Team:Colombia/Notebook/Journal
From 2012.igem.org
The Journal
Chitinase
To determine which chitinase is going to be used, we screened chitinase from 3 different species since the chitinase of different Vibrio spp. was not suitable or the strain was found (Alivibrio fischeri ES114, Alivibrio fischeri M11):
- Arabidopsis thaliana
- Colletotrichum spp.
- Trichoderma sbpp.
For each one we obtained accesion numbers:
- Arabidopsis thaliana: [http://www.ncbi.nlm.nih.gov/nuccore/AY099810.2 AY099810.2]
- Colletotrichum spp. : [http://www.ncbi.nlm.nih.gov/nucest/GW342409.1 GW342409.1]
- Trichoderma spp. : [http://www.ncbi.nlm.nih.gov/nucest/BM077089.2 BM077089.2]
Each one was tested using codon usage for bacteria (Translation table 11) to determine if the chitinase could be used. All of the sequences were suitable for use.
We selected Colletotrichum spp. and Trichoderma spp. sequences to design primers.
Primers
- Colletotrichum spp. :
- Trichoderma spp. :
Ralstonia solanacearum
June 8
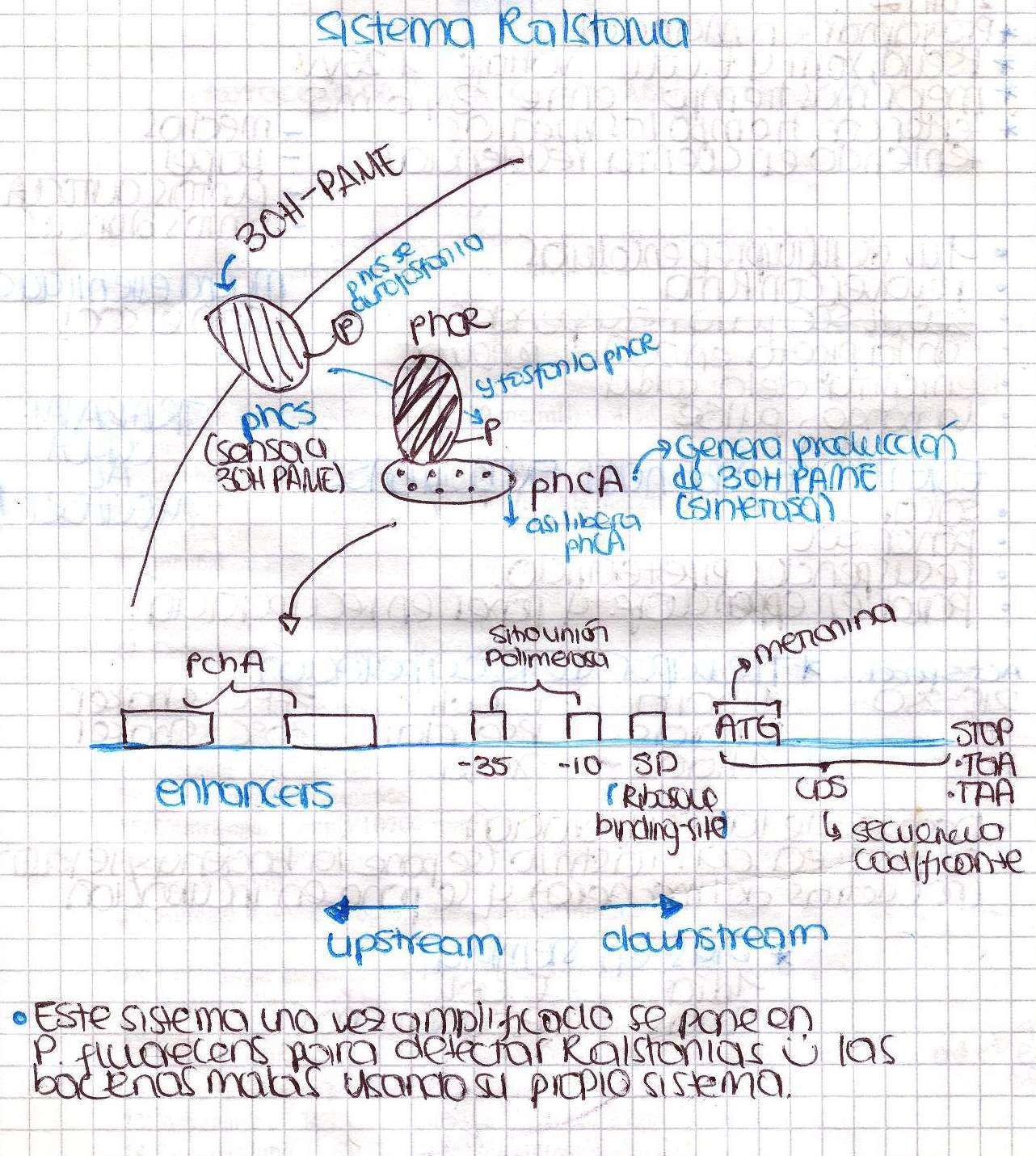
Today we have our first meeting! We basically talked about the Ralstonia solanacearum detection system and make a little research to find a proper culture medium. Here is a little sketch we made.
