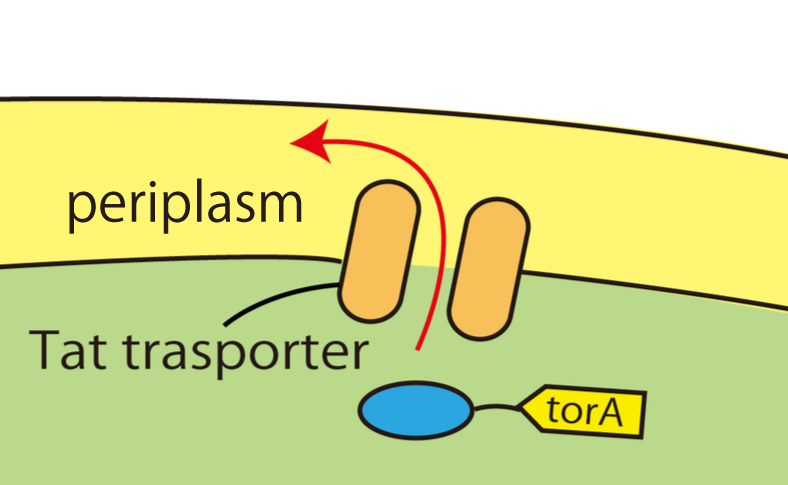
Wild E.coli have many secretion systems. In order to enable proteins to go through inner membrane, we decided to use one of these systems, called Twin Arginate Translocation (Tat) pathway. Tat pathway is more suitable than other secretion systems for our project, because by Tat pathway E.coli secrete proteins into the periplasm keeping proteins’ conformation and function. The following is the mechanism of Tat pathway; a Tat transporter recognizes TorA signal, and proteins having a TorA signal at their N terminals only can get into the periplasm.
We improved usability of TorA signal BioBrick
We tried to combine TorA signal with FT protein. Searching previous iGEM projects, we actually found TorA signals as iGEM parts (such as [http://partsregistry.org/Part:BBa_K638402 BBa_K638402]) submitted by other teams. These parts, however, have two big problems. One problem is that these parts do not have RBS. So iGEMers who use the parts have to spend additional processing time. The other problem is that stop codon appear between signal region and target coding sequence when these parts are combined with some other parts by standard or 3A assembly. In short, when iGEMers use these parts, TorA-fusion protein would not be expressed at all, or only TorA would be expressed.
To solve these problems, we produced new applicable TorA signal [http://partsregistry.org/Part:BBa_K797002 BBa_K797002] (Fig.2-2). Our part has two improvements. [http://partsregistry.org/Part:BBa_K797002 BBa_K797002] contains RBS and indels to prevent the emergence of stop codon between signal region and target cording sequence. We sequenced [http://partsregistry.org/Part:BBa_K797002 BBa_K797002] and confirmed that stop codon did not appear when we used it in Standard and 3A assembly. Using green fluorescent protein (GFP) as a target protein, we constructed plasmid of plac-RBS-TorA-GFP-DT and introduced it into E.coli. After that, we observed the TorA-GFP-fusion-expressing cells (Fig.2-3) suggesting that the TorA-GFP fusion was successfully expressed. This meant RBS in our TorA signal worked well and stop codon didn't appear after assembly.
Kil protein enables proteins to go through outer membrane
With Tat secretion pathway, FT protein can be transported into the periplasm. Next, FT protein needs to move to the outside of E.coli. For secretion, we used kil protein, which is derived from λ phage. Kil protein makes holes in the outer membrane of E.coli. So in our project, we introduced kil gene into E.coli. But the function of outer membrane is essential for E.coli to survive. Overexpression of kil gene, therefore, causes cell death. For this reason, we must check whether kil gene is harmful or not under our condition.
