Team:CU-Boulder/Project
From 2012.igem.org

Contents |
Project 1: Model System for studying quorum activation of pathogenicity
Quorum sensing is a system of bacterial communication that coordinates gene expression based on population density. Bacteria secrete a signaling molecule, such as N-Acyl Homoserine Lactones (AHL), and each bacteria has a extracellular receptor that specifically binds the signaling molecule which activates transcription factors specific to certain genes. Many pathogenic bacteria use quorum sensing to activate their virulent factors, as it is not energetically favorable for a single bacterium to release its virulent factors until it is in a large enough population for infect a host. For example, Pseudomonas Aeruginosa is an opportunistic pathogen that contributes to sepsis and its virulent proteins are not expressed constitutively but in a density dependent manner.(1)(2). In our project the CU-Boulder iGEM team aims to disrupt quorum sensing in order to reduce pathogenicity of otherwise virulent bacteria.
It has been well documented in research that bacteria have used quorum sensing signals to activate pathogenic factors. The gram negative bacteria P. aeruginosa synthesizes AHLs as a bacterial density indicator, and upon reaching a specific threshold, pathogenic factors are activated (Iglewski et al. 2000). P. aeruginosa is a common opportunistic infection that occurs most frequently in immunocompromised people (Iglewski et al. 2000). Though this same quorum sensing mechanism for activation of pathogenic traits has also been demonstrated in Salmonella, E. coli, and Y. pestis.(-----). Quorum sensing factors also play a role in activating multi-bacterial responses such as biofilm production, and food rot. (…) In order to study how to inhibit these pathogenic effects of quorum sensing, we needed a system that would respond to quorum signals, and was a safe alternative to the pathogenic bacteria. Therefore, we characterized the model quorum sensing system of Allivibrio fisheri, which luminesces in response to AHLs.
Allivibrio fischeri is a gram-negative bacteria that is a biological model for both quorum-sensing and bioluminescence research. This rod-shaped bacteria participates in symbiotic relationships with various marine organisms, most notably the Bobtail squid. This squid hosts Allivibrio fisheri and uses its light to hide the squid's shadow in shallow waters to avoid potential predators. We decided to use Allivibrio fisheri WH1 strain as our model organism because it is a safe organism to handle, it has a LuxI/LuxR quorum sensing system that it usees to detect the concentration of AHLs, and it responds with luminescence at a visible wavelength when the threshold level of AHLs is reached.
When LuxR detects AHLs it activates the rest of the Lux operon. The Lux operon (below) consists of 5 essential genes: LuxA, LuxB, LuxC, LuxD, and LuxE.

LuxC, LuxD, and Lux E form the fatty acid reductase enzyme complex that synthesizes fatty acid aldehydes with acyl-CoA as the starting substrate. The 3 gene products form a complex conglomerate (below) to increase kinetic efficiency. LuxD is the transferase, LuxE is the synthetase, and LuxC is the reductase.

LuxA and LuxB form a dimer that reduces O2 while oxidizing the aldehyde and FMNH2. This oxidation produces the blue-green light (~490nm). Another representation of the overall reaction is shown below:

Also, it has been shown that the LuxG gene, although not essential, does increase brightness by functioning as a flavin reductase.(http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2258676/)
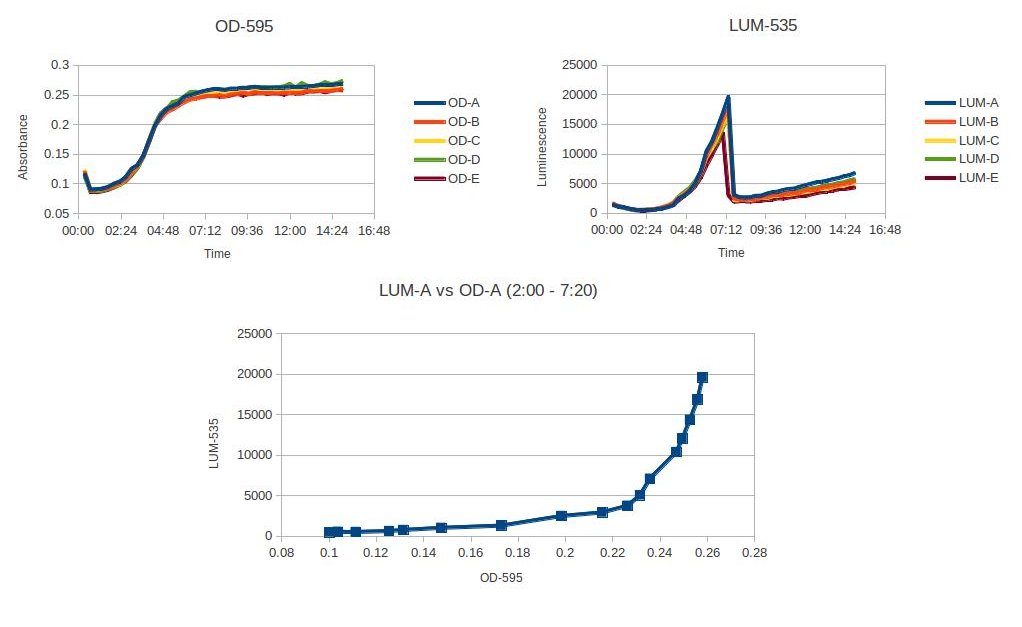
We utilized Allivibrio fisheri because the luminosity is easy to measure with a high degree of sensitivity and low background noise. The organism is visually luminous at a wavelength that can be perceived by the human eye, and we can therefore use the organism as a model for reporting when pathogenic factors would have been initiated in other bacteria. Using a plate reader, we characterized at what OD the A. fisheri starts to glow, and characterized how the density of the organism affected the luminosity. (Figure)
Basic Idea: Transform an E. Coli with the 5 essential Lux genes (LuxABCDE) as well as Lux G. This Lux brick will be under the control of a UV induced promoter system and a C1 lambda inverter system. This means that while the bacteria is exposed to UV light, the C1 protein will repress C1 regulated promoter that is upstream of the Lux brick, and no light will be produced. On the flip side, when the bacteria is not exposed to UV light, no C1 will be made and the C1 regulated promoter will be activated and produce the Lux brick and the culture will luminesce. To further optimize the system, we have expressed the individual Lux genes in varying levels to see which of the 6 is the rate limiting step in bioluminescence.