June 12
We decided to prepare Casamino acid-Peptone-Glucose (CPG) media, which is a rich medium and we also have all the components in the lab. This is the recipe for 1L of CPG:
| Reactives | Amount (g) |
|---|---|
| Casamino acid | 1 |
| Peptone | 10 |
| Glucose | 5 |
| Agar | 15 |
June 13
We growth in solid medium a Ralstonia solanacearum strain from the REVCO, it belongs to the phytopathogen bacteria strain store of the LAMFU, it will fully growth in 2 days, so we have to wait until then…everything is pretty easy so far.
June 15
We were going to extract DNA from Ralstonia tomorrow, so today we spend a lot of time preparing the solutions, but our Ralstonia strain didn’t growth, we are kind of upset. We also design the primers to amplify the promoter of xpsR(PxpsR) We used the sequence of Ralstonia solanacearum AW (gi:3132834)as this promoter sequence has been previously characterized by [http://www.ncbi.nlm.nih.gov/pubmed/9573161 Huang et al. in 1998.]
June 16
Today is a new day, we decided to growth a different accession of Ralstonia solanacearum and see what happens.
June 18
Our bacterium is alive! it not longer matter that today’s holyday and we are at lab…We are finally extracting DNA tomorrow, so we inoculated 5 ml of liquid CPG with a few colonies from the solid culture and left growth ON.
June 19
Today was DNA extraction day! We used the Xam’s DNA extraction protocol(see protocols) and here it is, our Ralstonia’s DNA, we think is a little bit degraded but still amplifiable…
June 20
Today we supposed to amplify all the genes for the detection system but PxpsR (those primers haven’t arrive yet). We used Fermentas Pfu (see protocols) and the Ta for each gene was calculated taking the lower primer Tm of each couple and subtracting it 2 degrees (phcS=61°C, phcR=57°C, phcA=54°C). Nonetheless, none amplified. We are no longer sure if the primers anneal in all Ralstonia strains, we haven’t consider that before.
June 21
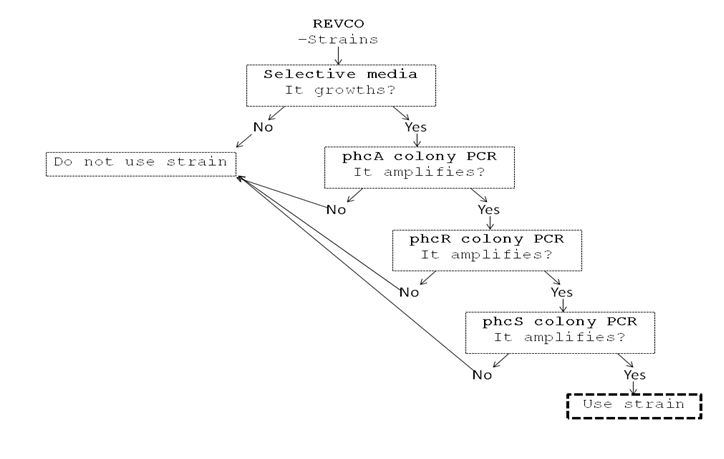
We performed a primer search in NCBI and confirmed our suspects … there are some base changes at the 3’ of our primers in the sequence of different strains of R. solanacearum…that will explain why it didn’t amplified a thing yesterday. The most conserved sequence appear to be PxpsR and then phcA, so, while PxpsR primers arrive, we will perform a screening of differtent Ralstonia solanacearum strains using phcA, then we will try to amplify the other genes from positive strains. Here is our decision tree.