Kil protein had no significant effect on E.coli's growth
We made the construction plac-RBS-kil-double terminator, whose backbone is pSB3C5. After culturing for 18hr at 37℃, we removed the supernatant, and diluted it to OD600=0.1. Then we resuspend it. And then, we added 0/0.001/0.01/0.1/1mM IPTG to each. While culturing again at 37℃, we measured OD600, which indicate the density of E.coli. The figure below shows the results. These results indicated that the expression of plac-RBS-kil-DT (pSB3C5) makes no effect on the survival of E.coli.
We established the new biobrick for the useful secretion system. When we apply these systems, torA signal and kil protein, to Flower Fairy E.coli, we can make E.coli secrete FT protein without cell lysis!!(Fig.2-7)
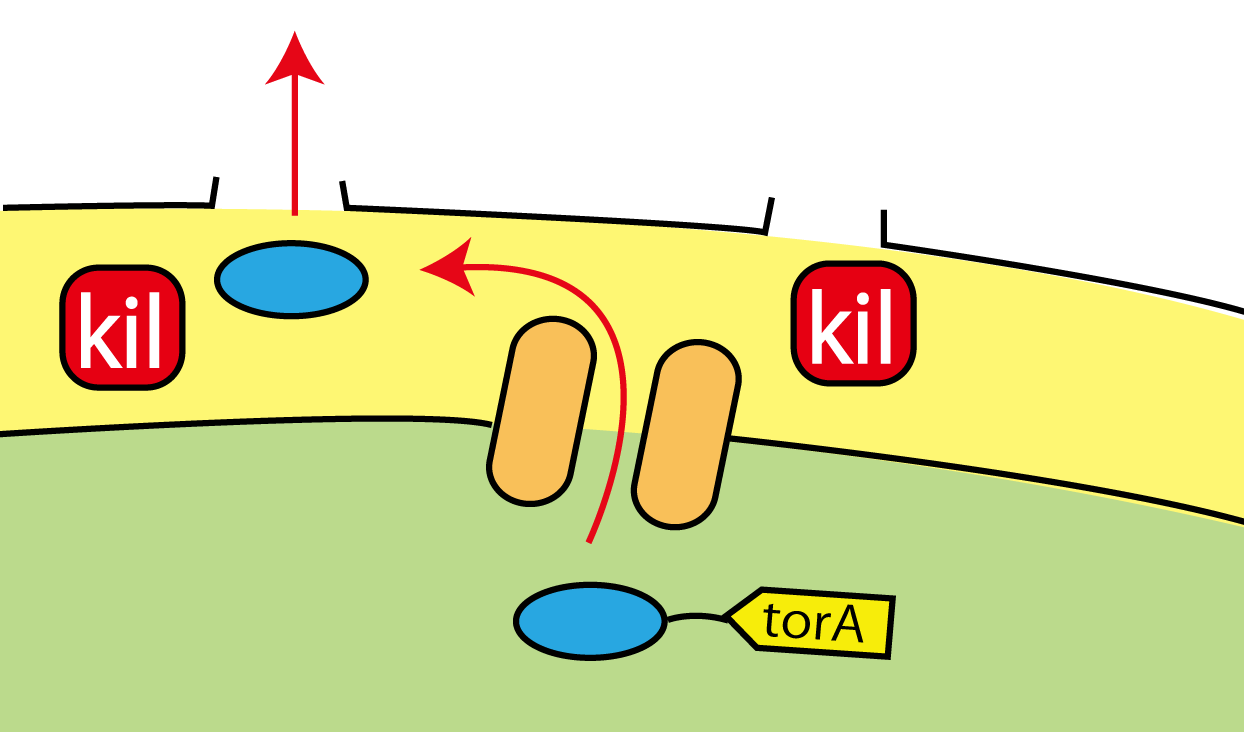
1.Tat transporters are piercing in the inner membrane. They recognize TorA signal, and transport into the periplasm only proteins that have TorA signal at their N terminals. 2. Kil protein makes holes in the outer membrane of E.coli and the proteins go through the outer membrane.
