Team:Peking/Modeling/Luminesensor/Optimization
From 2012.igem.org
TheTinaChen (Talk | contribs) |
|||
| (12 intermediate revisions not shown) | |||
| Line 10: | Line 10: | ||
<h3 id="title1">Parameter Analysis & Optimization</h3> | <h3 id="title1">Parameter Analysis & Optimization</h3> | ||
<p> | <p> | ||
| - | After modeling the | + | After modeling the prototype <i>Luminesensor</i>, we attempted to optimize it in a rational way. We have tuned the parameters both up and down, one by one, and finally discovered four parameters which predominantly influence the performance of the <i>Luminesensor</i>. |
</p> | </p> | ||
<table style="width:600px;"><tr> | <table style="width:600px;"><tr> | ||
| Line 32: | Line 32: | ||
<div> | <div> | ||
<p class="description"> | <p class="description"> | ||
| - | Figure | + | Figure 1. Results of Parameter Analysis. "Up" means tuning the parameter up to 10 times. This figure implied that tuning up these parameters can optimize the Luminesensor. |
</p> | </p> | ||
</div> | </div> | ||
</div> | </div> | ||
<p> | <p> | ||
| - | Within the two parameters to enhance contrast, K<sub>2</sub> (vivid association equilibrium constant) is related to the association | + | Within the two parameters to enhance contrast, K<sub>2</sub> (vivid association equilibrium constant) is related to the association of Vivid protein and K<sub>5</sub> (dimerized LexA binding equilibrium constant) is related to the cooperative binding to DNA. |
<br /> | <br /> | ||
</p> | </p> | ||
| Line 44: | Line 44: | ||
<div> | <div> | ||
<p class="description"> | <p class="description"> | ||
| - | Figure | + | Figure 2. A molecular structure of VVD protein marking the position of residue 74. As shown in the figure, residue 74 (originaly isoleucine) resides in the vicinity of FAD (flavin adenine dinucleotide) molecule (the chromophore of VVD protein) and residue cystine 108. When VVD is excited by light, a covalent bound will form between residue cystine 108 and C4 of FAD, leading to VVD's conformational change to its light-state. The mutation I74V will interact with Cystine 108 and FAD molecule to destablize the adduct form, accelarating VVD's decay back to its dark-state. |
</p> | </p> | ||
</div> | </div> | ||
| Line 52: | Line 52: | ||
<div> | <div> | ||
<p class="description"> | <p class="description"> | ||
| - | Figure | + | Figure 3. A molecular structure of VVD protein marking the position of residue 135. As shown in the figure, residu 135(originally methionine) also resides near the FAD molecule. M135I is proposed to change the electronic environment of FAD molecule and somehow enhance VVD dimerization (possibly by stablizing light-state). The exact molecule mechanism is unclear. |
</p> | </p> | ||
</div> | </div> | ||
</div> | </div> | ||
<p> | <p> | ||
| - | By searching the data of mutant | + | By searching the data of mutant vivid protein in literature, we finally focused on these mutants: M135I in vivid dimerization domain to enhance K2 (vivid association equilibrium constant) and I74V of amino acids surrounding Cys108 to enhance k1 (vivid decay rate constant)<sup><a href="#ref1" title="Zoltowski, B.D., Vaccaro, B., and Crane, B.R. (2009). Mechanism-based tuning of a LOV domain photoreceptor. Nat. Chem. Biol. 5: 827: 834 ">[1]</a></sup>. The experimental results verified our prediction (See <a href="/Team:Peking/Project/Luminesensor/Characterization" title="">Project Luminesensor Characterization</a>). |
