Team:TU Munich/Notebook/Labjournal
From 2012.igem.org
VolkerMorath (Talk | contribs) (→2 Quick Change mutagenis to remove SpeI from pYES2_RFC25 MCS 1.2) |
VolkerMorath (Talk | contribs) (→2 Quick Change mutagenis to remove SpeI from pYES2_RFC25 MCS 1.2) |
||
| Line 267: | Line 267: | ||
''' Transformation into ''E.coli'' Xl1-Blue''' | ''' Transformation into ''E.coli'' Xl1-Blue''' | ||
'''Operation Sequence''' | '''Operation Sequence''' | ||
| + | * This operation sequence was applied to the PCR prducts of P33, P13 and P14 respectively. | ||
* melting of 100 µl Ca-competent ''E.coli'' XL1-Blue cells | * melting of 100 µl Ca-competent ''E.coli'' XL1-Blue cells | ||
| - | * addition of 1 µl of the | + | * addition of 1 µl of the PCR product |
* incubation for 30 min on ice | * incubation for 30 min on ice | ||
* heat shock for 5 min at 37 °C | * heat shock for 5 min at 37 °C | ||
Revision as of 11:58, 29 June 2012
Contents |
Monday, 18th June
XXXX
Investigator: Daniela, Saskia
Tuesday, 19th June
XXXX
Investigator: Daniela, Saskia
Wednesday, 20th June
XXXX
Investigator: Daniela, Saskia
Thursday, 21th June
XXX
Investigator: Daniela, Saskia
Friday, 22th June
1 Transformation of E.coli XL1-Blue with pKS2µHyg-PAL-4Cl-CHS
Investigator: Ingmar, Volker
Aim of the experiment: Plasmid amplification
Operation Sequence:
- melting of 100 µl Ca-competent E.coli XL1-Blue cells
- addition of 1 µl of the Plasmid pKS2µHyg-PAL-4Cl-CHS
- incubation for 30 min on ice
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubate at 37°C and 180 rpm for 30 min
- plate 100 µl on an Amp-LB-plate
- sediment the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspend the sediment in 100 µl LB-medium and plate it as well on an Amp-LB-plate
Saturday, 23th June
2 Quick Change mutagenis to remove NgoMIV from pYES2
Investigator: Ingmar, Volker
Aim of the experiment: Generation of an RFC 25 compatible version of the pYes2 Vector.
PCR
Reaction batch
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 4 µl | Plasmid P7 pYes2_RFC25 MCS 1.1 template |
| 0.5 µl | 1:10 dilution of O38 (10 pmol/µL) |
| 0.5 µl | 1:10 dilution of O39 ((10 pmol/µL) |
| 17 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Turbo DNA polymerase (2.5 U / µl) |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 30 sec |
| 2 | 15 | 95°C | 30 sec |
| 55°C | 1 min | ||
| 68°C | 6 min |
- The vector resulting from the PCR-product was named pYes2_RFC25 MCS 1.2.
- Digestion of the parental DNA with DpnI: Addition of 1 µl DpnI to the PCR batch and incubate for 1 h at 37 °C.
Transformation into E.coli Xl1-Blue Operation Sequence
- melting of 100 µl Ca-competent E.coli XL1-Blue cells
- addition of 1 µl of the Plasmid P7 pYes2_RFC25 MCS 1.2
- incubation for 30 min on ice
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubate at 37°C and 180 rpm for 30 min
- plate 100 µl on an Amp-LB-plate
- sediment the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspend the sediment in 100 µl LB-medium and plate it as well on an Amp-LB-plate
Sunday, 24th June
1 Miniprep of E.coli XL1-Blue with pKS2µHyg-PAL-4Cl-CHS
Investigator: Ingmar, Volker
Aim of the experiment: Plasmid purification
Operation Sequence:
- A single clone of E.coli XL1-Blue with pKS2µHyg-PAL-4Cl-CHS was picked an transferred to 5 ml LB Amp on saturday evening. Incubation overnight at 37°C 180 rpm.
