Team:Bielefeld-Germany/Project/Database
From 2012.igem.org
Contents |
One important thing while working in such a bit group like ours is the organization of probes. Therefore we designed our own database where we can make an entry for each probe we use. On this page you can find all information about this dadabase.
Design of the database
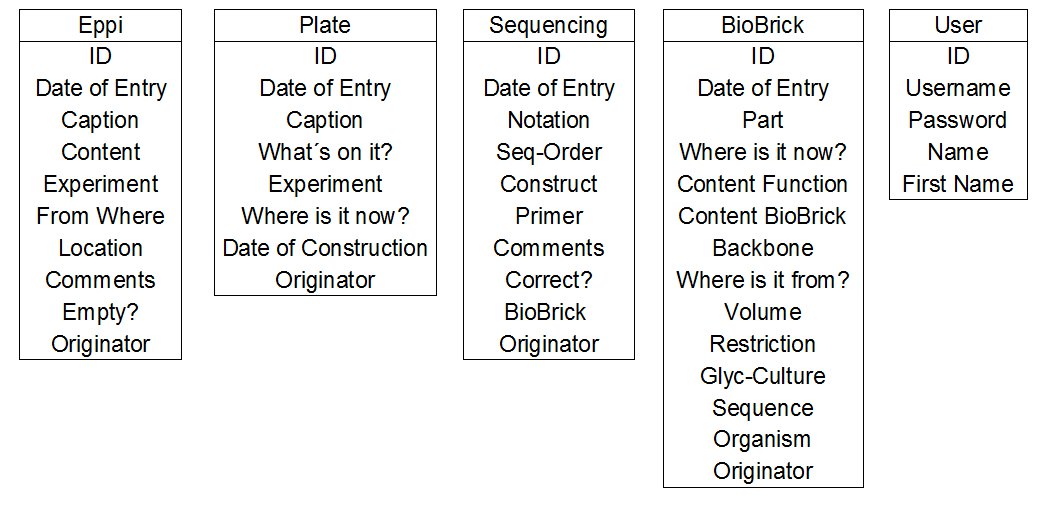
- initial design : First of all we think about the tables and the listings of the different tables we need. Here you can see our first abstract thoughts. This part of the design of a database is called the initial design of a database.
We need the User table for the login and for the allocation of entries to the team members. In the User table we erect a ID specific for each team member. This ID can be used as a foreign key in the other tables.
Additionally we want to realize the following particularities:
- For the Plates table we want to write a script witch checks if the date of construction is older than two weeks. If the week is longer ago than these two weeks, the entry will bee marked in red automatically. So we have a easy way to check if all our plate are fresh.
- For the BioBricks table we want to name our BioBricks automatically. Therefore we write a script, witch assigned a BioBrick number, according to the affiliation to either eukaryotic, bacterial, fungal or others.
- For the Eppis table it is very important, that you can always find out were a probe is. So we decides that we have to write a script witch controls if you insert a place of storage to you database entry, otherwise this entry couldn´t bee saved.
- One other very important point was, that the editing of entries is secured against mistakes, for example that one person make an entry and the other deleted it by mistake. Therefore we write a delete script witch deleted an entry not really. It saves the entry as a copy in the datebase but just shows ne newest copy of an entry. Witch this script t is possible to rebuild an entry if it is necessary.
- We did the same for the editing script.
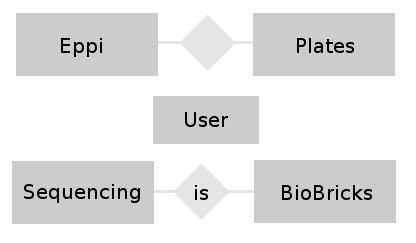
- conceptual design: In this phase of the design we try to find relations between the different tables and build a entity-relationship-model. This model helps to visualize the entities (our different tables) together with their attributes (the entries belonging to a table) and the relationship between these entities.
- We want to combine the tables Eppi with Plates and BioBrick with Sequencing.
- And of course is the User table combined to all tables because of the foreign keys.
In the graphic you can see our very easy E-R-Model.
- logically design : After this we think about the logically design of our database. In this phase the conceptual design is implemented with the help of a concrete data model. For Example as a relational design. We decided to use such a design.
For the Eppi table the relational design is shown exemplary.
| Eppi ( | ID INT(11), |
| DataID INT(11), | |
| Caption TEXT, | |
| Content TEXT, | |
| Experiment VARCHAR(255), | |
| Where2 VARCHAR(255) , | |
| Location TEXT, | |
| Comments TEXT, | |
| Empty ENUM ('N', 'J'), | |
| Create_User INT(11), | |
| Create_Date DATETIME, | |
| Edit_Date DATETIME, | |
| Delete ENUM ('N', 'J') | |
| Delete_User INT(11), | |
| Delete_Date DATETIME ) |
Programming the Database
Used Languages
For the programming language we used PHP. PHP is a general-purpose server-side scripting language and is originally designed for Web development to produce dynamic Web pages. It is one of the first developed server-side scripting languages to be embedded into an HTML source document and can also be used with many relational database management systems (RDBMS). As RDBMS and therefor for the construction of the tables in the database we used MySQL. MySQL is the world's most used open source RDBMS for the use in web applications. The SQL phrase stands for Structured Query Language. For the design of the tables and the look of them we use HTML. HTML stand for HyperText Markup Language and it is the main markup language for displaying web pages and other information that can be displayed in a web browser. So that we don´t have to design individually each table we use Cascading Style Sheets (CSS. This is a style sheet language used for describing the look and formatting of a document written in a markup language like HTML.
Realization of the design
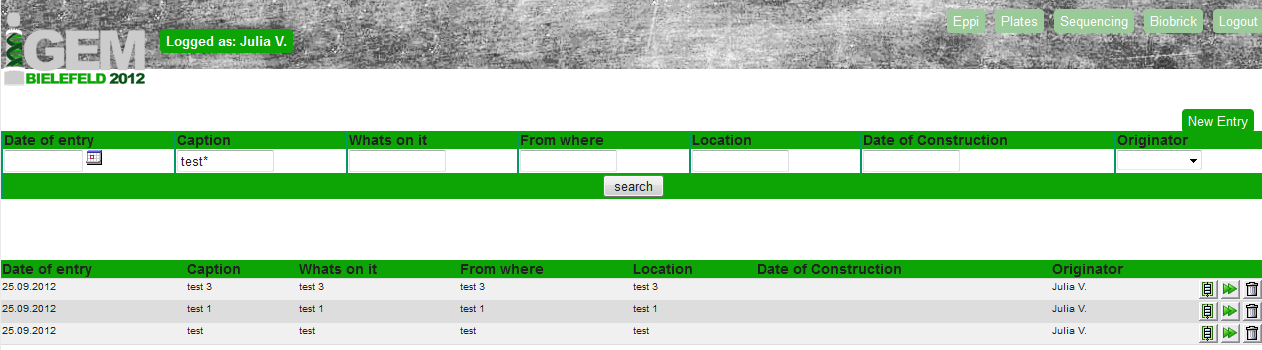
If you enter to our database you first have to log in, therefore you get a User name and a password (both saved in the User table).
After the successful log in you have the opportunity to enter to either the Eppi, the Plates, the Sequencing or the BioBrick table. The User table is only visible for the administrator.
On top of the page you see the search tool. With using _ for one letter after the search text and * for any number of letters after the search text you can specific can look of entries in the selected table.
Important Functions
Here we listed some important scripts we used. (click to the black arrows to watch the next one)
| 55px | | | | | | | | | | |
 "
"