Membrane Localization
Reconstructed Vectors
Vectors used in the project for membrane protein expression are modified versions of pRSFDuet-1, pETDuet-1,pACYCDuet-1(NOVAGEN), which are originally regulated by T7 promoter.
Considering our multiple-enzyme system, it is necessary to use those compatible plasmids. However, T7 promoter is hard to be fine-tuned in E. coli. To stress the advantage of the system, we recruited a weaker promoter, araBAD promoter, which could moderately response to a wide range of inducer (L-Arabinose) concentration. So we replaced T7 promoter in original plasmids with araBAD promoter (Fig.1).
All proteins we used are inserted into those three reconstructed vectors and under control of araBAD promoter.
 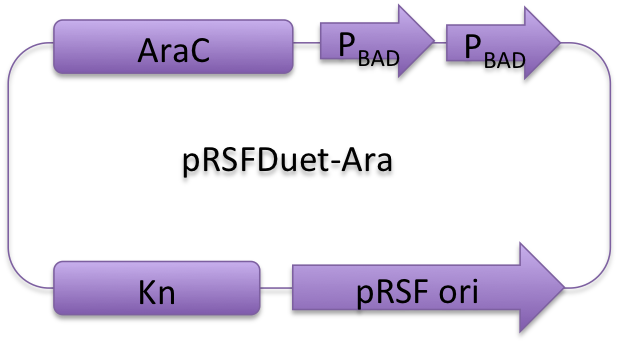
Fig.1 : Profile of reconstructed plasmids in Membrane Magic Project
Membrane Protein Construction
To construct membrane assemblies, we must make sure that our device is directed to inner membrane of E.coli.
The first step is to choose an membrane anchor upon which other components could be built. To avoid toxicity caused by foreign membrane proteins, we chose phosphatidylglycerol:: prolipoprotein diacylglyceryl transferase (Lgt) as transmembrane domain in the project. Lgt is an inner membrane protein of E.coli with seven transmembrane segments and has been successfully over expressed in E. coli without causing harm to cells.
SsDsbA is the signal recognition particle (SRP)-dependent signaling sequence of DsbA. SsDsbA-tagged proteins are exported to the periplasm through the SRP pathway. With ssDsbA fused to the N-terminus, engineered membrane proteins are expected to be anchored onto inner membrane of E.coli (Fig.2 ).
 Fig.2 : Construction sketch of membrane assemblies Membrane Localization Test
To justify the localization ability of fusion membrane proteins mentioned above, we constructed a novel fusion protein called BlaLG. β-lactamase is fused to the N-terminus of Lgt and GFP is fused to the C-terminus of Lgt. The fusion protein in under control of araBAD promoter (Fig.3 ). http://partsregistry.org/wiki/index.php?title=Part:BBa_K771401 BBa_K771401
 Fig.3 : Details of fusion protein BlaLG for membrane localization
Fluorescence Test
To visualize the localization of fusion protein BlaLG, we adopted fluorescence test. GFP fused to the C terminus of BlaLG enables us to have a closer look at where the membrane proteins are localized (Fig.3).
Under laser confocal microscope, we can observe the location of the fluorescence, thus to confirm the exact subcellular localization of the fusion protein.