June 22
Today we prepared the selective media for the screening; we had to make substantial variations due to a lack of compounds in our lab. This was the final composition:
| Reactives | Amount |
|---|---|
| Mannitol | 1g |
| Na2HPO4 | 3g |
| KH2PO4 | 3g |
| NH4Cl | 1g |
| MgSO4 | 0.25g |
| FeSO4 | 5mg |
| Crystal Violet | 3mg |
| Cycloheximide | 5mg |
| Chloramphenicol | 1mg |
| Bacitracine | 0.25mg |
| Agar | 15g |
| Distilled water | 1L |
June 24
We decided to make a pilot experiment in order to standardize the conditions of phcA amplification before to start with the massive screening . We choose #37 strain randomly for the standardization.
June 26
In order to standardize the conditions of phcA amplification before to start with the massive screening , we determine the annealing temperature (71°C) performing an in silico PCR (FAST PCR) and used that temperature to calculate a temperature gradient (form 64 to 71), reactions were carry out with or without DMSO. The results weren’t expected, none of the temperatures amplified.
June 27
We perfomed phcA PCRs using a boiling of the solid culture as a source of DNA. The results remained the same. Due to results we decided to change of strains and we growth on solid 3 new accessions.
June 28
The primers for the promoter of xpsR finally arrive! We intented to amplify this promoter due to the conservation of the secuence using the DNA previously extracted and using a Ta of 46°C. We expected a band of 389pb but we couldn’t even seen primer dimmers.
June 29
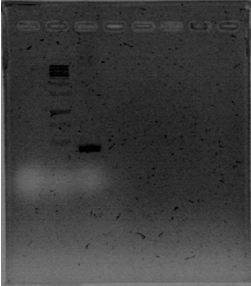
We are finally in the right direction! Today we massively performed PCR of the 3 new strains to amplify PxpsR using a temperature gradient (form 45 to 69). All but one seems to amplify in almost all the temperatures!
July 3
Now that we know that PxpsR amplifies, we attach to our decision tree and continue with phcA, we only used the strains where PxpsR amplifies. We used the temperature gradient previously named for phcA. Fortunately for us all the strains were positive.
July 5
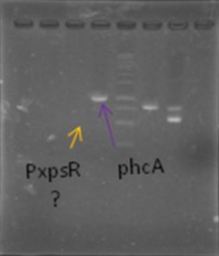
Today we try to amplify both PxpsR and phcA with Pfu in order to clone in the backbone (pBS1C3), as both genes had amplified previously at 64°C we choose that temperature as the Ta, although it didn’t work for PxpsR.
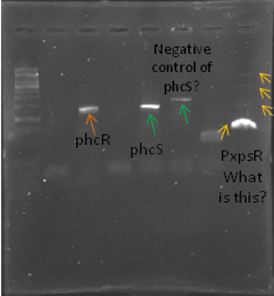
We also try to amplify phcR and phcS using a gradient temperature from 57 to 67 from the strain #75 that always amplify better than the others. Surprisingly both genes amplified just fine in all the temperatures, but there was double band, even in the highest temperatures, so we will have to cut band after amplifying with Pfu.
Finally we digest (see protocols) phcA with EcoRI and SpeI, and the backbone with EcoRI, SpeI and DpnI, and ligate (see protocols) with T4 ligase all the night.
July 6
Today we amplify PxpsR, phcR and phcS with Pfu, we used the lowest temperature for PxpsR (45°C), but once again PxpsR didn’t amplify as we expected (more than 1 band).
We also transform by electroporation (see protocols) the ligation of phcA into the backbone, we hope to see colonies in the morning.