Improvement of secretion system
Now we can let E.coli to secrete FT protein to the outside by using Tat pathway and kil protein. But E.coli secrete only a small amount of FT protein when they use Tat transporters which they inherently have. To make E.coli secrete enough amount of FT protein, we needed to improve the efficiency of their secretion systems. To reach this goal, we used two genes. One is composed of TatA, TatB, and TatC, which composes Tat transporter. Another is phage-shock protein A (pspA), which wild E.coli have. When their inner membrane is damaged, pspA gene is expressed ,and pspA maintains membrane potential and H+ concentration gradient between the periplasm and cytoplasm. It is known that pspA promote trasport of proteins to the periplasm through the detail mechanisms are unknown. As the next step of secretion, by the extra induction of these genes, we tried to increase the amount of secresion. We constructed Tat secretion cassette with constitutive promoter (BBa_K797004).(Fig.2-8) This part includes TatA, B and C proteins coding region and pspA. By using this part, the amount of Tat transporter is and pspA can be increased . In short, we can make E.coli secrete more proteins with TorA.(Fig.2-9)
Kyoto 2012 suggests this new way of secretion, Tat pathway and kil protein, and provides iGEMers with these genes regulated by constitutive promoter. We checked the sequence of TatABC [http://partsregistry.org/Part:BBa_K797000 (BBa_K797000)] and the sequence of pspA [http://partsregistry.org/Part:BBa_K797001 (BBa_K797001)] individually, and then, we made Tat construction composed of constitutive promoter [http://partsregistry.org/Part:BBa_J23107 (BBa_J23107)], TatABC [http://partsregistry.org/Part:BBa_K797000 (BBa_K797000)], pspA [http://partsregistry.org/Part:BBa_K797001 (BBa_K797001)] and double terminator [http://partsregistry.org/Part:BBa_B0015 (BBa_B0015)]. We performed electrophoresis of this cassette confirmed the length of our parts and sequenced them partially.
3.PENETRATION

The third step is PENETRATION. In order to induce flower formation, FT protein from E.coli must enter into plant cells. This is because FT protein upregulates other proteins which lead to flower formation in plant cells.
However, normally proteins cannot penetrate a cell membrane of a plant. Therefore, we needed a method to send FT protein into plant cells.
Thanks to the advice from Doctor Washida, we found a method for the penetration of cell membrane. In this method, we use polyarginines called R9 peptides.
R9 peptide enables FT protein penetrate membranes by endocytosis
R9 peptide consists of nine arginine residues(Fig.3-1). It is known as a kind of CPP (Cell Penetrating Peptide). Arginine-rich peptides induce macropinocytos, a type of endocytosis. From this, we judged that R9 peptides were suitable for cell membrane penetration.
This is the mechanism of how R9 peptides work(Fig3-2).
Firstly, R9 peptides adhere to a cell membrane of a plant because of their hydrophobic character.
Secondly, the cell responds to this stimulus and starts to endocytose.
Finally, proteins around the endocytosing region of the cell are taken into the cell.
R9 peptides seem to work effectively whether or not R9 peptides are fused with target proteins.
However, there are few examples about endocytosis of plants, so we needed to check the function of R9 when used on plant cells.
Given the R9 system, it induces endocytosis whether R9 peptides and target proteins are fused or not.
Considering this system, target proteins near the R9 peptides are taken into cells by endocytosis.
So, we assumed that the system would induce endocytosis in both cases where R9 peptides and target proteins were fused and not fused.
However, as we wanted to introduce proteins as efficiently as possible, we deliberated whether we should fuse R9 and target proteins.
We thought the shorter the distance between R9 peptides and target proteins is, the more easily they are taken into a plant cell. Therefore, we expected that fusing R9 peptide and a target protein would lead to higher efficiency.