- Using a Quiagen kit a miniprep of the overnight culture was done.
2 Quick Change mutagenis to remove NgoMIV from pYES2
Investigator: Ingmar, Volker
Aim of the experiment: Removal of a NgoMIV restriction site in the backbone of pYes2.
Operational sequence:
- A single clone of E. coli pYes2_RFC25 MCS 1.2 was picked an transferred to 5 ml LB Amp. Incubation overnight at 37°C 180 rpm.
3 Transformation of 2 Biobricks into E. coli XL1-Blue
Investigator: Jeffery Truong
Aim of the experiment: Transformation of Biobricks into E. coli for plasmid propagation for PCR with new RFC pre- and suffix primer in order to do protein fusions.
- 2 Biobricks from the distribution kit were used: First, LexA (BBa_K105005, Plate 3 Well 9E) in the pSB1A2 plasmid with ampicillin resistance and second, the heme oxygenase (BBa_I15008, Plate 2 Well 13J) in the pSB2K3 plasmid with kanamycin resistance.
- 10 µL of autoclaved H2O were added to each well on the distribution kit. The well immediately turned red which means that one does it right.
- The now resuspended DNA liquids were transferred into a new ERG on ice.
- CaCL2 competent E. coli XL1-Blue cells from the stock were gently defrezed on ice.
- For each Biobrick 100 µL cells were used and pooled together with 2 µL of plasmid DNA in a ERG on ice.
- Incubation on ice for 30 min.
- 5 min heat shock at 37 °C.
- Each ERG now is transferred in a new ERG prefilled with 1 mL of LB-medium and incubated in a cell-culture shaker at 37 °C for 45 min.
- 100 µL of these cell suspension were plated on antibiotic selection plates (Ampicillin for LexA and Kanamycin for heme oxygenase).
- The rest of the cell suspension is centrifuged at 13000 rpm for 60 s and the supernatant is discarded.
- The pellet is resuspended with 100 µL for each ERG and is plated on another antibiotic selection plate
- These 4 plates were put at 37 °C overnight
Monday, 25th June
1 Miniprep of E. coli XL1-Blue with pYes2_RFC25 MCS 1.2
Investigator: Alois, Martin
Aim of the experiment: proof of successful removal of NgoMIV in the backbone of pYes2
Operation Sequence:
- Mini prep of pYes2 1.2. The resulting purified DNA is P33.
- Control digest of pYes2_RFC25 MCS 1.2 and p13 (+ analytical gel electrophoresis: 90 V, 1 h:
* 15 µl ddH20 * 2 µl NEBuffer4 * 0,5 µl NgoMIV * 2,5 µl pYes2 1.2/p13 * 37°C, 1 h.
2 Quick Change mutagenis to remove SpeI from pYES2_RFC25 MCS 1.2
Investigator: Ingmar, Volker
Aim of the experiment: Generation of an RFC 25 compatible version of the pYes2 Vector.
PCR
Reaction batch
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 4 µl | Plasmid P33 template |
| 0.5 µl | 1:10 dilution of O44 (10 pmol/µL) |
| 0.5 µl | 1:10 dilution of O45 ((10 pmol/µL) |
| 17 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Turbo DNA polymerase (2.5 U / µl) |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 30 sec |
| 2 | 15 | 95°C | 30 sec |
| 55°C | 1 min | ||
| 68°C | 6 min |
- The procedure was furthermore applied to P13 and P14.
- The vector resulting from the PCR-product was named pYes2_RFC25 MCS 1.3.
- Digestion of the parental DNA with DpnI: Addition of 1 µl DpnI to the PCR batch and incubate for 1 h at 37 °C.
Transformation into E.coli Xl1-Blue Operation Sequence
- This operation sequence was applied to the PCR prducts of P33, P13 and P14 respectively.