Possible Promoter systems:
Circadian clock - KaiABC/SasA/RpaA is the simplest oscillator found in cyanobacteria, Synechococcus elongatus
Blue Light Induced - YcgE/YcgF is endogenous to E. Coli. We have the system built and are testing it now.
Red light Induced - Cph8/PCB/OmpR is a system that won iGEM in 2006 but every team that has used it since has run into problems
Project 1: Experiments
1. '''Assemble the LuxABCDE operon + LuxG''' The Cambridge 2010 "E. Glowi" team submitted the Lux operon to the registry, BBa_K325905, but several attempts at transforming E. Coli with this part failed either due to its large size (6478bp) or due to an inaccurate registry submission. Several other teams have tried to use their part since 2010 but none have been successful. For this reason, our team is BioBricking each Lux gene individually and submitting them to the registry.
2. '''Optimize the Lux System''' Once the Lux brick is working in a cell, we ligated each Lux gene to a different strength promoter to maximize luminescence while conserving energy. Because LuxC is the scaffolding on which the LuxCDE reductase complex is built, we started by up regulating LuxC under a high constitutive promoter.
3. '''Regulate the system under a UV induced promoter''' We ligated a UV induced promoter to a C1 lambda inverter system which is upstream of the optimized Lux Brick. This is our final project as the Lux Brick will only luminesce in low light conditions (night) and the system will be repressed in high light conditions (daytime). Visit our Modeling page to see how we used rapidly degrading LVA tags to ensure a quickly responding system.
Project 1: Results
The figure above shows the plate reader data characterizing Allivibrio fisheri's luminosity at a specific OD. The luminosity starts to increase at OD .23, and then after the A. fisheri becomes stationary, the luminescence sharply decreases. This experiment was read on a plate reader every 20min with 5 trials. A. fisheri was grown up for 4 hours in a room temperature shaker. Then pelleted and washed three times with LB + NaCl. Diluted to an OD of .05 and placed in the plate reader.
Project 2: Biofilm Prevention
Quorum Sensing in LuxR containing Bacteria
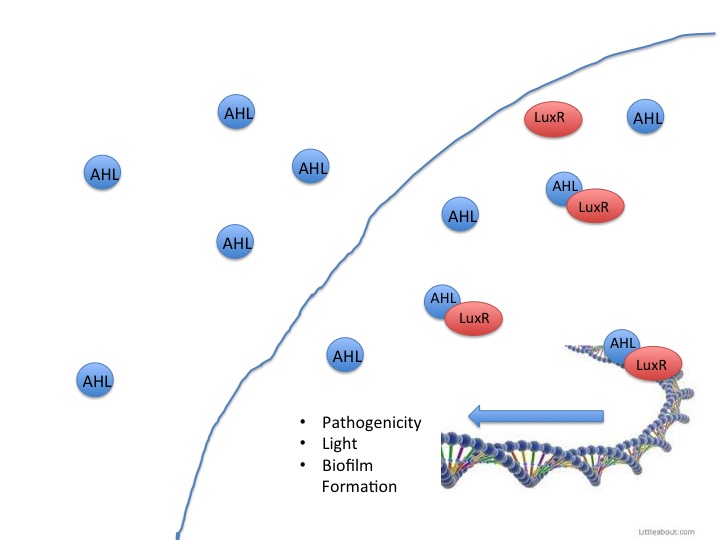
This year the CU iGEM team has harnessed the quorum sensing factors endogenous in the Salmonella enterica serovar typhimurium LT2 strain to detect AHLs produced by other bacteria. Neither Salmonella nor E. coli have been shown to produce detectable levels of AHLs, so they were model organisms for the detection of other AHL producing bacteria such as Vibrio fisheri, or pathogenic bacteria such as Yersinia pestis. AHLs are small molecules synthesized by most gram negative bacteria, and are able to freely diffuse throughout the membrane. When the concentration of AHL producing bacteria in a specific area increases, the total concentration of AHLs diffusing into the cytoplasm also increases. Once a threshold level of AHLs are in the cytosol, transcription factors like the Vibrio fisheri LuxR or Salmonella enterica SdiA activate gene synthesis for quorum factors to be produced. In pathogenic bacteria, these signals have been shown to activate transcription of pathogenic factors.
