Team:HokkaidoU Japan/Notebook/aggregation Week 10
From 2012.igem.org
(3e) |
(3rd digestion-transformation) |
||
| Line 76: | Line 76: | ||
#Incubated at 37C for hrs. | #Incubated at 37C for hrs. | ||
</p> | </p> | ||
| + | |||
| + | |||
| + | ==Estimation of concentration of eCFP-RBS-pSB1A2 and pBAD-RBS-pSB1A2== | ||
| + | <p> | ||
| + | No.2,5 means the colony number of colony PCR of eCFP-RBS-pSB1A2. | ||
| + | |||
| + | |||
| + | [[image:|thumb|electrophoresis result]] | ||
| + | |||
| + | |||
| + | We estimated the concentration of eCFP-RBS-pSB1A2 is 30 ng/ul and pBAD-RBS-pSB1A2 is 50ng/ul. | ||
| + | </p> | ||
| + | |||
| + | |||
==Digestion of eCFP-RBS-pSB1A2 and pBAD-RBS-pSB1A2== | ==Digestion of eCFP-RBS-pSB1A2 and pBAD-RBS-pSB1A2== | ||
| Line 84: | Line 98: | ||
{|class="hokkaidou-table-digestion" | {|class="hokkaidou-table-digestion" | ||
|- | |- | ||
| - | |DNA solution ( | + | |DNA solution ( 30ng/ul) |
| - | | | + | |22 ul |
|- | |- | ||
| - | | | + | |XbaI |
|1 ul | |1 ul | ||
|- | |- | ||
| - | | | + | |PstI |
|1 ul | |1 ul | ||
|- | |- | ||
| - | | | + | |10xM buffer |
| - | | | + | |3 ul |
|- | |- | ||
|DW | |DW | ||
| - | | | + | |3 ul |
|- | |- | ||
|Total | |Total | ||
| - | | | + | |30 ul |
|} | |} | ||
| Line 109: | Line 123: | ||
{|class="hokkaidou-table-digestion" | {|class="hokkaidou-table-digestion" | ||
|- | |- | ||
| - | |DNA solution ( | + | |DNA solution ( 50ng/ul) |
| - | | | + | |3 ul |
|- | |- | ||
| - | | | + | |SpeI |
|1 ul | |1 ul | ||
|- | |- | ||
| - | | | + | |PstI |
|1 ul | |1 ul | ||
|- | |- | ||
| - | | | + | |10xH buffer |
|2 ul | |2 ul | ||
|- | |- | ||
|DW | |DW | ||
| - | | | + | |13 ul |
|- | |- | ||
|Total | |Total | ||
| Line 129: | Line 143: | ||
| - | control ( | + | control (eCFP-pSB1A2) |
{|class="hokkaidou-table-digestion" | {|class="hokkaidou-table-digestion" | ||
|- | |- | ||
|DNA solution (30 ng/ul) | |DNA solution (30 ng/ul) | ||
| - | | | + | |5 ul |
|- | |- | ||
|XbaI | |XbaI | ||
| + | |1 ul | ||
| + | |- | ||
| + | |SpeI | ||
|1 ul | |1 ul | ||
|- | |- | ||
|10xM buffer | |10xM buffer | ||
|2 ul | |2 ul | ||
| - | |||
| - | |||
| - | |||
|- | |- | ||
|DW | |DW | ||
| Line 173: | Line 187: | ||
| - | [[image: | + | [[image:|thumb|digestion result]] |
| Line 179: | Line 193: | ||
</p> | </p> | ||
| - | |||
| - | |||
| - | |||
| - | + | ==Ethanol precipitation of digestion products (eCFP-RBS and pBAD-RBS-pSB1A2) and estimation of concentration== | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | ==Ethanol precipitation of digestion products (eCFP and RBS-pSB1A2) and estimation of concentration== | + | |
<p> | <p> | ||
#Added 5 ul of NaoAc, 1.5 ul of glycogen and 125 ul of 100% ethanol. | #Added 5 ul of NaoAc, 1.5 ul of glycogen and 125 ul of 100% ethanol. | ||
| - | |||
| - | |||