So, we tried to fuse R9 peptides and target proteins to increase penetration efficiency
We transformed E.coli, fusing R9 pepetides and target protein's gene in line. By doing this, we tried to make E.coli express R9::GFP fusion protein.
In order to visualize the function of R9, we tried to prepare R9::GFP fusion protein. However, E.coli expressing the R9::GFP fusion protein was not effectively cultivated.(Fig3-3). Then, we checked whether R9 peptides had a bad effect on the expression of R9::GFP fusion protein by Western blotting and RT-PCR. Although we succeeded in confirming the existence of the mRNA (Fig3-4), we did not find the proteins (Fig3-5). These results are probably caused by poor translation or quick breakdown of the protein.
We used the existing GFP generator part, [http://partsregistry.org/Part:BBa_I746915 BBa_I746915].
Fig.3-4 Western blotting for checking the expression. No band of R9::GFP fusion protein. |
We used R9 and GFP separately and to check the R9 peptides function
It was a question whether R9 peptides work properly and introduce FT protein into plant cells. Although endocytosis is often observed in animal cells, there are few examples of plant cell’s endocytosis. We, therefore, needed to verify the function of R9 peptides. For this reason, we performed the following experiment(Fig3-6).
First, we removed the cuticles from Arabidopsis thaliana's leaves by scratching them. Second, we devided the leaves into two groups, and soaked one into a solution of only GFP, and the other into that of GFP and R9. After 5 minutes we washed cells by PBS in order to wash away GFP and R9 peptides from around the leaves. After that, we contrasted leaves having soaked into the only GFP solution with ones having soaked into the solution of R9 and GFP mixture. Then we succeeded in getting the figure of GFP fluorescence (Fig3-7).
The control groups on the left were soaked in only GFP, and the experimental group on the right were soaked in GFP and R9. These fluorescence indicated that R9 peptides properly kept GFP in or on the plant cells. This figure strongly suggests that R9 peptides work successfully and GFP penetrate cell membrane, because this was taken by a confocal microscopy and seen as cross sections.

We can make FT protein penetrate cell membrane of plants by R9 peptide function!.
4. Activation

The final step is; Activation. We verified whether FT protein made by E.coli work normally in plant cells.
Actually, FT protein is a transcriptional factor and we can check the ability of FT protein by observing the transcription levels of genes up-regulated by FT protein.
The detail function of FT
FT protein increases transcript activities of flowering factors which lead to flower formation in a plant cell. Let us introduce the function mechanism of FT protein. Actually, FT function is to up-regulate genes relating to bloom flowers. at the stage of flowering of plants, the expressions are incresed in these genes, FUL, SEP3, and so on. These genes work on shoot apex and change the form of shoot apex in order to start flowering. We focused on the amount of FUL and SEP3 because they are the most popular genes among them.
Injecting FT and verifying its function by RT-PCR
To evaluate the function of FT produced by our E.coli, we tried to check the amount of flowering factors' mRNA upregulated by FT. Experimental procedure is as follows.
1. We prepared solutions. One is composed of R9 peptides and GFP, and the other is composed of R9 peptides and FT. GFP protein is used as a control group. 2. We injected these solutions into each leaf by a syringe. 3.After incubation, we tried to perform RT-PCR in order to check up-regulation of flowering factors.
We used leaf cells of Arabidopsis thaliana, insted of cells of a shoot apex. This is because shoot apex' cells are too small to observe. FT protein upregulates FUL and SEP3 genes in leaf cells, too. GFP is suitable for control experiments because GFP's molecular weight(27kDa) is relatively similar to that of FT(20kDa.)

30mg of leaves of Arabidopsis thaliana before bolting were used for one sample. FT or GFP protein and R9 peptide were diluted in PBS (pH7.4), 50ug/L and 500ug/uL each. Leaves were soaked into FT-R9 or GFP-R9 solution for 5min. and incubated for 16hr. in PBS(pH7.4.) After incubation, leaves were freezed with liquid nitrogen and glinded immediately.
