Team:Peking/Project/Luminesensor/Characterization
From 2012.igem.org
Sensitivity
1. Set-up & Brief Procedure
We tested the sensitivity of Luminesensor by examine the light-dependent transcriptional activity of a GFP-ssrA reporter. ssrA is a protein tag that induces fast degradation of protein, which in our case facilitated the observation of transcriptional activity. Based on the consideration of guaranteeing accuracy and precision, our setup (Figure 2):
Figure 2. Setup
consists of three central parts: light source, incubator and 48-well plate. On account of high sensitivity, protecting the system from the preventable light exposure with the purpose of acquiring the accurate results which is the true reflection of our sensitivity is necessary. In order to solve the problems, we focus on two foremost aspects: utilizing attenuators to weaken the light intensity and using tin foil to avoid light leakage. For more details about how we conducting the experiment, see experiment procedures. In our experiments, illumination with different light intensity conditions at 460nm peak light from blue LED arrays for 16 hours show marked light-depressed reporter gene transcription, which indicates that under different blue light exposure conditions, there was hardly any light-induced reporter gene transcriptional activity. But when in the dark environment (packaged with three layers of aluminum foil), our systems showed extremely high GFP expression (Figure 3)
Figure 3. Expression of GFP.
There are a couple of plausible and conceivable reasons for why our system possesses such high sensitivity that can sense unbelievably weak light intensity. First, it includes the unusually stable, photo-activated state of VVD, which causes the system to be extremely sensitive to light. Secondly, without relying on the addition of chromophores, which might impede the growth of cells, our system can directly regulate gene expression without trade-off.
2. Results and Conclusions
Cells exposed to different light intensity expressing Luminesensor showed manifest light-repressed reporter gene transcription. As shown in the figure, all of the cells with dissimilar attenuators showed incredible repression efficiency (figure 4: luminance measurement of different attenuator).

Figure 4. luminance measurement of different attenuator.
It proves that once the system is exposed to natural light, reporter gene transcription will be absolutely depressed regardless of the attenuators we attached to it. In other words, all of the systems protected from equal to or lower than natural-light-intensity light blocked light-depressed gene expression as expected. Besides, in the fifth sets of data, which was entirely in the dark state, the expression of GFP ran up to a high degree of 50,000. Taking everything into account, our luminesensor does possess high sensitivity that across several orders of magnitude. Our next work plan is to improve our devices and attenuators in order to find out the threshold point in which the light-depressed gene expression will present linear results.
We are impartially proud of accomplishing the goal of controlling gene expression using light without excessive or unnecessary energy or substrates, and we have respectable and appropriate reasons to have complete faith in our system; that it has imposed specific innovative challenges of light-dependent process for the future researchers.
Orthogonal Test
Abstract
It is our biggest concern whether lexA408VVD works indepently with endogenous LexA. Two sections of testing expriments were carried out simultaneously. To our satisfication, the results of different support in vivo orthogonality.
Amazingly sensitive as LexA-VVD system is, its application was limited for the wildspread of endogenous lexA repressor in cells. We have to maximum its biological orthogonality to get a really plug-and-play device. Therefore our part can work effciently in widely used host strains, e.g. DH5&alpha, BL21(DE3).
We had found from literature that LexA408, a lexA variant, works independently with wild-type lexA in E.coli; they recognize different sequences and have orthogonal DNA binding specfities.
We then designed the LexA408-VVD fusion protein and a series of its exclusive promoters fused to op408. These novel clones were achieved by point mutations from their wild-typel counterparts. To make things easier, we named the protein lexA408-VVD.
In our experiments, protein expression work was conducted in wild-type BL21(de3) strain. GFP was selected as a reporter and was fused downstream to op408. In this condition, GFP expression has a negative relation with repression activity of the protein. To be more specific, higher level of green fluorescent indicates weaker repression effect; lower expression of GFP stands for stronger repression.
Experimental Design
To prove the orthogonality, facts that LexA408-VVD and endogenous LexA work totally independently are needed. Considering practical efficiency, three points of evidence are to collect:
- 1. LexA-responsive promoters are repressed in wild-type strains; LexA408-VVD-responsive promoters are not blocked in wild-type strains;
- 2. LexA408-VVD efficiently represses its target in blue illumination, while it does not repress targets in total dark.
Figure 5. Shows observable color of plates where cell cultures growing under totally dark or light conditions. Different colors are used to indicate three levels of GFP expression.