July 7
Nothing growth into the plate! We are not sure if the restriction enzymes aren’t working well or the T4 ligase is failing… We test the restriction enzymes using as target the salicylic acid biobrick, we performed different combinations of enzymes (PstI-EcoRI, EcoRI-SpeI, XbaI-PstI, SpeI-XbaI) that excises a fragment of 1500bp. Conclusion: Everything seems to be fine with the enzymes, it must be the T4 ligase.
We also repeat the PCR for PxpsR this time using as Ta 50°C, it amplified!
July 10
We are back in the game, we have a different ligase and we are making new electrocompetent cells!
July 12
Today we are going to transform phcA+BB, phcR+BB, phcR+BB and PxpsR+BB in our just prepared competent cells. Tomorrow we will make passes of the colonies.
July 13
There are not colonies in the plates…maybe the cells aren’t that competent, we will try again with other cells…
Mathematical Models
Up to June 6
Up to June 6 we organized the variables and parameters of our design, formulated the differential equations for each of the molecules of the model and set out to look for parameter values in literature. This link is the depurated version summary of the model that should be included in the Modeling section of the wiki.
July 3
Today’s subject was the parameters to be used at differential equations that had been previously developed. In order to do this, a set of parameters, to be searched from different sources, had been assigned to each member of the team. According to the results obtained it was established to categorize each group in a range as it is shown below:
- α group: Since no value was found it would be supposed at first that all values are the same.
- γ group: For this parameters, Gabriel proposed to bear in mind the lifetime of a E. coli. As a backup, rates of normal destruction in proteins (in general) and salicylic acid could be looked for.
- Kinetic constants of reaction: We will focus on ChiA and look for any number of cellular kinetic trying to find a maximum.
- βx group: Search concentrations of the most produced protein in E. coli, RubisCo and the least produced to establish a range.
- Parameters of export and import: To reference export and import rates of any protein and then try to establish a range.
July 10
The following document has all the information we could find about parameter approximate values. It is in spanish, but it should be translated and included in the Modeling section of the wiki. File.
Our first matlab code was written today too. It is a two file code:
Code for ecuaDif.m
%Codigo que contiene todas las ecuaciones diferenciales de los procesos %dados en los 3 plasmidos
function y=ecuaDif(t,v) % Funcion que devuelve un vector y con todas las ecuaciones, tiene como parametro un vector con los parametros y un vector v
% con las variables.
QQ=10;
%------ Variables%------
Ao=v(1); %Cocentration of chitinase outside the cell Ai=v(2); %Concentration of chitinase inside the cell P=v(3); %Concentration of chitiporin C=v(4); %Concentratio of chitin binding protein (CBP) CS=v(5); %Concentration the complex CBP-s R=v(6); %Concentration of LuxR Ii=v(7); %Concentration of LuxI inside the cell Io=v(8); %Concentration of LuI outsied the cell IR=v(9); %Concentration of the complex LuxI-LuxR CI=v(10);%Concentration of the protein CI HA=v(11);%Concentration of HipA7 HB=v(12);%Concnetratio of HipB AS=v(13);%Concentration of salicylic acid Q=v(14); %Concentratio of quitin monomers
%----Parameters----%
%All the parameter came in a vector given by other function.