<br /> | <br /> | ||
</p> | </p> | ||
| Line 63: | Line 63: | ||
<img src="/wiki/images/2/2b/Peking2012_luminesensor_wiki.png" alt="Figure 8" /> | <img src="/wiki/images/2/2b/Peking2012_luminesensor_wiki.png" alt="Figure 8" /> | ||
<div> | <div> | ||
| - | <p class="description">Figure | + | <p class="description">Figure 4. Experiment result: Effects of introduced mutations on the contrast (on/off ratio) of <i>Luminesensor</i>. We can see that the mutation on M135 obviously improves the contrast (on/off ratio) of <i>Luminesensor</i>, which validate our modeling prediction. |
</p> | </p> | ||
</div> | </div> | ||
| Line 71: | Line 71: | ||
</p> | </p> | ||
<div class="floatC"> | <div class="floatC"> | ||
| - | <img src=". | + | <img src="/wiki/images/a/ad/Peking2012_Design_LexA408mutations.png" alt="Figure 9. Mutation 3" title="Figure 9. Mutation 3" style="width:500px;" /> |
<div> | <div> | ||
<p class="description"> | <p class="description"> | ||
| - | Figure | + | Figure 5. A molecular structure of N-terminal domain of LexA protein. The arrow points to the position of residue 40, 41 and 42. In LexA408 protein, three point mutations P40A, N41S and A42S are introduced into the N-terminal domain. These three mutations near the DNA binding surface of the protein will change its binding specificity. LexA408 will recognize a symmetrically altered sequence different from the one recognized by wild type LexA. |
| + | |||
</p> | </p> | ||
</div> | </div> | ||
| Line 80: | Line 81: | ||
</div> | </div> | ||
<div class="PKU_context floatR"> | <div class="PKU_context floatR"> | ||
| - | <h3 id=" | + | <h3 id="title2">Reference</h3> |
<p></p> | <p></p> | ||
<ul class="refer"><li id="ref1"> | <ul class="refer"><li id="ref1"> | ||
Latest revision as of 02:49, 27 September 2012
Parameter Analysis & Optimization
After modeling the prototype Luminesensor, we attempted to optimize it in a rational way. We have tuned the parameters both up and down, one by one, and finally discovered four parameters which predominantly influence the performance of the Luminesensor.
| Function | Parameter | Description | Remark |
| Reduce responsing time | k1 | Vivid lighting decay rate constant | Mainly on process from Light to Dark |
| k3 | rate constant of monomer LexA releasing from specific binding site | ||
| Enhance contrast | K2 | Vivid association equilibrium constant | More dimerization provides more binding opportunity |
| K5 | dimered LexA binding equilibrium constant | More binding affinity |
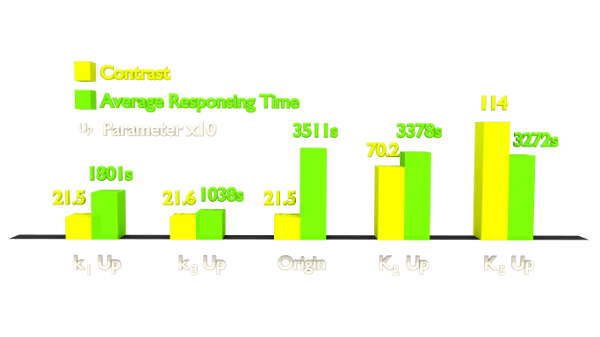
Figure 1. Results of Parameter Analysis. "Up" means tuning the parameter up to 10 times. This figure implied that tuning up these parameters can optimize the Luminesensor.
Within the two parameters to enhance contrast, K2 (vivid association equilibrium constant) is related to the association of Vivid protein and K5 (dimerized LexA binding equilibrium constant) is related to the cooperative binding to DNA.