Detailed Methods
Experiments were carried out in parallel: taking photos of plates as visual evidence; measuring GFP to quatitify the effect. Detailed protocols are as follows:
- 1. Visible display:
-
- Inoculate a single colony into 1.5ml centifuge tube, shaking cultivate cells at 250 rpm/min for 2-3 h at 30℃, then the cell culture are prepared for streaking;
- Streak cell culture onto right plates;
- Ioncubate in totally dark and light state at 30℃ for 1 or 2 days;
- Take photos of right plates;
- 2. Data collection:
-
- Inoculate a proper colony in liquid media;
- Shaking cultivate cells until platform stage at 30 ℃;
- Dilute celll cultures with selective media in proportion of 1:500;
- Shaking cultivate cells until platform stage at 30 ℃ in fully dark or under blue light;
- Harvest cells by centrifugation, resuspend pellets in PBS;
- Measuring GFP expression with 96-well-plate reader;
Results
Below is a collage of our plates. The plates are placed in groups at 30℃ in either total dark or blue illumination. Visual results fit well with Figure1.Visual look of plates were positive evidence for us.
Figure 6. Plates display. Streaking certain strain onto selected media.
Apart from those photos, more accurate results were obtained with Fluorescent Microplate Reader. Below are histograms based on our experimmental data. They clarify how fluoresence changes with lighting condition in different strains.
Figure 7. GFP expression under different promoters. The tests were carried out in strains capable of expressing LexA408-VVD.
It repeartedly turned out that:
- 1. reporters under wild-type promoters hardly express in either illuminated or dark state; their colonies are yellow. In contrast, GFPs fused to 408 operators were not blocked in strains withour lexA408-VVD, thus they looks bright green.
- 2. BL21 (DE3) strains containing both lexA408-VVD and its reporters twist swiftly and sensitively from one states to the other, giving high definition output including lighting condition.
To sum up, LexA408-VVD works independently with endogenous lexA. What’s more, thet are unrelated to most currently prevalent ribosome-mRNA, mRNA -rRNA orthogonal pairs, etc. Our devices are to help enrich logic gates system and enable the establishment of more complexed network in synthetic biology. We feel proud thinking that more and more r resaerchers will benefit and like it, our lexA408-VVD.
Optimization
To test whether our designated mutations would optimize our luminesensor in respect to reversibility and the on/off ratio, we co-transformed the ColE-GFP (GFP driven by luminesensor repressible promoter ColE) plasmid with four versions of our luminesensor plasmid into BL21 (ΔLexA ΔSulA): the original LexA-VVD(WT), LexA-VVD(74), LexA-VVD(135), and LexA-VVD(74+135). The resulting four strains were designated LV-WT, LV-74, LV-135, LV-74-135, respectively. The overnight culture of the four strains were diluted 500 times and divided into two groups: one exposed to blue light and the completely wrapped with aluminum foil. After incubation for 16 hours, GFP expression levels were measured. As shown in figure X, LV-135 has an increased on/off ratio compared to the original LexA-VVD, while the LV-74 and LV-74-135 show reduced on/off ratios, which is in accordance with our model .
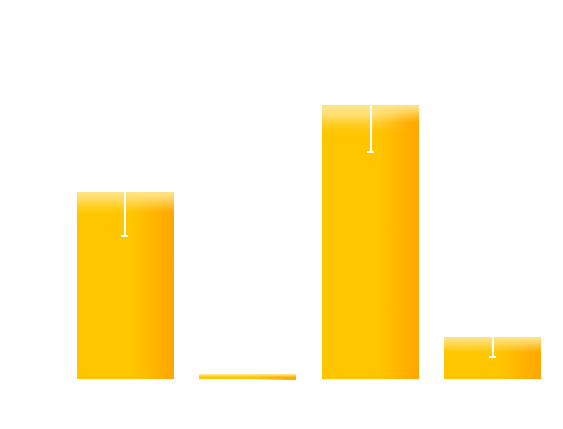
Figure 8. The on/off ratio of GFP expression level of the four strains.
In order to determine whether the LexA-VVD(135) has enhanced reversibility in comparison to the original LexA-VVD, the temporal change of GFP expression level of dilated overnight culture of the strains LV-WT and LV-135 under blue light were measured at 2 hour intervals for 26 hours. As shown in figure Y, the GFP expression level of both of the two strains began to rise after incubating at dark for about 10 hour. We speculated that the GFP expression level of the two strains is mainly determined by the equilibrium between GFP production and degradation and the dimerized LexA-VVD(WT) and LexA-VVD(135) dissociate in a shorter time scale compared to the time needed to establish the equilibrium of the GFP production and degradation.
Figure 9. The temporal change of GFP expression of the LV-WT and LV-135
Conclusion
In the end, we have finally constructed a LexA-VVD fusion protein that will function as a transcription repressor in response to blue light. And since we have introduced mutations into the LexA DNA binding domain, the Luminesensor proved orthogonal to endogenous bacteria SOS system. Futher introduced mutations in VVD photosensitive domain significantly improved the dynamic range of our optogenetic module. Equipped with this optimized LexA408-VVD74 Luminesensor, we are now able to apply it in real bacteria system to achieve amazing accomplishments.
Reference
reference here
 "
"