#Centrifuged in 15000 rpm, 15 min at 4C. | #Centrifuged in 15000 rpm, 15 min at 4C. | ||
| + | #Remove supernatant and added 220 ul of 70% ethanol. | ||
| + | #Centrifuged in 15000 rpm, 10 min at 4C. | ||
#Remove supernatant and air drying in room temperature then added 5 ul of DW. | #Remove supernatant and air drying in room temperature then added 5 ul of DW. | ||
| - | [[image: | + | [[image:|thumb|ethanol precipitation result]] |
| - | We estimated the concentration of ethanol presipitation products.The concentration of Insert DNA solution is about 20 ng/ul and Vector DNA solution is about | + | We estimated the concentration of ethanol presipitation products.The concentration of Insert DNA solution is about 20 ng/ul and Vector DNA solution is about 50 ng/ul. |
</p> | </p> | ||
| - | ==Ligation of eCFP and RBS-pSB1A2== | + | ==Ligation of eCFP-RBS and pBAD-RBS-pSB1A2== |
<p> | <p> | ||
{|class="hokkaidou-table-ligation" | {|class="hokkaidou-table-ligation" | ||
|- | |- | ||
| - | |Vector DNA ( | + | |Vector DNA (50 ng/ul) |
| - | |1 ul | + | |1.5 ul |
|- | |- | ||
| - | |Insert DNA (20 ng/ul | + | |Insert DNA (20 ng/ul) |
|4 ul | |4 ul | ||
|- | |- | ||
|Ligation Mighty Mix | |Ligation Mighty Mix | ||
|6 ul | |6 ul | ||
| + | |- | ||
| + | |DW | ||
| + | |0.5 ul | ||
|- | |- | ||
|Total | |Total | ||
| - | | | + | |12 ul |
|} | |} | ||
| Line 241: | Line 248: | ||
</p> | </p> | ||
| - | ==Transformation of eCFP-RBS-pSB1A2== | + | ==Transformation of pBAD-RBS-eCFP-RBS-pSB1A2== |
<p> | <p> | ||
| - | #Mixed 2 ul eCFP-RBS-pSB1A2 ligation product to 50 ul of thawed competent cells on ice. | + | #Mixed 2 ul pBAD-RBS-eCFP-RBS-pSB1A2 ligation product to 50 ul of thawed competent cells on ice. |
#Incubated on ice for 30 min. | #Incubated on ice for 30 min. | ||
#Mixed 350 ul of LB. | #Mixed 350 ul of LB. | ||
| Line 249: | Line 256: | ||
#Plated 300 ul of the culture onto first dish and spread. | #Plated 300 ul of the culture onto first dish and spread. | ||
#Mixed 450 ul of LB to 50 ul of the culture and plated 300 ul of it onto second dish and spread. | #Mixed 450 ul of LB to 50 ul of the culture and plated 300 ul of it onto second dish and spread. | ||
| - | #Incubated the plates at 37C for | + | #Incubated the plates at 37C for hours. |
</p> | </p> | ||
Revision as of 21:40, 3 September 2012
September 3rd
Colony PCR of eCFP-RBS-pSB1A2
| DNA solution | 4 ul |
| Kapa-Taq(Taq polymerase) | 5 ul |
| Forward Primer(EX-F primer) | 0.5 ul |
| Reverse Primer(PS-R down primer) | 0.5 ul |
| Total | 10 ul |
| Number | Degree | Second |
| 1 | 95 | 120 |
| 2 | 95 | 30 |
| 3 | 68.9 | 30 |
| 4 | 72 | 60 |
| 5 | 72 | 60 |
| 6 | 4 | HOLD |
Cycle:2~4 x 35
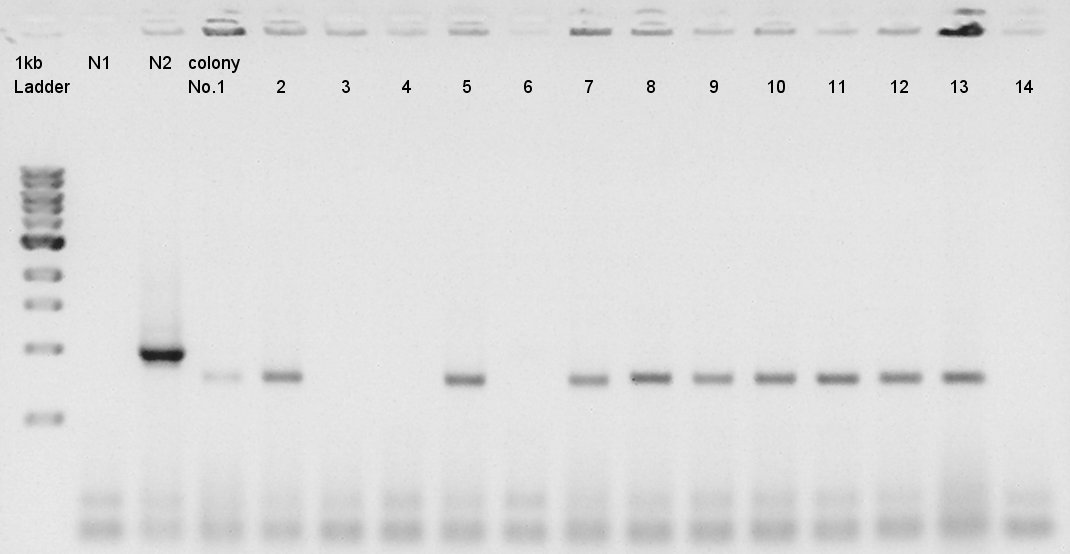
We used N1 (DW only) and N2 (ptetR-RBS-eCFP-dT-pSB1A2) as controls. Desired product is about 776bp.