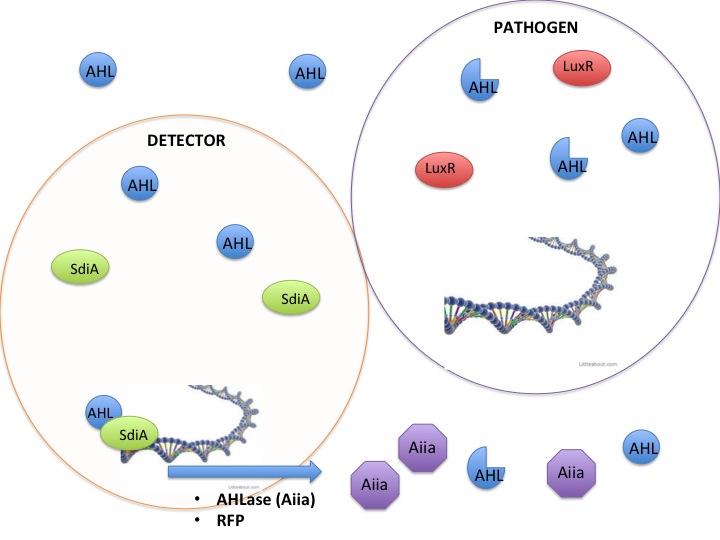
Detection and silencing of AHL producing bacteria
The SdiA transcription factor has been shown to be more sensitive to lower concentrations and a greater diversity of AHLs than its LuxR homolog. We took advantage of this extra-sensitivity to create a detection system for AHL producing bacteria, while additionally attaching the detection system to the secretion of potent AHLases such as the Aiia enzyme, and a reporter protein RFP. Using this construct we tested whether we could disrupt the quorum sensing signal AHLs in order to keep them from reaching the threshold concentration to activate the less sensitive LuxR receptor. The V. fisheri species and other pathogenic bacteria use the LuxR receptor, therefore inhibition of AHLs would keep V. fisheri from producing light. Keeping the V. fisheri from producing light is an analogous model to keeping pathogenic bacteria from being stimulated by AHLs to release their pathogenesis factors.
Inhibiting quorum sensing in bacteria has been shown to not only inhibit the transcription of pathogenesis factors, but also to keep biofilms from forming. In the history of iGEM teams have used secreted enzymes to digest biofilms that have already been made, but our use of quorum sensing inhibitors have gone a step further, and are being tested in their ability to keep bacteria from making biofilms in the first place.
Project 2: Experiments
1. Test whether the SdiA pSrgE cassette is more sensitive to extracellular AHLs than the LuxR LuxpR cassette. This will be done by placing an RFP behind the cassette, and using the plate reader to determine the fluorescence at a given concentration of RFP.
2. Use the SdiA pSrgE cassette’s sensitivity to inhibit the LuxR LuxpR system. Attach an AHLase and YFP to the more sensitive SdiA/pSrgE cassette. Co-culture the AHLase/YFP construct with a LuxR/LuxpR-RFP bacteria. The culture should turn fluoresce yellow, and not turn red.
3. Test the SdiA pSrgE cassette’s inhibition effects with Vibrio fisheri acting as a pathogen analogue. Co-culture the SdiA/pSrgE/YFP/Aiia cells with V. fisheri and use the plate reader to show that co-cultures will keep V. fisheri from emitting light at given ODs.
Project 2: Results
Applications
Medal Consideration
Bronze
- ___ Complete Judging form
- ___ Present Poster and a talk at the iGEM Jamboree: North American West Regional Jamboree at Stanford University on Oct.16, 2012
- ___ At least one new submitted BioBrick part or a new application of an existing part with quantitative documentation
 Team Registration: April 11, 2012
Team Registration: April 11, 2012
 Team Wiki
Team Wiki
Silver
- ___ Demonstrate that a newly submitted BioBrick part works as expected
- ___ Characterize the operation of at least one new BioBrick Part and enter this information in the Registry
Gold
Complete one or more of the following
- ___ Improve the function of an existing BioBrick part and enter it's information in the registry
- ___ Help another iGEM team by, for example, characterizing a part, debugging a construct, or modeling or simulating their system
- ___ Outline and detail a new approach to an issue of Human Practice in synthetic biology as it relates to your project, such as safety, security, ethics, or ownership, sharing, and innovation
 "
"