alfA=0.4; %Basal concentration of Chitinase inside the cell (micromolar)
alfP=0.4; %Basal concnetration of chitoporin
alfC=0.4 ; %Basal concentration of the CBP
alfR=0.4; %Basal concentration of LuxR
alfI=0.4; %Basal concentration of LuxI
alfCI=0.4; %Basal concentration of CI
alfHA=0.4; %Basal concnetration of HipA7
alfHB=0.4; %Basal concnetration of HipB
alfAS=0.4; %Basal concnetration of Salycilic acid
gammaA=0.01; %Degradation of Chitinase inside the cell
gammaP=0.01; %Degradation of chitoporin
gammaC=0.01; %Degradation concentration of the CBP
gammaR=0.01; %Degradation of LuxR
gammaI=0.01; %Degradation of LuxI
gammaCI=0.01; %Degradation of CI
gammaHA=0.01; %Degradation of HipA7
gammaHB=0.01; %Degradation of HipB
gammaAS=0.01; %Dergradation of Salycilic acid
gammaCS=0.01; %Degradation of the complex CS
mCS=5; %Kinetic constant for the formation of the complex CS
mCSQ=5; %Kinetic constant of the reaction of the complex CS with the chitin
mAQQ=5; %Kinetic constant for the reaction of the chitinase and th chitin
mIR=5; %Kinetic constant for the formation of the complex LuxILuxR
mI=5; %Constant that represent the union of the complex LuxILuxR with the promoter
mHAHB=5; %Kinetic constant for the inhibition of HipA7
betaP=10; %Max production of the chitoporin
betaA=10; %Max production of chitinase
betaI=10; %Max production of LuxI
betaCI=10; %Max production of CI
betaHB=10; %Max peoduction of HipB
betaHA=10; %Max production of HipA7
betaAS=10; %Max production of Salicylic acid
kS=1; %Constant k of the hill ecuation for the promoter promoted by S
kIR=0.5; %Constant k of the hill equiation for the promorer prmoted by the complex luxIluxR
kCI=0.1; %Cosntant k of the hill equation for the promoter promoted by CI
hS=1; %Hill constant for the promoters promoted by S
hIR=3; %Hill constant for the promoter promoted by the complex IR
hCI=2.3; %Hill constant fot the promoter CI
eA=0.5; %Exportation factor of the chitinase
jQ=0.8; %Importation factor of the chitin monomers
deltaA=0.2; %Difusion factor of the chinitanse outside the cell
eI=0.5; %Exportation factor of LuxI
jI=0.8; %Importation factor of LuxI
deltaI=0.2;%Difusion of LuxI outside the cell
Stotal= 1; %Total concentration of the sensor in the cell
numcel=100; %number of cells
%---Equations---%
S=Stotal-CS;
dC=alfC- gammaC*C - mCS*C*S; %Change of CBP
dCS=mCS*C*S- mCSQ*CS*Q-gammaCS*CS; %Change of the complex CS
dP=alfP - gammaP*P + (betaP*(S^hS))/(kS+(S^hS));%Change of chitoporin
dAi=alfA- gammaA*Ai+ (betaA*(S^hS))/(kS+(S^hS))- eA*Ai; %Change of chitinase inside the cell
dAo= eA*Ai-deltaA*Ao- mAQQ*Ao*QQ; %Change of chitinase outside the cell
dQ= jQ*P*(mAQQ*QQ*Ao)-mCSQ*CS*Q; %Change of chitin monomer inside the cell
dIi= alfI+ (betaI*(S^hS))/(kS+(S^hS)) -gammaI*Ii +jI*Io- eI*Ii- mIR*Ii*R; %Change of LuxI inside the cell
dIo= numcel*(eI*Ii-jI*Io)-deltaI*Io; %Change of LuxI outside the cell
dIR= mIR*Ii*R - mI*IR; %Change of the complex LuxI luxR
dR= alfR-gammaR*R -mIR*Ii*R; %Change of LuxR
dCI= alfCI -gammaCI*CI+ (betaCI*(CI^hCI))/(kCI+(CI^hCI))+(betaCI*(IR^hIR))/(kIR+(IR^hIR));%Change of CI
dHB=alfHB-gammaHB*HB+(betaHB*(CI^hCI))/(kCI+(CI^hCI))+(betaHB*(IR^hIR))/(kIR+(IR^hIR))-mHAHB*HA^2*HB^2; %Chanche of HipB
dHA=alfHA-gammaHA*HA+ (betaHA*(CI^hCI))/(kCI+(CI^hCI))-mHAHB*HA^2*HB^2; %Change of HipA7
dAS=alfAS-gammaAS*AS +(betaAS*(CI^hCI))/(kCI+(CI^hCI)); %Change of Salicylic acid
y1(1)=dC;
y1(2)=dCS;
y1(3)=dP;
y1(4)=dAi;
y1(5)=dAo;
y1(6)=dQ;
y1(7)=dIi;
y1(8)=dIo;
y1(9)=dIR;
y1(10)=dR;
y1(11)=dCI;
y1(12)=dHB;
y1(13)=dHA;
y1(14)=dAS;
y=y1';
end
 "
"