We confirmed that about 70% of ligated DNA formed our desired construct. We selected No.2 and 5 colony for incubation and store No.7 and 8 colony mixture at 4C.
Incubation of eCFP-RBS-pSB1A2 for mini-prep
- Prepared 2 ml LBA into culture tubes.
- Re-suspended 2 colony mixture (No.2 and No.5 respectively).
- Incubated at 37C for hrs.
Estimation of concentration of eCFP-RBS-pSB1A2 and pBAD-RBS-pSB1A2
No.2,5 means the colony number of colony PCR of eCFP-RBS-pSB1A2. [[image:|thumb|electrophoresis result]] We estimated the concentration of eCFP-RBS-pSB1A2 is 30 ng/ul and pBAD-RBS-pSB1A2 is 50ng/ul.
Digestion of eCFP-RBS-pSB1A2 and pBAD-RBS-pSB1A2
To make a construct of pBAD-RBS-eCFP-RBS-pSB1A2, we digested eCFP-RBS-pSB1A2 with XbaI & PstI and pBAD-RBS-pSB1A2 with SpeI & PstI. And we digested eCFP-pSB1A2 with XbaI & SpeI as a control for confirmation of the ability to digest. Insert (eCFP-RBS-pSB1A2)
| DNA solution ( 30ng/ul) | 22 ul |
| XbaI | 1 ul |
| PstI | 1 ul |
| 10xM buffer | 3 ul |
| DW | 3 ul |
| Total | 30 ul |
Vector(pBAD-RBS-pSB1A2)
| DNA solution ( 50ng/ul) | 3 ul |
| SpeI | 1 ul |
| PstI | 1 ul |
| 10xH buffer | 2 ul |
| DW | 13 ul |
| Total | 20 ul |
control (eCFP-pSB1A2)
| DNA solution (30 ng/ul) | 5 ul |
| XbaI | 1 ul |
| SpeI | 1 ul |
| 10xM buffer | 2 ul |
| DW | 11 ul |
| Total | 20 ul |
| Number | Degree | Minute |
| 1 | 37 | 120 |
| 2 | 60 | 15 |
| 3 | 4 | HOLD |
[[image:|thumb|digestion result]]
From this image, we confirmed that DNA were digested into fragments and all of restriction enzyme worked.
Ethanol precipitation of digestion products (eCFP-RBS and pBAD-RBS-pSB1A2) and estimation of concentration
- Added 5 ul of NaoAc, 1.5 ul of glycogen and 125 ul of 100% ethanol.
- Centrifuged in 15000 rpm, 15 min at 4C.
- Remove supernatant and added 220 ul of 70% ethanol.
- Centrifuged in 15000 rpm, 10 min at 4C.
- Remove supernatant and air drying in room temperature then added 5 ul of DW.
Ligation of eCFP-RBS and pBAD-RBS-pSB1A2
| Vector DNA (50 ng/ul) | 1.5 ul |
| Insert DNA (20 ng/ul) | 4 ul |
| Ligation Mighty Mix | 6 ul |
| DW | 0.5 ul |
| Total | 12 ul |
Ligation reaction time was in detail below.
| Degree | Minute |
| 16 | 30 |
| 65 | 10 |
| 4 | Hold |
Transformation of pBAD-RBS-eCFP-RBS-pSB1A2
- Mixed 2 ul pBAD-RBS-eCFP-RBS-pSB1A2 ligation product to 50 ul of thawed competent cells on ice.
- Incubated on ice for 30 min.
- Mixed 350 ul of LB.
- Prepared and Labeled two plastic plates with LB plate medium which contained appropriate antibiotics (LBA).
- Plated 300 ul of the culture onto first dish and spread.
- Mixed 450 ul of LB to 50 ul of the culture and plated 300 ul of it onto second dish and spread.
- Incubated the plates at 37C for hours.
 "
"