Total RNA was extracted by phenol-chloroform extraction. cDNA was synthesized by reverse transcription and used as templates of RT-PCR. TUBULIN was used for internal control of mRNA expression.
Lane1: TUBULIN (GFP-R9 treated) amplicon 61bp
Lane2: TUBULIN (FT-R9 treated)
Lane3: FUL (GFP-R9 treated) amplicon 132bp
Lane4: FUL (FT-R9 treated)
Lane5: SEP3 (GFP-R9 treated) amplicon 87bp
Lane6: SEP3 (FT-R9 treated)
Lane7: AP1 (GFP-R9 treated) amplicon 958bp
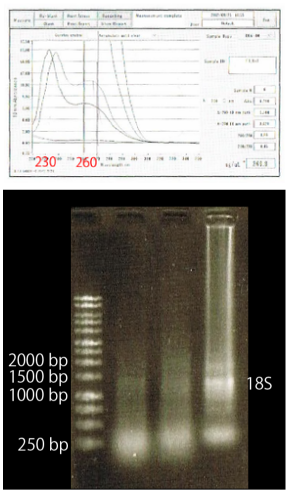
As shown above, we got no correct ampicon band, even from tubulin, which is a high expressed gene we used as internal control. One possible reason of this failure was the poor quality of extracted RNA in this experiment. To check this, we compared the total RNA by electrophoresis, shown in fig.4-3.
As shown in the Fig.4-3, this time, RNA samples were degradated.
To add to this, the waveforms of them had a law peak at 260nm(Fig.4-4.)
Establishing an effective way of RNA purification
We found that RNA purification method needed to be improved in ordrer to perform RT-PCR successfully .
We improved following things;
1. Total leaf volume was increased.
2. Samples ware freezed with liquid nitrogen and suspended in ISOGEN more rapidly.
3. We centrifuged samples and collected supernatant twice after adding ISOGEN.
After the improvement, 260nm peak became higher and the degradation is minimized(Fig.4-4A.)
Then we used better quality of RNA for reverse transcription and retried RT-PCR.
Fig.4-4C shows the result of this RT-PCR.

A: the waveform of RNA purified in the previous way. it indicated that RNA purity is low
B: Improved RNA waveform
C: Electrophoresis of RNA. RNA concentration is adjusted. These band showed all sample were purified correctly D: Lane1: 100bp ladder Lane2: TUBULIN Lane3: FUL Lane4: SEP3 Lane5: AP1
Conducting RT-PCR and qPCR again by using new RNA purification
To confirm the exact differnece between experimental group(FT+) and negative control(FT-), we conducted qPCR and compared the relative RNA expression of each gene. However, there was not significant difference between experimental group and negative control.
Checking FT protein purification by Western Blotting
We could not confirm FT function though enough quality of RNA was extracted. Therefore, we questioned about FT protein quality, and performed western blotting to confirm FT protein is successfully purified.
Unfortunately, the band of FT was not detected. Because of the time shortage, we could not conduct further experiment. We have to reconsider about FT purification and now we are trying.
Achievement
1.Expression
- Mutate FT sequence
- Standardize FT as an iGEM part
- Confirm expression of FT protein in E.coli
2.Secretion
- Modify TorA signal to be easy to use the signal more
- Construct Tat secretion cassette that contains Constitutive promoter, RBS, TatABC, pspA, double terminator
- Standardize kil gene
- Multiply TatABC in order to strengthen Tat secretion system--not yet
3.Penetration
- Keep GFP in or around plant cells using R9 peptide
- Introduce FT in plant cells using R9 peptide--not yet
4.Activation
- Get high quality of RNA
- Amplify genes successfully
- Check the function of FT--not yet
At PENETRATION step, GFP::R9 fusion protein connected at GFP’s N-terminal was not expressed. However, if GFP is tagged with R9 at C-terminal, that fusion protein may be successfully expressed. This is because there is an example where proteins tagged with arginine at C- terminals were correctly expressed whereas proteins tagged at N-terminals were not([http://partsregistry.org/wiki/index.php/Part:BBa_K249005 BBa_K249005] ). By using GFP::R9 fusion protein connected at GFP's C-terminal, we might improve the efficiency of macropinocytos, a type of endocytosis.