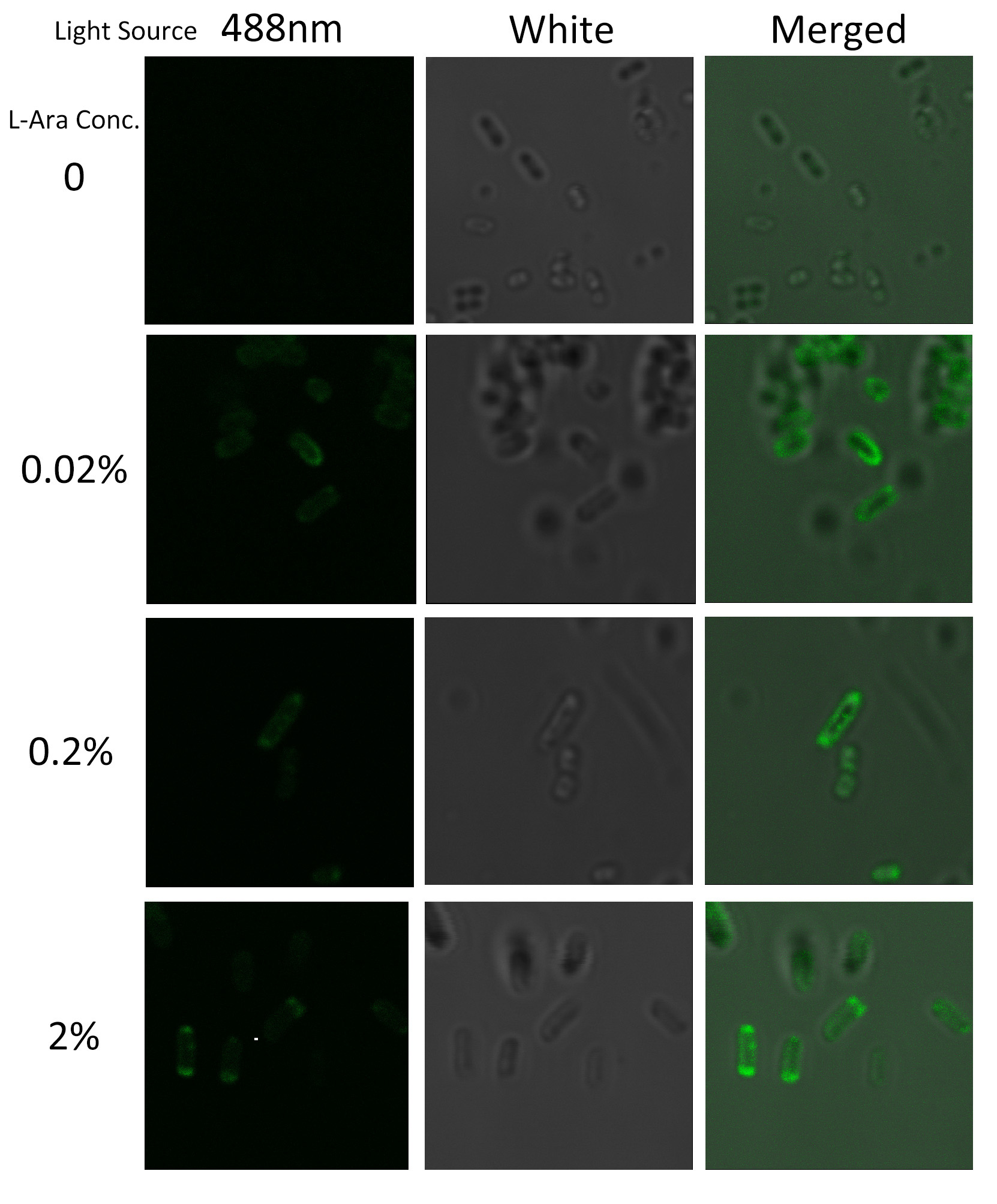 Fig.4 :Bacteria carrying BlaLG induced at different concentration of L-arabinose. It is observed that green fluorescence intensity of E.coli margin is higher than that of cytoplasm. Fig.4 proves that BlaLG has been localized to membrane.
Antibiotics Test
In this section, we test the membrane localization ability of fusion membrane protein by observing host cells' growth phenotype under different level of Ampicillin. Besides, we find the optimal inducer (L-arabinose) concentration for expressing membrane proteins.
β-lactamase,a bacterial enzyme which must be exported to the periplasm in order to confer significant resistance to β-lactam antibiotics, such as ampicillin. Note that N-terminus of Lgt faces the periplasm and C-terminus faces the cytoplasm (Fig.3). Hence, if our fusion membrane protein is correctly anchored to membrane, β-lactamase is expected to be functional and host cells should be able to grow on culture media containing ampicillin.