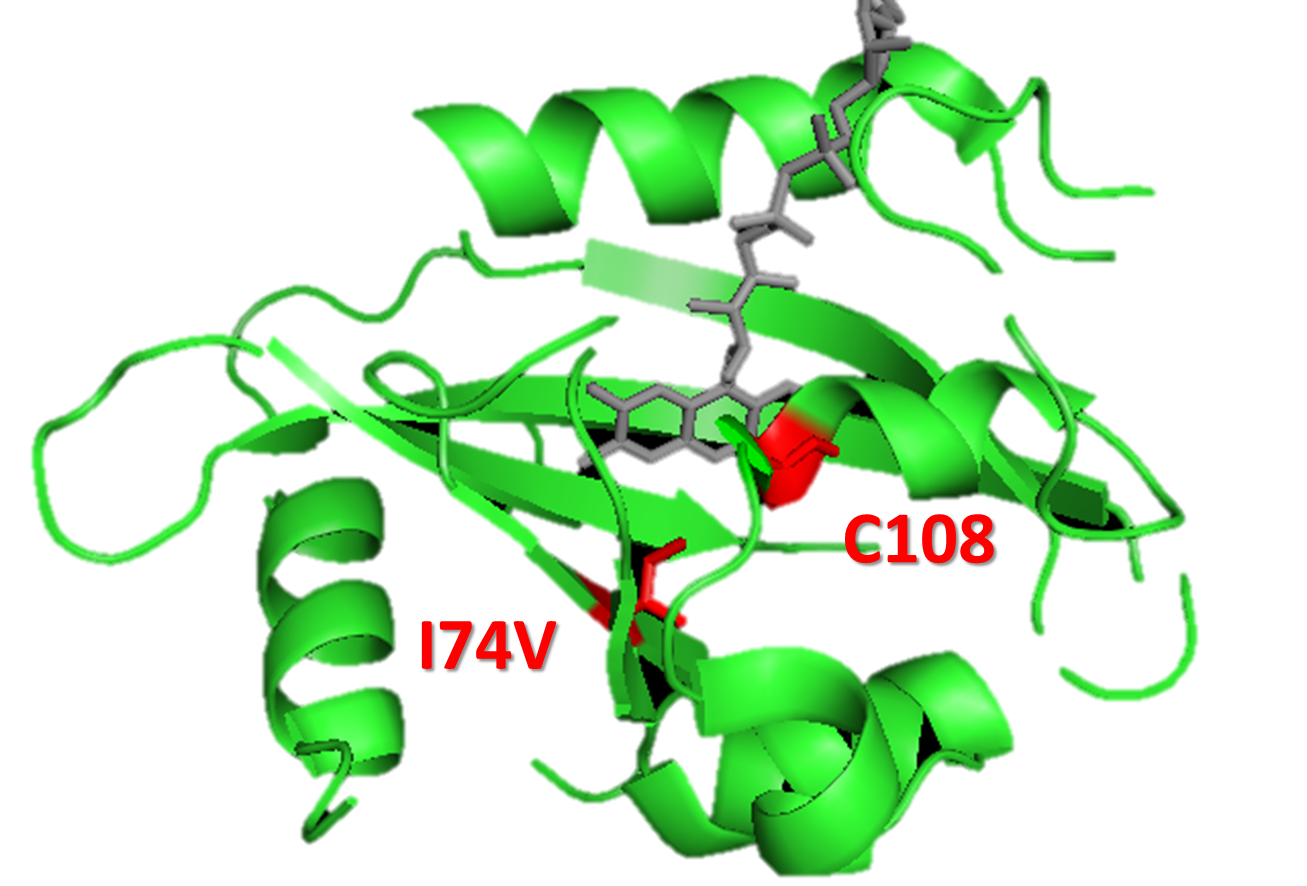
Figure 2. A molecular structure of VVD protein marking the position of residue 74. As shown in the figure, residue 74 (originaly isoleucine) resides in the vicinity of FAD (flavin adenine dinucleotide) molecule (the chromophore of VVD protein) and residue cystine 108. When VVD is excited by light, a covalent bound will form between residue cystine 108 and C4 of FAD, leading to VVD's conformational change to its light-state. The mutation I74V will interact with Cystine 108 and FAD molecule to destablize the adduct form, accelarating VVD's decay back to its dark-state.