- melting of 100 µl Ca-competent E.coli XL1-Blue cells
- addition of 1 µl of the PCR product
- incubation for 30 min on ice
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubate at 37°C and 180 rpm for 30 min
- plate 100 µl on an Amp-LB-plate
- sediment the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspend the sediment in 100 µl LB-medium and plate it as well on an Amp-LB-plate
Picking of E. coli cells on antibiotic selection plates: pSB1A2 plasmid with BBa_K105005 (LexA) and pSB2K3 plasmid BBa_I15008 (heme oxygenase)
Investigator: Jeffery Truong
Aim of the experiment: Picking colonies from transformed E. coli XL1-Blue, 4x picked for each Biobrick.
- pSB1A2 plasmid with BBa_K105005 (LexA): Colonies were on both ampicillin selection plates, the one with diluted cell suspension and the one with concentrated E. coli cell suspension. Typical E. coli colony morphology. Picking was performed on the plate with diluted cell suspension.
- pSB2K3 plasmid BBa_I15008 (heme oxygenase): Colonies were only on the kanamycin selection plate with concentrated cell suspension. The one with diluted susepension was empty. Typical but very small E. coli colonies. Picking was performed from the first plate.
- Picked pipette tips was transferred into a special cell-culture tubes with air-permeable, but sterile cover. In each tube 4 mL of LB-medium + ampicillin (???)(for pSB1A2) or kanamycin (35 mg/mL) (for pSB2K3).
- 4 colonies for each Biobrick was picked; total: 8 tubes overnight culture.
- These tubes were transferred in a cell culture shaker at 37 °C and were incubated overnight
Tuesday, 26th June
Quick Change mutagenis to SpeI from pYes2_RFC25 MCS 1.2
Investigator: Ingmar, Volker
Aim of the experiment: Removal of a SpeI restriction site in the backbone of pYes2.
Operational sequence:
- A single clone of E. coli pYes2_RFC25 MCS 1.3 was picked an transferred to 6 ml LB Amp. Incubation overday at 37°C 180 rpm.
- Using a Quiagen kit a miniprep of the overnight culture was done.
PCR of PAL, 4CL, CHS, OMT (Coumaryl-CoA)
Investigator: Daniela, Mary
Determination of concentration of plasmids (Nanodrop): c(pKS2µHyg-PAL-4CL-CHS) = 500 ng/µl c (pOMT) = 20 ng/µl
PCR:
| Name of tube | Enzyme | consensus (+)/ consensless (-) | used Oligos |
| CHS - | CHS | - | O13 and O24 |
| CHS + | CHS | + | O23 and O24 |
| PAL - | PAL | - | O15 and O16 |
| PAL + | PAL | + | O22 and O16 |
| OMT - | OMT | - | O17 and O26 |
| OMT + | OMT | + | O25 and O26 |
| 4CL - | 4CL | - | O19 and O20 |
| 4CL + | 4CL | + | O21 and O20 |
Reaction batch
| volume | reagent |
| 5 µl | 10x Pfu Ultra II buffer |
| 4 µl | dNTP's (each 2.5 mM) |
| 0.5 µl | Pfu Ultra II (2.5 U/µL) |
| 5 µl | 1:10 dilution of used forward primers (10µM) |
| 5 µl | 1:10 dilution of used reversed primers (10µM) |
| 1 µl | DNA (pKS2µHyg-PAL-4CL-CHS 50 ng/µL or pOMT 20 ng/µL) |
| 29.5 µL | bidest. sterile Water |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 5 min (and adding Pfu Ultra after 3 min) |
| 2 | 30 | 95°C | 30 sec |
| 46°C | 2.5 min | ||
| 72°C | 1.5 min | ||
| 3 | 72°C | 5 min |
PCR purification
- Purification was done using QIAquick PCR Purification Kit (250)
Analytical Gel Electrophoresis:
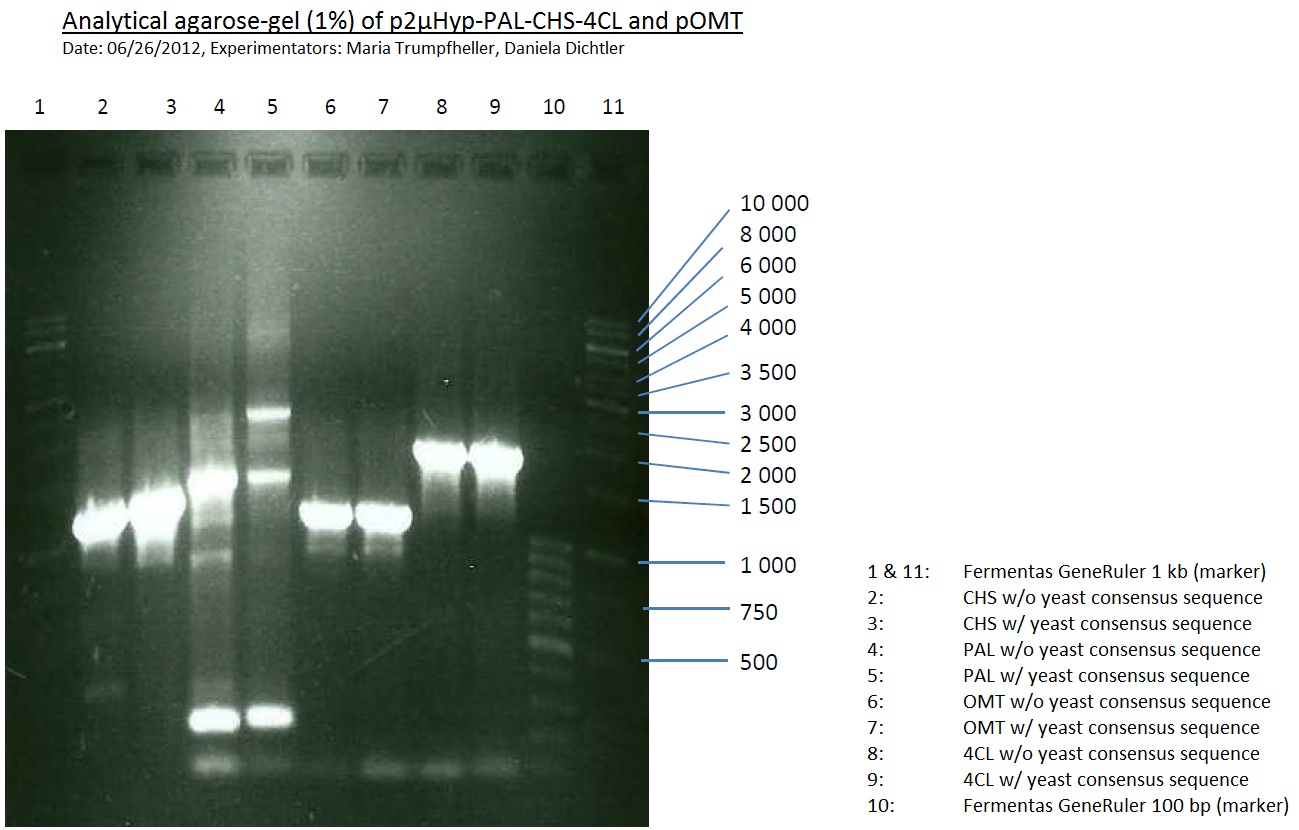
-> going on with CHS, 4CL and OMT; the PCR of PAL will be repeated with an temperature-gradient
Miniprep and analytical gel of picked transformed overnight culture with pSB1A2 plasmid with BBa_K105005 (LexA) pSB2K3 plasmid BBa_I15008 (heme oxygenase)
Investigator: Jeffery Truong, Georg Schützinger
Aim of the experiment: Plasmid isolation from the picked transformed overnight E. coli cells with pSB1A2 plasmid with BBa_K105005 (LexA) pSB2K3 plasmid BBa_I15008 (heme oxygenase).