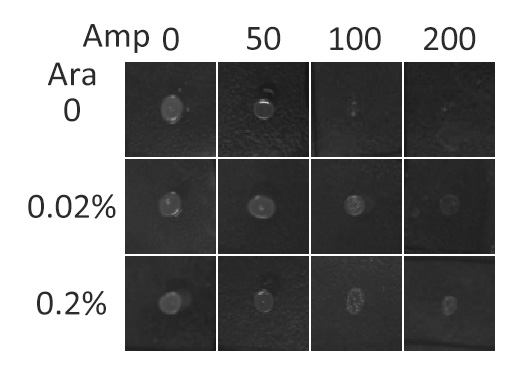 Fig.5 :Growth phenotypes of E. coli cells expressing BlaLG on LB agar media with concentration of inducer L-arabinose from 0 to 0.2%. We increased the concentration of ampicillin from 0 to 200 (μg/ ml).Bacteria carrying BlaLG were able to grow at ampicillin concentration of 200 μg/ ml with induction Fig.5 showed that our fusion protein has been correctly localized to membrane. Besides, a very low L-arabinose concentration at 0.02% is already enough to induce sufficient amount of membrane protein.
To further characterize features of the Membrane Scaffold System and find optimal inducer concentration, we test the growth condition of E.coli carrying gene of BlaLG under different concentration of L-Arabinose and Ampicillin. We expect to see more colonies grown at higher Ampicillin concentration if the inducer concentration is optimal.
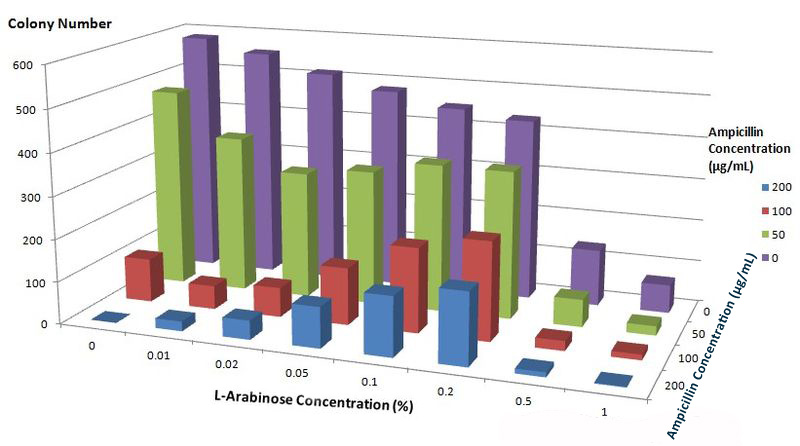 Fig.6 :Growth condition of E. coli cells expressing BlaLG on LB agar media with concentration of L-arabinose and Ampicillin. Single colony is picked and cultivated at 37℃ until OD value reaches 0.7. Bacteria cultures are diluted by 1:100000 and coated onto plates containing different concentration of L-Arabinose and Ampicillin. Growth condition is measured through counting colonies on plates. The result apparently showed that at L-Arabinose concentration of 0.1% and 0.2%, more bacteria could grow on high concentration of Ampicllin. Thus, L-Arabinose concentration of 0.1% and 0.2% best suits membrane protein expression in Project Membrane Magic.
Reference
1.Pailler, J., W. Aucher, et al. (2012). "Phosphatidylglycerol:: prolipoprotein diacylglyceryl transferase (Lgt) of Escherichia coli has seven transmembrane segments, and its essential residues are embedded in the membrane." Journal of bacteriology 194(9): 2142-2151.
2.Schierle, C. F., M. Berkmen, et al. (2003). "The DsbA signal sequence directs efficient, cotranslational export of passenger proteins to the Escherichia coli periplasm via the signal recognition particle pathway." Journal of bacteriology 185(19): 5706-5713.
3.Skretas, G. and G. Georgiou (2010). "Simple genetic selection protocol for isolation of overexpressed genes that enhance accumulation of membrane-integrated human G protein-coupled receptors in Escherichia coli." Applied and environmental microbiology 76(17): 5852-5859.
|