At Activation, in order to see whether FT is usable or not, we measured the relative RNA expression of FUL and SEP3 by qPCR. However, there was not any significant difference between experimental group and negative control.
We found that FT was not successfully purified and there is the room for improvement in that.
In addition of this, several reasons can be considered.
The cause of failure may be that FT protein was not active or decomposed in the cell or even that FT protein didn't enter plant cells.
If we perform Western blotting of FT in plant cell after putting FT into it with R9, we will be able to understand whether the problem was FT protein or R9 peptides.
If the cause was that our FT protein was not active though FT had entered the cell, FT will be confirmed. In that case, we have to consider the possibilities that FT needs Post-translational modification or Arabidopsis thaliana that we used was old.
When FT needs Post-translational modification, we have to do more research about difference of Post-translational modification between E.coli and Arabidopsis thaliana.
When Arabidopsis thaliana that we used was old, it might have originally expressed enough FT, so our FT might not be necessary to induce FUL and SEP3.
If the cause was that FT didn’t enter plant cell or FT was decomposed in the cell, FT will not be confirmed. When FT didn’t enter plant cell, we have to reconstruct experimental system, for example, by using other kinds of R9 since there exist more effective R9s. Wnen FT was decomposed in the cell, we have to investigate the mechanism of decomposing protein.
Future Works
We noticed only flowering and florigen in this time but there are many other plant hormones. We made translocation pathway from E.coli into plant cells, so we will be able to introduce plant hormones into plant cells if E.coli can make them. It means we can control plant growth in any stage through genetically engineered E.coli. In the future that is not so far, we will be able to meddle in plants' growth――germinating, elongation, flowering, and fructification. We human will finally accomplish a technology that control plants perfectly.
Moreover, R9 peptide functions not only plant cell. R9 peptide works on animal cell similarly. It means that we found a pathway into any kinds of cells. R9 peptide tag enables us to introduce proteins into any cells, so we will be able to control all living cells using this technology.
Biosafety
We cooperated with KAIT-Japan and the mark on the left indicates Biosafety Level of our parts.
[1][http://www.ncbi.nlm.nih.gov/pubmed/15695452 Microsugar Chang et al.(2005) "Cellular internalization of fluorescent proteins via arginine-rich intracellular delivery peptide in plant cells" Plant Cell Physiol, 46(3), 482–488]
[2][http://www.ncbi.nlm.nih.gov/pubmed/16155177 Paula Teper-Bamnolker and Alon Samach1.(2005) "The flowering integrator FT regulates SEPALLATA3 and FRUITFULL accumulation in Arabidopsis leaves" The Plant Cell, 17, 2661–2675]
[3][http://www.ncbi.nlm.nih.gov/pubmed/16099980 Philip A. Wigge et al.(2005) "Integration of spatial and temporal information during floral induction in Arabidopsis" Science, 309(5737), 1056-1059]
[4][http://www.mdpi.com/1424-8247/3/4/961/htm Sara Trabulo et al.(2010) "Cell-penetrating peptides—mechanisms of cellular uptake and generation of delivery systems" Pharmaceuticals, 3, 961-993]