- The LB-medium with antibiotics of every tube was opaque which means that the picked cells were successfully inoculated.
- Centrifugation step at 5000 rpm for 10 min at 16 °C.
- Every single step now was performed on ice.
- Miniprep (Qiagen Qiaprep spin) after manufacturer's protocoll.
- Analytical restriction master mix was prepared after following scheme (Using XbaI and PstI):
- 4.4 µL XbaI
- 4.4 µL PstI
- 17.6 µL Tango-buffer (10x)
- 132 µL ELGA H2O
- 17.5 µL from the master mix was poooled together with 2.5 µL of plasmid DNA. That corresponds to 2.5&µL of plasmid DNA, 0.25 µL XbaI, 0.25 µL PstI, 2 µL Tango-buffer (10x), 15 µL ELGA H2O.
- Incubation at 37 °C for 120 min on a ERG heating unit.
- BUT: error was performed during preparing the digested plasmid DNA for analytical gelelectrophoresis in the dilution step. 1:10 dilution of analyctical probe with DNA loading buffer:
- 3.3 µL sample (contains already 1x loading buffer!)
- 0.7 µL loading buffer (?x)
- 6 µL TAE-buffer
- Should have done: 3&µL sample + 1 µL loading buffer (10x) + 6 µL TAE-buffer (1x)
- DNA-ladder preperation: 10 µL ladder stock solution + 10 µL DNA loading buffer + 80 µL TAE-buffer. 10 µL of this solution was pipetted in one gel pocket of the prepared 1% agarose gel including ehtiudiumbromid.
- 20 µL of each samples were also pipetted into the gel pockets.
- The gel pockets were pipetted after following scheme:
| LexA (colony 1) | LexA (colony 2) | LexA (colony 3) | LexA (colony 4) | DNA-ladder | Heme oxygenase (colony 1) | Heme oxygenase (colony 2) | Heme oxygenase (colony 3) | Heme oxygenase (colony 4) |
- Gel electrophoresis at 90 V
- After 20 min the resolution was still poor; 20 min longer.
- Analytical Gel OK:
Friday, 29th June
preparative digest of PCR-products of 4CL, CHS and OMT
investigator: Katrin, Mary
each digestion will dure 2.5h at 37°C
- CHS: digestion with Xba1 and HF-Age1 (both NEB)
| volume | reagent |
| 25µl | PCR-product |
| 5µl | Buffer NEB4 |
| 0.5µl | BSA |
| 1µl | Xba1 (NEB; 20u/µl) |
| 1µl | HF-Age1 (NEB; 20u/µl) |
| 17.5µl | bidest. sterile H2O |
- OMT: digestion with Xba1 and HF-Age1 (both NEB)
| volume | reagent |
| 25µl | PCR-product |
| 5µl | Buffer NEB4 |
| 0.5µl | BSA |
| 1µl | Xba1 (NEB; 20u/µl) |
| 1µl | HF-Age1 (NEB; 20u/µl) |
| 17.5µl | bidest. sterile H2O |
- 4CL: digestion with Xba1 and Pst1 (both Fermentas)
| volume | reagent |
| 25µl | PCR-product |
| 5µl | Buffer Tango |
| 2µl | Xba1 (Fermentas; 10u/µl) |
| 3µl | Pst1 (Fermentas; 10u/µl) |
| 15µl | bidest. sterile H2O |
repetition of PCR of PAL with an temperature-gradient
Reaction batch
| volume | reagent |
| 6 µl | 10x Pfu Ultra II buffer |
| 4.8 µl | dNTP's (each 2.5 mM) |
| 0.6 µl | Pfu Ultra II (2.5 U/µL) |
| 6 µl | 1:10 dilution of used forward primers (10µM) |
| 6 µl | 1:10 dilution of used reversed primers (10µM) |
| 1.2 µl | DNA (pKS2µHyg-PAL-4CL-CHS 50 ng/µL or pOMT 20 ng/µL) |
| 35.4 µL | bidest. sterile Water |
 "
"