Figure 3. A molecular structure of VVD protein marking the position of residue 135. As shown in the figure, residu 135(originally methionine) also resides near the FAD molecule. M135I is proposed to change the electronic environment of FAD molecule and somehow enhance VVD dimerization (possibly by stablizing light-state). The exact molecule mechanism is unclear.
By searching the data of mutant vivid protein in literature, we finally focused on these mutants: M135I in vivid dimerization domain to enhance K2 (vivid association equilibrium constant) and I74V of amino acids surrounding Cys108 to enhance k1 (vivid decay rate constant)[1]. The experimental results verified our prediction (See Project Luminesensor Characterization).
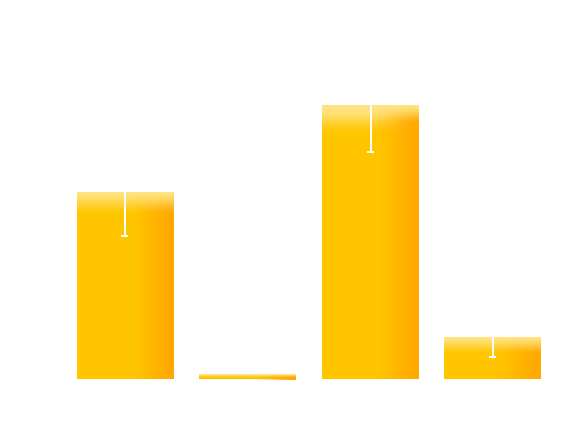
Figure 4. Experiment result: Effects of introduced mutations on the contrast (on/off ratio) of Luminesensor. We can see that the mutation on M135 obviously improves the contrast (on/off ratio) of Luminesensor, which validate our modeling prediction.
We also chose LexA408, the mutant of LexA, in order to increase K5 (dimered LexA binding equilibrium constant)[2], even though our main reason for choosing LexA408 over the wild-type LexA is due to the bio-orthogonality.
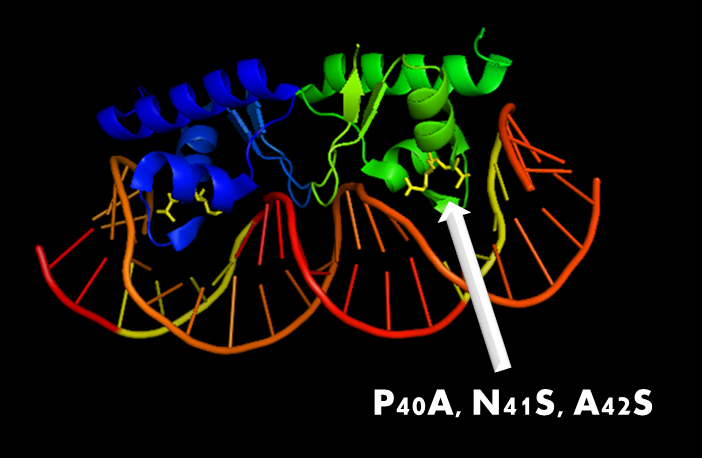
Figure 5. A molecular structure of N-terminal domain of LexA protein. The arrow points to the position of residue 40, 41 and 42. In LexA408 protein, three point mutations P40A, N41S and A42S are introduced into the N-terminal domain. These three mutations near the DNA binding surface of the protein will change its binding specificity. LexA408 will recognize a symmetrically altered sequence different from the one recognized by wild type LexA.
Reference
- 1. Zoltowski, B.D., Vaccaro, B., and Crane, B.R. (2009). Mechanism-based tuning of a LOV domain photoreceptor. Nat. Chem. Biol. 5: 827: 834
- 2. Dmitrova, M., Younes-Cauet, G., Oertel-Buchheit, P., Porte, D., Schnarr, M., Granger-Schnarr, M.(1998) A new LexA-based genetic system for monitoring and analyzing protein heterodimerization in Escherichia coli. Mol. Gen. Genet., 257: 205: 212
 "
"