[5][http://www.ncbi.nlm.nih.gov/pubmed/15147914 Unnamalai N, Kang BG, Lee.(2004) "Cationic oligopeptide-mediated delivery of dsRNA for post-transcriptional gene silencing in plant cells" FEBS Lett 21, 566(1-3), 307-10]
[6][http://www.ncbi.nlm.nih.gov/pubmed/22683878 Tracy Palmer and Ben C. Berks.(2012) "The twin-arginine translocation (Tat) protein export pathway" Nat Rev Microbiol, 10(7), 483-96]
[7][http://www.ncbi.nlm.nih.gov/pubmed/14966662 Choi JH, Lee SY.(2004) "Secretory and extracellular production of recombinant proteins using Escherichia coli" Appl Microbiol Biotechnol, 64(5), 625-35]
[8][http://www.ncbi.nlm.nih.gov/pubmed/9042754 Miksch G, Fiedler E, Dobrowolski P, Friehs K.(1997) "The kil gene of the ColE1 plasmid of Escherichia coli controlled by a growth-phase-dependent promoter mediates the secretion of a heterologous periplasmic protein during the stationary phase" Arch Microbiol, 167(2-3), 143-50]
[9][http://www.ncbi.nlm.nih.gov/pubmed/11854367 Seibel BA, Walsh PJ.(2002) "Trimethylamine oxide accumulation in marine animals: relationship to acylglycerol storage" J Exp Biol, 205(Pt 3), 297-306]
[10][http://www.ncbi.nlm.nih.gov/pubmed/11123687 Thomas JD, Daniel RA, Errington J, Robinson C.(2001) "Export of active green fluorescent protein to the periplasm by the twin-arginine translocase (Tat) pathway in Escherichia coli" Mol Microbiol, 39(1), 47-53]
[11][http://www.ncbi.nlm.nih.gov/pubmed/3139642 Suit JL, Luria SE.(1988) "Expression of the kil gene of the ColE1 plasmid in Escherichia coli Kilr mutants causes release of periplasmic enzymes and of colicin without cell death" J Bacteriol, 170(10), 4963-4966]
[12][http://www.ncbi.nlm.nih.gov/pubmed/14702305 DeLisa MP, Lee P, Palmer T, Georgiou G.(2004) "Phage shock protein PspA of Escherichia coli relieves saturation of protein export via the Tat pathway" J Bacteriol, 186(2), 366-373]
[13][http://www.ncbi.nlm.nih.gov/pubmed/16099979 Araki, T et al.(2005) “FD, a bZIP protein mediating signals from the floral pathway integrator FT at the shoot apex” Science 309(5737), 1052–1056]
BioBricks are useful for us because we can look for required BioBrick parts from their registry and recombine genes easily. When we want to introduce many parts into one plasmid, however, we have to repeat the process; restrict enzyme digestion and ligation. It takes us too much time and sometimes we lose time for other experiments.
We want to reduce the time required for the recombination of genes and get time for verification of the expression and the effect of genes.
Golden Gate assembly is one of the ways to make it possible.
Some teams like 2011 WHU-China have used this assembly. But they didn't seem to spread Golden Gate Assembly through other iGEM teams. So, we created plasmid backbone parts [http://partsregistry.org/Part:BBa_K797013 "BBa_K797013"] to make it easier to use Golden Gate assembly.
We also created software which designs primers for Golden Gate assembly.
What's Golden Gate Assembly
Golden Gate Assembly is developed by Carola Engler, Ramona Gruetzner, Romy Kandzia and Sylvestre Marillonnet.[2]
This method enables us to introduce plural gene segments into one plasmid all at once.
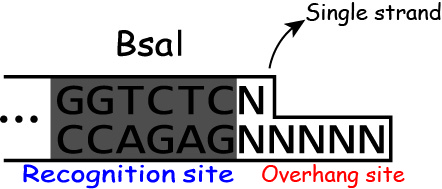
Golden Gate Assembly takes advantage of the characteristic restrict enzyme "BsaI".
Most restrict enzyme cuts its own recognition sites.
But BsaI recognizes the sequence "GGTCTC"(Figure 1) and cuts downstream of the recognition site as shown in the Figure 2.
And BsaI activity is independent of the sequences of the downstream of the
recognition site.
Restrict enzyme digestion and ligation are completed by just one PCR because once DNA is cut and ligated irreversibly, the recognition site of BsaI disappears. We create the segments which have complementary ligation sites so that we can introduce plural DNA segments into one plasmid at the same time and arrange them in the way we like. (Figure 4) Learn more about [http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0005553 Golden Gate Assembly].
Plasmid backbone [http://partsregistry.org/Part:BBa_K797013 BBa_K797013]
Before we started assembly, we had to make proper gene segments for Golden Gate assembly. The DNA segments we created had four bp for ligation, one base pair spacer, BsaI recognition sites and four base pair at the both ends.
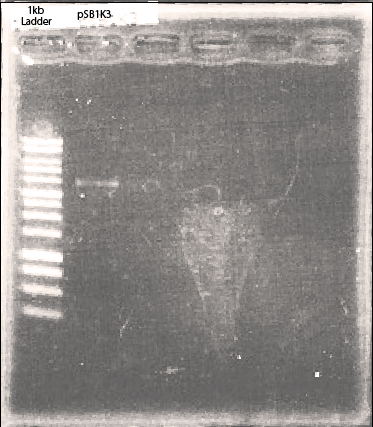
We amplified plasmid backbone(psB1K3).
After amplification, we assembled these DNA segments referring to the protocol.(See the part of Golden Gate Assembly)
After we confirmed this part has the restriction cites and restriction enzyme cutting sites of BsaI. After this, we mixed DpnI with enzyme-treated plasmid and conducted ligation. Other iGEM teams can use this backbone plasmid for their Golden Gate assembly.
Golden Pass
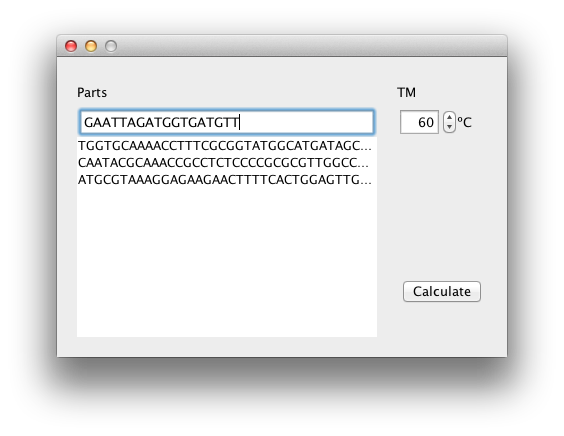
We created software which design primers to create DNA segments for Golden Gate assembly. This software gives you the sequences of primers when you input the sequences of the parts and melting temperature.
How to use
- input the sequences of the parts starting from 5' terminal
- set melting temperature
- click the "calculate"
[http://igemkyotogoldenpass.appspot.com/ Use the web version of GoldenPass...]
We created the plasmid backbone [http://partsregistry.org/Part:BBa_K797013 BBa_K797013] and software to design primers for Golden Gate assembly. They make it easier to use Golden Gate assembly. We want other teams to use this parts and software to use precious time efficiently.
[1][http://www.ncbi.nlm.nih.gov/pubmed/18985154 Carola Engler, Romy Kandzia, Sylvestre Marillonnet.(2008) "A One Pot, One Step, Precision Cloning Method with High Throughput Capability" PLoS ONE 3(11), e3647.]
[2][http://www.ncbi.nlm.nih.gov/pubmed/19436741 Carola Engler, Ramona Gruetzner, Romy Kandzia, Sylvestre Marillonnet.(2009) "Golden Gate Shuffling: A One-Pot DNA Shuffling Method Based on Type IIs Restriction Enzymes" PLoS ONE 4(5), e5553. doi:10.1371/journal.pone.0005553 doi:10.1371/journal.pone.0003647]
 "
"